class: center, middle, inverse, title-slide # 2.7 — Inference for Regression ## ECON 480 • Econometrics • Fall 2021 ### Ryan Safner<br> Assistant Professor of Economics <br> <a href="mailto:safner@hood.edu"><i class="fa fa-paper-plane fa-fw"></i>safner@hood.edu</a> <br> <a href="https://github.com/ryansafner/metricsF21"><i class="fa fa-github fa-fw"></i>ryansafner/metricsF21</a><br> <a href="https://metricsF21.classes.ryansafner.com"> <i class="fa fa-globe fa-fw"></i>metricsF21.classes.ryansafner.com</a><br> --- class: inverse # Outline ### [Hypothesis Testing](#3) ### [Digression: p-Values and the Philosophy of Science](#33) ### [Hypothesis Testing by Simulation, with *infer*](#54) ### [Classical Statistical Inference (What R Calculates)](#79) ### [The Use and Abuse of `\(p\)`-values](#93) --- class: inverse, center, middle # Hypothesis Testing --- # Estimation and Hypothesis Testing I .pull-left[ .smallest[ - We want to **test** if our estimates are .hi[statistically significant] and they describe the population - this is the “bread and butter” of using inferential statistics .bg-washed-green.b--dark-green.ba.bw2.br3.shadow-5.ph4.mt5[ .green[**Examples**]: - Does reducing class size actually improve test scores? - Do more years of education increase your wages? - Is the gender wage gap between men and women 23%? ] ] ] .pull-right[ .center[  ] - .hi-purple[All modern science is built upon statistical hypothesis testing, so understand it well!] ] --- # Estimation and Hypothesis Testing II .smallest[ - Note, we can test a lot of hypotheses about a lot of population parameters, e.g. - A population mean `\(\mu\)` - <span class="green">**Example**: average height of adults</span> - A population proportion `\(p\)` - <span class="green">**Example**: percent of voters who voted for Trump</span> - A difference in population means `\(\mu_A-\mu_B\)` - <span class="green">**Example**: difference in average wages of men vs. women</span> - A difference in population proportions `\(p_A-p_B\)` - <span class="green">**Example**: difference in percent of patients reporting symptoms of drug A vs B</span> - We will focus on hypotheses about .hi-purple[population regression slope] `\((\hat{\beta}_1)\)`, i.e. the .hi-purple[causal effect]<sup>.magenta[†]</sup> of `\(X\)` on `\(Y\)` ] .footnote[<sup>.magenta[†]</sup> With a model this simple, it's almost certainly **not** causal, but this is the ultimate direction we are heading...] --- # Null and Alternative Hypotheses I - All scientific inquiries begin with a .hi[null hypothesis] `\((H_0)\)` that proposes a specific value of a population parameter - Notation: add a subscript 0: `\(\beta_{1,0}\)` (or `\(\mu_0\)`, `\(p_0\)`, etc) -- - We suggest an .hi[alternative hypothesis] `\((H_a)\)`, often the one we hope to verify - Note, can be multiple alternative hypotheses: `\(H_1, H_2, \ldots , H_n\)` -- - Ask: .hi-purple["Does our data (sample) give us sufficient evidence to reject `\\(H_0\\)` in favor of `\\(H_a\\)`?"] - Note: **the test is *always* about** `\(\mathbf{H_0}\)`! - See if we have sufficient evidence to reject the status quo --- # Null and Alternative Hypotheses II - Null hypothesis assigns a value (or a range) to a population parameter - e.g. `\(\beta_1=2\)` or `\(\beta_1 \leq 20\)` - .hi-purple[Most common is `\\(\beta_1=0\\)`] `\(\implies\)` `\(X\)` has no effect on `\(Y\)` (no slope for a line) - Note: always an equality! -- - Alternative hypothesis must mathematically *contradict* the null hypothesis - e.g. `\(\beta_1 \neq 2\)` or `\(\beta_1 > 20\)` or `\(\beta_1 \neq 0\)` - Note: always an inequality! -- - Alternative hypotheses come in two forms: 1. .hi-purple[One-sided alternative]: `\(\beta_1 >H_0\)` or `\(\beta_1< H_0\)` 2. .hi-purple[Two-sided alternative]: `\(\beta_1 \neq H_0\)` - Note this means either `\(\beta_1 < H_0\)` or `\(\beta_1 > H_0\)` --- # Components of a Valid Hypothesis Test - All statistical hypothesis tests have the following components: -- 1. A .hi-purple[null hypothesis, `\\(H_0\\)`] -- 2. An .hi-purple[alternative hypothesis, `\\(H_a\\)`] -- 3. A .hi-purple[test statistic] to determine if we reject `\(H_0\)` when the statistic reaches a "critical value" - Beyond the critical value is the "rejection region", sufficient evidence to reject `\(H_0\)` -- 4. A .hi-purple[conclusion] whether or not to reject `\(H_0\)` in favor of `\(H_a\)` --- # Type I and Type II Errors I .pull-left[ .smallest[ - Sample statistic `\((\hat{\beta_1})\)` will rarely be exactly equal to the hypothesized parameter `\((\beta_1)\)` - Difference between observed statistic and true parameter could be because: - .hi-turquoise[Parameter is *not* the hypothesized value] - `\(H_0\)` is *false* - .hi-turquoise[Parameter is truly hypothesized value but *sampling variability* gave us a different estimate] - `\(H_0\)` is *true* - .hi-purple[We cannot distinguish between these two possibilities with any certainty] ] ] .pull-right[ .center[  ] ] --- # Type I and Type II Errors II .pull-left[ .smaller[ - We can interpret our estimates probabilistically as commiting one of two types of error: 1. .hi[Type I error (false positive)]: rejecting `\(H_0\)` when it is in fact true - Believing we found an important result when there is truly no relationship 2. .hi[Type II error (false negative)]: failing to reject `\(H_0\)` when it is in fact false - Believing we found nothing when there was truly a relationship to find ] ] .pull-right[ .center[  ] ] --- # Type I and Type II Errors III .center[  ] - Depending on context, committing one type of error may be more serious than the other --- # Type I and Type II Errors IV .center[  ] .smallest[ - Anglo-American common law *presumes* defendant is innocent: `\(H_0\)` ] -- .smallest[ - Jury judges whether the evidence presented against the defendant is plausible *assuming the defendant were in fact innocent* ] -- .smallest[ - If highly improbable (beyond a “reasonable doubt”): sufficient evidence to reject `\(H_0\)` and convict ] --- # Type I and Type II Errors V .left-column[ .center[  William Blackstone (1723-1780) ] ] .right-column[ > "It is better that ten guilty persons escape than that one innocent suffer." - Type I error is worse than a Type II error in law! ] .source[Blackstone, William, 1765-1770, *Commentaries on the Laws of England*] --- # Type I and Type II Errors VI .center[  ] --- # Type I and Type II Errors VI .center[  ] --- # Significance Level, `\(\alpha\)`, and Confidence Level `\(1-\alpha\)` - The .hi[significance level, `\\(\alpha\\)`], is the probability of a **Type I error** `$$\alpha=P(\text{Reject } H_0 | H_0 \text{ is true})$$` -- - The .hi[confidence level] is defined as .hi[`\\((1-\alpha)\\)`] - Specify *in advance* an `\(\alpha\)`-level (0.10, 0.05, 0.01) with associated confidence level (90%, 95%, 99%) -- - The probability of a **Type II error** is defined as `\(\beta\)`: `$$\beta=P(\text{Don't reject } H_0 | H_0 \text{ is false})$$` --- # `\(\alpha\)` and `\(\beta\)` .center[  ] --- # Power and p-values - The statistical .hi[power of the test] is `\((1-\beta)\)`: the probability of correctly rejecting `\(H_0\)` when `\(H_0\)` is in fact false (e.g. convicting a guilty person) `$$\text{Power} = 1- \beta = P(\text{Reject }H_0|H_0 \text{ is false})$$` -- - The .hi[`\\(p\\)`-value] or .hi[significance probability] is the probability that, if the null hypothesis were true, the test statistic from any sample will be *at least as extreme* as the test statistic from *our* sample `$$p(\delta \geq \delta_i|H_0 \text{ is true})$$` - where `\(\delta\)` represents some test statistic - `\(\delta_i\)` is the test statistic we observe in our sample - More on this in a bit --- # p-Values and Statistical Significance - After running our test, we need to make a *decision* between the competing hypotheses - Compare `\(p\)`-value with *pre-determined* `\(\alpha\)` (commonly, `\(\alpha=0.05\)`, 95% confidence level) - If `\(p<\alpha\)`: .hi-purple[statistically significant] evidence sufficient to *reject* `\(H_0\)` in favor of `\(H_a\)` - Note this does **not** mean `\(H_a\)` is true! We merely have *rejected* `\(H_0\)`! - If `\(p \geq \alpha\)`: *insufficient* evidence to reject `\(H_0\)` - Note this does **not** mean `\(H_0\)` is true! We merely have *failed* to *reject* `\(H_0\)`! --- class: inverse, center, middle # Digression: p-Values and the Philosophy of Science --- # Hypothesis Testing and the Philosophy of Science I .left-column[ .center[  Sir Ronald A. Fisher (1890—1962) ] ] .right-column[ > "The null hypothesis is never proved or established, but is possibly disproved, in the course of experimentation. Every experiment may be said to exist only in order to give the facts a chance of disproving the null hypothesis." 1931, *The Design of Experiments* ] --- # Hypothesis Testing and the Philosophy of Science I .pull-left[ .smallest[ - Modern philosophy of science is largely based off of hypothesis testing and .hi-purple[falsifiability], which form the "Scientific Method"<sup>.magenta[†]</sup> - For something to be "scientific", it must be .hi-purple[falsifiable], or at least .hi-purple[testable] - Hypotheses can be *corroborated* with evidence, but always *tentative* until falsified by data in suggesting an alternative hypothesis > **"All swans are white"** is a hypothesis rejected upon discovery of a single black swan ] ] .pull-right[ .center[ .polaroid[] ] ] .footnote[<sup>.magenta[†]</sup> Note: economics is a very different kind of "science" with a different methodology!] --- # Hypothesis Testing and p-Values - Hypothesis testing, confidence intervals, and p-values are probably the hardest thing to understand in statistics .center[ <iframe src="https://fivethirtyeight.abcnews.go.com/video/embed/56150342" width="640" height="360" scrolling="no" style="border:none;" allowfullscreen></iframe> [Fivethirtyeight: Not Even Scientists Can Easily Explain P-values](https://fivethirtyeight.com/features/not-even-scientists-can-easily-explain-p-values/)] --- # Hypothesis Testing: Which Test? I - Rigorous course on statistics ([**ECMG 212**](http://ryansafner.com/courses/ecmg212) or **MATH 112**) will spend weeks going through different types of tests: - Sample mean; difference of means - Proportion; difference of proportions - Z-test vs t-test - 1 sample vs. 2 samples - `\(\chi^2\)` test --- # Hypothesis Testing: Which Test? II .center[  ] --- # There is Only One Test - Fortunately, some clever statisticians realized [“**there is only one test**”](https://allendowney.blogspot.com/2011/05/there-is-only-one-test.html) and some built a nice `R` package called `infer` 1. **Calculate** a statistic, `\(\delta_i\)`<sup>.magenta[†]</sup>, from a sample of data 2. **Simulate** a world where `\(\delta\)` is null `\((H_0)\)` 3. **Examine** the distribution of `\(\delta\)` across the null world 4. **Calculate** the probability that `\(\delta_i\)` could exist in the null world 5. **Decide** if `\(\delta_i\)` is statistically significant .footnote[<sup>.magenta[†]</sup> `\\(\delta\\)` can stand in for any test-statistic in any hypothesis test! For our purposes, `\\(\delta\\)` is the slope of our regression sample, `\\(\hat{\beta}_1\\)`.] --- # Elements of a Hypothesis Test .center[  [Alan Downey: “There is still only one test”](https://allendowney.blogspot.com/2016/06/there-is-still-only-one-test.html) ] --- # Hypothesis Testing with the infer Package I - R naturally runs the following hypothesis test on any regression as part of `lm()`: `$$\begin{align*} H_0: \; & \beta_1=0\\ H_1: \; & \beta_1 \neq 0 \end{align*}$$` - `infer` allows you to run through these steps manually to understand the process: -- 1. `specify()` a model -- 2. `hypothesize()` the null -- 3. `generate()` simulations of the null world -- 4. `calculate()` the `\(p\)`-value -- 5. `visualize()` with a histogram (optional) --- # Hypothesis Testing with the infer Package II .center[  ] --- # Hypothesis Testing with the infer Package II .center[  ] --- # Hypothesis Testing with the infer Package II .center[  ] --- # Hypothesis Testing with the infer Package II .center[  ] --- # Hypothesis Testing with the infer Package II .center[  ] --- # Hypothesis Testing with the infer Package II .center[  ] --- # Classical Inference: Critical Values of Test Statistic - .hi-purple[Test statistic `\\((\delta)\\)`]: measures **how far what we observed in our sample `\((\hat{\beta_1})\)` is from what we would expect if the null hypothesis were true `\((\beta_1=0)\)`** - Calculated from a sampling distribution of the estimator (i.e. `\(\hat{\beta_1})\)` - In econometrics, we use `\(t\)`-distributions which have `\(n-k-1\)` degrees of freedom<sup>.magenta[†]</sup> - .hi-purple[Rejection region]: if the test statistic reaches a .hi-purple["critical value"] of `\(\delta\)`, then we **reject** the null hypothesis .footnote[<sup>.magenta[†]</sup> Again, see last class's [appendix](/content/2.6-content#appendix) for more on the t-distribution. `\\(k\\)` is the number of independent variables our model has, in this case, with just one `\\(X\\)`, `\\(k=1\\)`. We use two degrees of freedom to calculate `\\(\hat{\beta_0}\\)` and `\\(\hat{\beta_1}\\)`, hence we have `\\(n-2\\)` df.] --- class: inverse, center, middle # Hypothesis Testing by Simulation, with *infer* --- # Imagine a Null World, where `\(H_0\)` is True .center[  Our world, and a world where `\(\beta_1=0\)` by assumption. ] --- # Comparing the Worlds I - From that null world where `\(H_0: \, \beta_1=0\)` is true, we **simulate** another sample and calculate OLS estimators again -- .pull-left[ ## Our Sample .smallest[ <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["term"],"name":[1],"type":["chr"],"align":["left"]},{"label":["estimate"],"name":[2],"type":["dbl"],"align":["right"]},{"label":["std.error"],"name":[3],"type":["dbl"],"align":["right"]},{"label":["statistic"],"name":[4],"type":["dbl"],"align":["right"]},{"label":["p.value"],"name":[5],"type":["dbl"],"align":["right"]}],"data":[{"1":"(Intercept)","2":"698.932952","3":"9.4674914","4":"73.824514","5":"6.569925e-242"},{"1":"str","2":"-2.279808","3":"0.4798256","4":"-4.751327","5":"2.783307e-06"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] ] -- .pull-right[ ## Another Sample .smallest[ <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["term"],"name":[1],"type":["chr"],"align":["left"]},{"label":["estimate"],"name":[2],"type":["dbl"],"align":["right"]},{"label":["std.error"],"name":[3],"type":["dbl"],"align":["right"]},{"label":["statistic"],"name":[4],"type":["dbl"],"align":["right"]},{"label":["p.value"],"name":[5],"type":["dbl"],"align":["right"]}],"data":[{"1":"(Intercept)","2":"647.8027952","3":"9.7147718","4":"66.6822452","5":"6.997699e-225"},{"1":"str","2":"0.3235038","3":"0.4923581","4":"0.6570499","5":"5.115104e-01"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] ] --- # Comparing the Worlds II .smallest[ - From that null world where `\(H_0: \, \beta_1=0\)` is true, let's **simulate 1,000** samples and calculate slope `\((\hat{\beta_1})\)` for each ] .quitesmall[ <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["sample"],"name":[1],"type":["int"],"align":["right"]},{"label":["slope"],"name":[2],"type":["dbl"],"align":["right"]}],"data":[{"1":"1","2":"-0.3027333296"},{"1":"2","2":"-0.3624481355"},{"1":"3","2":"0.6448518690"},{"1":"4","2":"-0.0745971847"},{"1":"5","2":"0.5969444290"},{"1":"6","2":"0.5505335318"},{"1":"7","2":"0.5927466147"},{"1":"8","2":"0.0572148658"},{"1":"9","2":"-0.0989989073"},{"1":"10","2":"0.8043957511"},{"1":"11","2":"-0.2438170364"},{"1":"12","2":"-0.6982879943"},{"1":"13","2":"0.9830641411"},{"1":"14","2":"0.3799867860"},{"1":"15","2":"-0.5600627540"},{"1":"16","2":"-0.8889196263"},{"1":"17","2":"1.0043191256"},{"1":"18","2":"-1.0952995001"},{"1":"19","2":"-0.3504492414"},{"1":"20","2":"-0.2644372417"},{"1":"21","2":"0.6735202365"},{"1":"22","2":"0.4258348785"},{"1":"23","2":"0.4257561509"},{"1":"24","2":"0.5485410213"},{"1":"25","2":"-0.0687439325"},{"1":"26","2":"0.2811513023"},{"1":"27","2":"-0.0385227840"},{"1":"28","2":"0.1491024436"},{"1":"29","2":"-0.1708824013"},{"1":"30","2":"0.3645308264"},{"1":"31","2":"0.1487846906"},{"1":"32","2":"-0.4704167216"},{"1":"33","2":"0.2000187396"},{"1":"34","2":"-0.7431958248"},{"1":"35","2":"-0.7524093345"},{"1":"36","2":"0.5922846484"},{"1":"37","2":"0.5364921247"},{"1":"38","2":"-0.2865572069"},{"1":"39","2":"1.0791355870"},{"1":"40","2":"0.0944318106"},{"1":"41","2":"-0.7922212533"},{"1":"42","2":"-0.3037078900"},{"1":"43","2":"0.1137069602"},{"1":"44","2":"0.0413982484"},{"1":"45","2":"0.2312012465"},{"1":"46","2":"0.1886313805"},{"1":"47","2":"0.1085040590"},{"1":"48","2":"-0.1074460833"},{"1":"49","2":"0.1366258816"},{"1":"50","2":"0.4829654509"},{"1":"51","2":"-0.0579507213"},{"1":"52","2":"0.1524977707"},{"1":"53","2":"-0.5264828719"},{"1":"54","2":"-0.5274925981"},{"1":"55","2":"1.0603685685"},{"1":"56","2":"-0.2190550697"},{"1":"57","2":"-0.1332383673"},{"1":"58","2":"-0.7412044394"},{"1":"59","2":"0.0658030770"},{"1":"60","2":"-0.1906063232"},{"1":"61","2":"-0.4053914941"},{"1":"62","2":"0.4320062259"},{"1":"63","2":"-0.3331640888"},{"1":"64","2":"0.5856592323"},{"1":"65","2":"0.1782529945"},{"1":"66","2":"0.1285081496"},{"1":"67","2":"-0.8226654100"},{"1":"68","2":"-0.7163772466"},{"1":"69","2":"-0.4969688217"},{"1":"70","2":"-0.1020706897"},{"1":"71","2":"-0.1633113938"},{"1":"72","2":"0.1609282359"},{"1":"73","2":"0.0180383888"},{"1":"74","2":"0.4506401663"},{"1":"75","2":"-0.6868421097"},{"1":"76","2":"-0.6809684741"},{"1":"77","2":"0.0816358684"},{"1":"78","2":"-0.1282760521"},{"1":"79","2":"0.1381140570"},{"1":"80","2":"-0.3547485496"},{"1":"81","2":"0.3584359218"},{"1":"82","2":"0.5336725817"},{"1":"83","2":"0.9146378308"},{"1":"84","2":"-0.2621153329"},{"1":"85","2":"0.2721682102"},{"1":"86","2":"-0.6159636093"},{"1":"87","2":"0.2529403107"},{"1":"88","2":"-0.0556586038"},{"1":"89","2":"-0.2212322538"},{"1":"90","2":"-0.4652816510"},{"1":"91","2":"-0.2937945686"},{"1":"92","2":"0.5985966733"},{"1":"93","2":"0.3344954834"},{"1":"94","2":"-0.5160325363"},{"1":"95","2":"0.3276724468"},{"1":"96","2":"-0.0564833668"},{"1":"97","2":"-0.0421615806"},{"1":"98","2":"-0.5811619827"},{"1":"99","2":"0.2193463838"},{"1":"100","2":"-0.5333419241"},{"1":"101","2":"-0.8431380688"},{"1":"102","2":"0.1506860439"},{"1":"103","2":"0.1637072816"},{"1":"104","2":"-0.1487773287"},{"1":"105","2":"-0.4373486251"},{"1":"106","2":"-0.0842235898"},{"1":"107","2":"0.0943201190"},{"1":"108","2":"-1.2236190643"},{"1":"109","2":"0.3436632069"},{"1":"110","2":"-0.3547939336"},{"1":"111","2":"0.3930861442"},{"1":"112","2":"0.2941607396"},{"1":"113","2":"-0.2595312818"},{"1":"114","2":"-0.4815194944"},{"1":"115","2":"-0.7165579781"},{"1":"116","2":"0.0167572690"},{"1":"117","2":"-0.5354641069"},{"1":"118","2":"-0.1267083518"},{"1":"119","2":"-0.1513303882"},{"1":"120","2":"0.1086866417"},{"1":"121","2":"0.6709136395"},{"1":"122","2":"-0.4377728570"},{"1":"123","2":"0.3809331235"},{"1":"124","2":"-0.1157013965"},{"1":"125","2":"-0.2418237400"},{"1":"126","2":"0.7078070024"},{"1":"127","2":"-0.1164991487"},{"1":"128","2":"0.4317451534"},{"1":"129","2":"-0.0458990595"},{"1":"130","2":"0.1116328590"},{"1":"131","2":"-0.1999808200"},{"1":"132","2":"-0.6065754780"},{"1":"133","2":"-0.4195832606"},{"1":"134","2":"-0.5828723048"},{"1":"135","2":"-0.6709104090"},{"1":"136","2":"-0.4285878659"},{"1":"137","2":"-0.4002455953"},{"1":"138","2":"0.0492101890"},{"1":"139","2":"0.4051208853"},{"1":"140","2":"0.6914279272"},{"1":"141","2":"-0.2190922702"},{"1":"142","2":"-0.2070134294"},{"1":"143","2":"0.3380788023"},{"1":"144","2":"-1.0639289107"},{"1":"145","2":"0.4194562518"},{"1":"146","2":"0.4857529052"},{"1":"147","2":"0.4096885468"},{"1":"148","2":"-0.0571035720"},{"1":"149","2":"0.0762046846"},{"1":"150","2":"-0.4321140263"},{"1":"151","2":"-0.6598680971"},{"1":"152","2":"0.4725596961"},{"1":"153","2":"-0.1974981090"},{"1":"154","2":"-0.8583200841"},{"1":"155","2":"-0.7183021394"},{"1":"156","2":"-0.0710560758"},{"1":"157","2":"0.6322994721"},{"1":"158","2":"0.0465411605"},{"1":"159","2":"0.0247907462"},{"1":"160","2":"-0.4736649946"},{"1":"161","2":"1.0651750483"},{"1":"162","2":"0.9381308507"},{"1":"163","2":"-0.1469675461"},{"1":"164","2":"0.7449440193"},{"1":"165","2":"0.6979671516"},{"1":"166","2":"1.6187923046"},{"1":"167","2":"-0.0046971701"},{"1":"168","2":"-0.1649930409"},{"1":"169","2":"-1.2552583181"},{"1":"170","2":"0.1771354334"},{"1":"171","2":"0.4989736964"},{"1":"172","2":"0.7383312051"},{"1":"173","2":"0.1637881407"},{"1":"174","2":"-0.2158255381"},{"1":"175","2":"-0.2763201693"},{"1":"176","2":"-0.1703996981"},{"1":"177","2":"0.0077105885"},{"1":"178","2":"0.6724211774"},{"1":"179","2":"0.2744105119"},{"1":"180","2":"-0.2881725867"},{"1":"181","2":"0.8074358831"},{"1":"182","2":"-0.7135852963"},{"1":"183","2":"-0.2468922495"},{"1":"184","2":"-0.5405710818"},{"1":"185","2":"-0.4998377738"},{"1":"186","2":"0.3793388292"},{"1":"187","2":"-0.0149121583"},{"1":"188","2":"-0.5635348827"},{"1":"189","2":"0.2562168257"},{"1":"190","2":"0.2447952369"},{"1":"191","2":"0.8683702113"},{"1":"192","2":"-0.9042100694"},{"1":"193","2":"-0.2832883497"},{"1":"194","2":"0.0056469255"},{"1":"195","2":"0.5793692441"},{"1":"196","2":"-0.3416621676"},{"1":"197","2":"0.2980236795"},{"1":"198","2":"0.2296454228"},{"1":"199","2":"0.3430989246"},{"1":"200","2":"0.9317725905"},{"1":"201","2":"0.0149048861"},{"1":"202","2":"1.1089261112"},{"1":"203","2":"-0.7922462995"},{"1":"204","2":"1.2110017589"},{"1":"205","2":"-0.4321451036"},{"1":"206","2":"-0.0286398795"},{"1":"207","2":"0.0445971887"},{"1":"208","2":"-0.4380067947"},{"1":"209","2":"0.4953999077"},{"1":"210","2":"-0.4436713420"},{"1":"211","2":"-0.6588995821"},{"1":"212","2":"0.4120906573"},{"1":"213","2":"0.4319351385"},{"1":"214","2":"-0.0755990991"},{"1":"215","2":"-0.6421513674"},{"1":"216","2":"0.0006774872"},{"1":"217","2":"-0.3274745847"},{"1":"218","2":"-0.3850812148"},{"1":"219","2":"-0.0395342413"},{"1":"220","2":"0.3780242481"},{"1":"221","2":"0.9133237564"},{"1":"222","2":"0.0715788966"},{"1":"223","2":"0.1788601015"},{"1":"224","2":"0.8488749702"},{"1":"225","2":"-1.1269719338"},{"1":"226","2":"-0.0676708921"},{"1":"227","2":"-0.1082583943"},{"1":"228","2":"-0.1695696717"},{"1":"229","2":"0.3494798868"},{"1":"230","2":"-0.2413092290"},{"1":"231","2":"-0.1844788349"},{"1":"232","2":"-0.2286293964"},{"1":"233","2":"0.3837137381"},{"1":"234","2":"-0.8109756146"},{"1":"235","2":"0.1548992826"},{"1":"236","2":"-0.8742979765"},{"1":"237","2":"0.7252951983"},{"1":"238","2":"0.7280670205"},{"1":"239","2":"-0.5596334603"},{"1":"240","2":"0.0959082591"},{"1":"241","2":"0.6035880457"},{"1":"242","2":"-1.2914046961"},{"1":"243","2":"-1.1390259251"},{"1":"244","2":"-0.3629237080"},{"1":"245","2":"0.7025054234"},{"1":"246","2":"-0.1722162786"},{"1":"247","2":"-1.2461677691"},{"1":"248","2":"-0.0792155746"},{"1":"249","2":"-0.3183164278"},{"1":"250","2":"0.5325205043"},{"1":"251","2":"-0.4677313243"},{"1":"252","2":"-0.6547476875"},{"1":"253","2":"0.9941310234"},{"1":"254","2":"0.4376542498"},{"1":"255","2":"-0.2977593164"},{"1":"256","2":"-0.2612792656"},{"1":"257","2":"-0.2790817845"},{"1":"258","2":"1.1010107390"},{"1":"259","2":"-0.5201719621"},{"1":"260","2":"0.7195326065"},{"1":"261","2":"0.1901959636"},{"1":"262","2":"-1.4045825990"},{"1":"263","2":"0.4412290796"},{"1":"264","2":"-0.3164639912"},{"1":"265","2":"-1.0789785047"},{"1":"266","2":"-1.0402595469"},{"1":"267","2":"-0.4845019509"},{"1":"268","2":"-1.1979573118"},{"1":"269","2":"0.1583899937"},{"1":"270","2":"-0.5700130593"},{"1":"271","2":"1.1281297386"},{"1":"272","2":"-0.5902223495"},{"1":"273","2":"0.1812005139"},{"1":"274","2":"-0.3072893490"},{"1":"275","2":"0.2716968062"},{"1":"276","2":"0.0102006470"},{"1":"277","2":"-0.0492102141"},{"1":"278","2":"-0.7896264717"},{"1":"279","2":"-0.0576766897"},{"1":"280","2":"-0.4360669545"},{"1":"281","2":"-0.6229681677"},{"1":"282","2":"0.2989971852"},{"1":"283","2":"-0.7533242740"},{"1":"284","2":"0.1356750930"},{"1":"285","2":"0.3657509752"},{"1":"286","2":"-0.0384944823"},{"1":"287","2":"-0.5377758326"},{"1":"288","2":"0.7557053194"},{"1":"289","2":"1.0111331607"},{"1":"290","2":"1.2276047813"},{"1":"291","2":"-0.3023652943"},{"1":"292","2":"-0.3835871756"},{"1":"293","2":"-0.3316231176"},{"1":"294","2":"0.3933882147"},{"1":"295","2":"-0.2363547293"},{"1":"296","2":"-0.0278210880"},{"1":"297","2":"0.1960812004"},{"1":"298","2":"-0.0672208329"},{"1":"299","2":"-0.3124411487"},{"1":"300","2":"-0.1320277023"},{"1":"301","2":"-0.0645041233"},{"1":"302","2":"-0.3035195887"},{"1":"303","2":"0.4128969089"},{"1":"304","2":"-0.1908608998"},{"1":"305","2":"-0.3528339182"},{"1":"306","2":"-0.3075531503"},{"1":"307","2":"-0.2236941643"},{"1":"308","2":"0.1897503066"},{"1":"309","2":"0.1725995717"},{"1":"310","2":"0.0661234288"},{"1":"311","2":"-0.5043405181"},{"1":"312","2":"0.1146217031"},{"1":"313","2":"0.3712050130"},{"1":"314","2":"-0.3902320748"},{"1":"315","2":"-0.6990006381"},{"1":"316","2":"0.3119577301"},{"1":"317","2":"-0.5031561987"},{"1":"318","2":"-0.2414996847"},{"1":"319","2":"0.2082278812"},{"1":"320","2":"0.6118164146"},{"1":"321","2":"-0.0029678223"},{"1":"322","2":"1.2134758553"},{"1":"323","2":"-0.3761444387"},{"1":"324","2":"0.3657926984"},{"1":"325","2":"0.4357372227"},{"1":"326","2":"-0.1062249183"},{"1":"327","2":"-0.1604572625"},{"1":"328","2":"-0.1748775875"},{"1":"329","2":"0.0844831925"},{"1":"330","2":"0.4827772235"},{"1":"331","2":"0.0175062456"},{"1":"332","2":"-0.2963983311"},{"1":"333","2":"0.5072565874"},{"1":"334","2":"-0.1015553567"},{"1":"335","2":"-0.7852142646"},{"1":"336","2":"0.1426630694"},{"1":"337","2":"-0.0095408760"},{"1":"338","2":"0.0256217487"},{"1":"339","2":"0.0163295631"},{"1":"340","2":"-0.4543958813"},{"1":"341","2":"-0.5283615843"},{"1":"342","2":"-0.0356401536"},{"1":"343","2":"0.5269037298"},{"1":"344","2":"0.0001523661"},{"1":"345","2":"0.0093125265"},{"1":"346","2":"0.1745565547"},{"1":"347","2":"-0.4415714193"},{"1":"348","2":"0.1002289472"},{"1":"349","2":"0.3705850215"},{"1":"350","2":"-0.5696994098"},{"1":"351","2":"0.4271106247"},{"1":"352","2":"0.2941078599"},{"1":"353","2":"0.1202864466"},{"1":"354","2":"0.7176024797"},{"1":"355","2":"0.5197124659"},{"1":"356","2":"-0.8967168402"},{"1":"357","2":"-0.7446170061"},{"1":"358","2":"-0.7609270813"},{"1":"359","2":"0.2643493885"},{"1":"360","2":"-0.6964666708"},{"1":"361","2":"-0.6471088575"},{"1":"362","2":"-0.1837290259"},{"1":"363","2":"1.1325575373"},{"1":"364","2":"0.2087691406"},{"1":"365","2":"0.1293827450"},{"1":"366","2":"0.3834041959"},{"1":"367","2":"0.4015490486"},{"1":"368","2":"0.1024085504"},{"1":"369","2":"0.0367956013"},{"1":"370","2":"-0.6959544040"},{"1":"371","2":"-0.2391991592"},{"1":"372","2":"-0.3294021408"},{"1":"373","2":"-0.2446487134"},{"1":"374","2":"-0.3108560644"},{"1":"375","2":"-0.1755633209"},{"1":"376","2":"0.1671181825"},{"1":"377","2":"0.0614592886"},{"1":"378","2":"0.8926627571"},{"1":"379","2":"-0.4032905210"},{"1":"380","2":"-0.2141471165"},{"1":"381","2":"0.0830308861"},{"1":"382","2":"0.6276670651"},{"1":"383","2":"-0.1848489143"},{"1":"384","2":"-0.2668088485"},{"1":"385","2":"1.1735869645"},{"1":"386","2":"-0.0166500067"},{"1":"387","2":"0.2283216982"},{"1":"388","2":"-0.5535911745"},{"1":"389","2":"-0.2866882792"},{"1":"390","2":"-0.3472958420"},{"1":"391","2":"0.0671247922"},{"1":"392","2":"0.6524367846"},{"1":"393","2":"-0.1238640100"},{"1":"394","2":"0.0426556871"},{"1":"395","2":"-0.2588256285"},{"1":"396","2":"-0.5535244439"},{"1":"397","2":"-0.3264337787"},{"1":"398","2":"0.5820149450"},{"1":"399","2":"-0.0205543342"},{"1":"400","2":"0.8563864635"},{"1":"401","2":"0.8365150891"},{"1":"402","2":"0.3322880371"},{"1":"403","2":"0.7426486264"},{"1":"404","2":"0.1079562420"},{"1":"405","2":"-0.6698144690"},{"1":"406","2":"-0.0247110486"},{"1":"407","2":"-0.3055947010"},{"1":"408","2":"0.1375954350"},{"1":"409","2":"0.3150780028"},{"1":"410","2":"-0.8905436779"},{"1":"411","2":"-0.9846236326"},{"1":"412","2":"-0.8903042720"},{"1":"413","2":"0.5484089806"},{"1":"414","2":"0.0750954345"},{"1":"415","2":"-0.5473397239"},{"1":"416","2":"0.2422685949"},{"1":"417","2":"-0.1971176114"},{"1":"418","2":"0.1208374534"},{"1":"419","2":"-0.8530589591"},{"1":"420","2":"0.2081873462"},{"1":"421","2":"-0.8215310834"},{"1":"422","2":"-0.6996630188"},{"1":"423","2":"-0.0728618752"},{"1":"424","2":"-0.4016514603"},{"1":"425","2":"0.0980436711"},{"1":"426","2":"0.0926039056"},{"1":"427","2":"-0.1627142331"},{"1":"428","2":"0.7041111535"},{"1":"429","2":"0.0211100541"},{"1":"430","2":"0.8289079292"},{"1":"431","2":"-0.0013622388"},{"1":"432","2":"0.5666425862"},{"1":"433","2":"0.3837323292"},{"1":"434","2":"0.4205816313"},{"1":"435","2":"-0.4015715511"},{"1":"436","2":"-0.4258884701"},{"1":"437","2":"-0.2873327854"},{"1":"438","2":"0.0152526314"},{"1":"439","2":"0.2150452311"},{"1":"440","2":"0.3764134467"},{"1":"441","2":"0.0308683799"},{"1":"442","2":"0.0164040095"},{"1":"443","2":"-0.3363879120"},{"1":"444","2":"-0.6554810195"},{"1":"445","2":"-0.3598477050"},{"1":"446","2":"-0.1038435831"},{"1":"447","2":"-0.8069886021"},{"1":"448","2":"0.0604618240"},{"1":"449","2":"0.5412274044"},{"1":"450","2":"-1.0612558257"},{"1":"451","2":"-0.4606285533"},{"1":"452","2":"0.8659856844"},{"1":"453","2":"0.9306864032"},{"1":"454","2":"-0.4547526058"},{"1":"455","2":"-0.2482355503"},{"1":"456","2":"0.1281455874"},{"1":"457","2":"0.4891567493"},{"1":"458","2":"0.6760389871"},{"1":"459","2":"0.1586468118"},{"1":"460","2":"0.5924346213"},{"1":"461","2":"0.1913659240"},{"1":"462","2":"0.0741025396"},{"1":"463","2":"0.1869577441"},{"1":"464","2":"0.2252605458"},{"1":"465","2":"-0.0623005162"},{"1":"466","2":"-0.1690578206"},{"1":"467","2":"0.9206248066"},{"1":"468","2":"0.7039994060"},{"1":"469","2":"-0.2823377387"},{"1":"470","2":"0.0528382323"},{"1":"471","2":"0.3347181426"},{"1":"472","2":"0.4415201486"},{"1":"473","2":"0.3817980908"},{"1":"474","2":"0.6338175022"},{"1":"475","2":"-0.4798090650"},{"1":"476","2":"0.8578781756"},{"1":"477","2":"-0.9646208974"},{"1":"478","2":"0.3503125078"},{"1":"479","2":"0.0495019676"},{"1":"480","2":"0.5035510557"},{"1":"481","2":"0.0042848109"},{"1":"482","2":"0.0052195850"},{"1":"483","2":"0.2730317876"},{"1":"484","2":"0.2801357602"},{"1":"485","2":"0.2988705657"},{"1":"486","2":"0.1153331980"},{"1":"487","2":"-1.0455004316"},{"1":"488","2":"0.8140055488"},{"1":"489","2":"-0.3532152486"},{"1":"490","2":"-0.3854903021"},{"1":"491","2":"0.1854140429"},{"1":"492","2":"0.0920787110"},{"1":"493","2":"0.2034564013"},{"1":"494","2":"-0.2400770869"},{"1":"495","2":"-0.8046583500"},{"1":"496","2":"-0.8607343262"},{"1":"497","2":"-0.4690288034"},{"1":"498","2":"-0.0034336450"},{"1":"499","2":"0.3928824135"},{"1":"500","2":"0.3140414314"},{"1":"501","2":"0.0242355194"},{"1":"502","2":"0.8982589392"},{"1":"503","2":"-0.0341252970"},{"1":"504","2":"-1.1996274543"},{"1":"505","2":"-0.1012627820"},{"1":"506","2":"0.1183849227"},{"1":"507","2":"-0.2727816981"},{"1":"508","2":"-0.2431202213"},{"1":"509","2":"0.6721913639"},{"1":"510","2":"-0.0383532926"},{"1":"511","2":"-0.2580846554"},{"1":"512","2":"0.1282465791"},{"1":"513","2":"-0.0718737707"},{"1":"514","2":"0.6194540072"},{"1":"515","2":"-0.1071835537"},{"1":"516","2":"0.0776329211"},{"1":"517","2":"0.7147754536"},{"1":"518","2":"-0.6845261217"},{"1":"519","2":"0.2402110287"},{"1":"520","2":"0.3422635014"},{"1":"521","2":"0.2672436459"},{"1":"522","2":"0.4621576945"},{"1":"523","2":"-0.1215657832"},{"1":"524","2":"0.3881504962"},{"1":"525","2":"-1.3996330859"},{"1":"526","2":"1.1006869475"},{"1":"527","2":"0.4314128841"},{"1":"528","2":"0.0440118659"},{"1":"529","2":"1.2218961290"},{"1":"530","2":"0.1751104631"},{"1":"531","2":"0.5989093797"},{"1":"532","2":"-0.5939368021"},{"1":"533","2":"0.4955615542"},{"1":"534","2":"0.0050548774"},{"1":"535","2":"-0.5221685728"},{"1":"536","2":"0.5550444119"},{"1":"537","2":"0.2126119757"},{"1":"538","2":"-0.3570058302"},{"1":"539","2":"-0.0379035553"},{"1":"540","2":"-0.5126292511"},{"1":"541","2":"-0.1919204571"},{"1":"542","2":"-0.7946401807"},{"1":"543","2":"-0.5674808730"},{"1":"544","2":"-0.0341769426"},{"1":"545","2":"-0.0869004667"},{"1":"546","2":"-0.2761832808"},{"1":"547","2":"0.1988987846"},{"1":"548","2":"-0.6126855001"},{"1":"549","2":"0.0700661031"},{"1":"550","2":"0.0344243095"},{"1":"551","2":"0.0601953352"},{"1":"552","2":"-0.1171982459"},{"1":"553","2":"0.5031346231"},{"1":"554","2":"-0.5092509202"},{"1":"555","2":"-0.6781676225"},{"1":"556","2":"-0.6974475969"},{"1":"557","2":"0.3006477154"},{"1":"558","2":"-0.2751240981"},{"1":"559","2":"0.1646550618"},{"1":"560","2":"-0.5146856217"},{"1":"561","2":"0.7664662660"},{"1":"562","2":"-0.1448586535"},{"1":"563","2":"-0.4260054281"},{"1":"564","2":"0.3401686503"},{"1":"565","2":"-0.3166383875"},{"1":"566","2":"0.0657232804"},{"1":"567","2":"0.3364540188"},{"1":"568","2":"0.3006580043"},{"1":"569","2":"0.1220658408"},{"1":"570","2":"0.2460132523"},{"1":"571","2":"0.2201073490"},{"1":"572","2":"1.3059297762"},{"1":"573","2":"-0.0460034741"},{"1":"574","2":"0.2332501528"},{"1":"575","2":"-0.2173059668"},{"1":"576","2":"0.0854683415"},{"1":"577","2":"-0.3990143570"},{"1":"578","2":"-0.4176239996"},{"1":"579","2":"-0.6515036938"},{"1":"580","2":"0.0767976460"},{"1":"581","2":"-0.0245793098"},{"1":"582","2":"-0.2841717431"},{"1":"583","2":"0.3527097886"},{"1":"584","2":"-0.1731758005"},{"1":"585","2":"1.1299261349"},{"1":"586","2":"-0.4183629169"},{"1":"587","2":"0.2500025686"},{"1":"588","2":"-0.5676051544"},{"1":"589","2":"1.5944497404"},{"1":"590","2":"0.2778887481"},{"1":"591","2":"0.0750093653"},{"1":"592","2":"-0.7970635500"},{"1":"593","2":"0.8687966191"},{"1":"594","2":"0.0713410254"},{"1":"595","2":"-0.6509660831"},{"1":"596","2":"0.4577346272"},{"1":"597","2":"0.6710046264"},{"1":"598","2":"0.2568451119"},{"1":"599","2":"-0.2979288013"},{"1":"600","2":"-0.3416704786"},{"1":"601","2":"0.5667511327"},{"1":"602","2":"0.1318638919"},{"1":"603","2":"-0.1688509325"},{"1":"604","2":"-0.4261687037"},{"1":"605","2":"0.4060822671"},{"1":"606","2":"-0.0138347145"},{"1":"607","2":"-0.5182375123"},{"1":"608","2":"0.0280836623"},{"1":"609","2":"0.3659595828"},{"1":"610","2":"-0.0920343106"},{"1":"611","2":"0.3654017824"},{"1":"612","2":"0.7761748561"},{"1":"613","2":"0.0512957026"},{"1":"614","2":"0.6126084089"},{"1":"615","2":"-1.2349005002"},{"1":"616","2":"0.0297989376"},{"1":"617","2":"0.5083440984"},{"1":"618","2":"0.2025761267"},{"1":"619","2":"0.3827669717"},{"1":"620","2":"0.1272168971"},{"1":"621","2":"0.1313859250"},{"1":"622","2":"-0.5462070682"},{"1":"623","2":"0.3551290675"},{"1":"624","2":"0.0310019573"},{"1":"625","2":"-0.4000376030"},{"1":"626","2":"-0.7916928170"},{"1":"627","2":"0.1128313866"},{"1":"628","2":"-1.1943355629"},{"1":"629","2":"0.4926341529"},{"1":"630","2":"-0.4223178548"},{"1":"631","2":"0.2402159467"},{"1":"632","2":"-0.2582461573"},{"1":"633","2":"-0.0840773642"},{"1":"634","2":"-0.4520462818"},{"1":"635","2":"-0.7034624515"},{"1":"636","2":"0.2537118060"},{"1":"637","2":"-0.5205715723"},{"1":"638","2":"-0.2918059576"},{"1":"639","2":"0.4865903160"},{"1":"640","2":"-0.7637468102"},{"1":"641","2":"0.7512910699"},{"1":"642","2":"-0.6965622481"},{"1":"643","2":"1.1630438364"},{"1":"644","2":"-0.1868705842"},{"1":"645","2":"0.9249851702"},{"1":"646","2":"-0.4648851764"},{"1":"647","2":"-1.0751035440"},{"1":"648","2":"0.4720155715"},{"1":"649","2":"0.4210588496"},{"1":"650","2":"0.7160714742"},{"1":"651","2":"-0.1958769016"},{"1":"652","2":"-0.7922958497"},{"1":"653","2":"-1.0641429838"},{"1":"654","2":"-0.3237658176"},{"1":"655","2":"0.9671506459"},{"1":"656","2":"0.0192937730"},{"1":"657","2":"0.1697004502"},{"1":"658","2":"0.8245136406"},{"1":"659","2":"0.2766495267"},{"1":"660","2":"-0.5702460408"},{"1":"661","2":"-0.0267371460"},{"1":"662","2":"0.3455103522"},{"1":"663","2":"0.0144956313"},{"1":"664","2":"-0.2256465103"},{"1":"665","2":"-0.3281169662"},{"1":"666","2":"0.5054256200"},{"1":"667","2":"-0.0695949511"},{"1":"668","2":"-0.9957726034"},{"1":"669","2":"-0.4974408574"},{"1":"670","2":"-0.4618787012"},{"1":"671","2":"-0.6424231339"},{"1":"672","2":"0.4841429352"},{"1":"673","2":"-0.1056009211"},{"1":"674","2":"0.0541637287"},{"1":"675","2":"0.4218038650"},{"1":"676","2":"0.7206450566"},{"1":"677","2":"-0.0714379610"},{"1":"678","2":"0.1418419761"},{"1":"679","2":"-0.9175881307"},{"1":"680","2":"0.4308382017"},{"1":"681","2":"-0.8826282710"},{"1":"682","2":"0.9499745194"},{"1":"683","2":"0.2156786231"},{"1":"684","2":"-1.0239948642"},{"1":"685","2":"0.0575217062"},{"1":"686","2":"-0.0670434581"},{"1":"687","2":"-1.2346383735"},{"1":"688","2":"0.2034602811"},{"1":"689","2":"0.4332059762"},{"1":"690","2":"-0.2084998855"},{"1":"691","2":"0.0192336248"},{"1":"692","2":"-0.3539581900"},{"1":"693","2":"-0.7048950555"},{"1":"694","2":"-0.0738256365"},{"1":"695","2":"-0.2340310906"},{"1":"696","2":"-0.4964976231"},{"1":"697","2":"0.3304338024"},{"1":"698","2":"-0.8476630442"},{"1":"699","2":"0.4055395086"},{"1":"700","2":"1.0646477438"},{"1":"701","2":"-0.1385415409"},{"1":"702","2":"0.2049458596"},{"1":"703","2":"-0.0136518386"},{"1":"704","2":"-0.9074165020"},{"1":"705","2":"0.1203822669"},{"1":"706","2":"-0.9068243257"},{"1":"707","2":"-0.1489956212"},{"1":"708","2":"1.2570572384"},{"1":"709","2":"0.3314187533"},{"1":"710","2":"-0.8679690665"},{"1":"711","2":"0.5473746950"},{"1":"712","2":"-0.6345918324"},{"1":"713","2":"0.5783902173"},{"1":"714","2":"-0.0006793636"},{"1":"715","2":"0.1765230028"},{"1":"716","2":"-0.0595782460"},{"1":"717","2":"0.2247211054"},{"1":"718","2":"-0.1581540610"},{"1":"719","2":"-0.1168624551"},{"1":"720","2":"0.2151934118"},{"1":"721","2":"0.2221649537"},{"1":"722","2":"0.8174355613"},{"1":"723","2":"-0.1738309846"},{"1":"724","2":"0.0492942366"},{"1":"725","2":"0.7362931040"},{"1":"726","2":"-0.3451720461"},{"1":"727","2":"0.7807247871"},{"1":"728","2":"0.7408486399"},{"1":"729","2":"-0.6186830869"},{"1":"730","2":"-0.2118967048"},{"1":"731","2":"0.6602445343"},{"1":"732","2":"-0.0500527589"},{"1":"733","2":"0.2731457630"},{"1":"734","2":"-0.9231768347"},{"1":"735","2":"0.3020594252"},{"1":"736","2":"0.1981622155"},{"1":"737","2":"-0.4613845839"},{"1":"738","2":"0.2013702449"},{"1":"739","2":"0.2405442306"},{"1":"740","2":"-0.3216353962"},{"1":"741","2":"0.0210136702"},{"1":"742","2":"-0.1445064228"},{"1":"743","2":"-0.1372819065"},{"1":"744","2":"-0.2239354293"},{"1":"745","2":"0.0249432226"},{"1":"746","2":"0.5717049208"},{"1":"747","2":"-0.8698616525"},{"1":"748","2":"0.7088744635"},{"1":"749","2":"0.0321248243"},{"1":"750","2":"0.1360448942"},{"1":"751","2":"-0.0357990838"},{"1":"752","2":"0.5217825786"},{"1":"753","2":"-0.1953273764"},{"1":"754","2":"-0.3952181669"},{"1":"755","2":"-0.2052035973"},{"1":"756","2":"-0.8213166351"},{"1":"757","2":"-0.5294360749"},{"1":"758","2":"0.2859349801"},{"1":"759","2":"0.4414938791"},{"1":"760","2":"-0.2169188794"},{"1":"761","2":"-0.2936558620"},{"1":"762","2":"0.1983003701"},{"1":"763","2":"1.3062267644"},{"1":"764","2":"-0.2952942375"},{"1":"765","2":"-0.1855339739"},{"1":"766","2":"-0.0152169624"},{"1":"767","2":"-0.7335753154"},{"1":"768","2":"0.2227222794"},{"1":"769","2":"0.0815052219"},{"1":"770","2":"0.0497252990"},{"1":"771","2":"-0.7700054946"},{"1":"772","2":"-0.2725907470"},{"1":"773","2":"0.4053100837"},{"1":"774","2":"0.5233098675"},{"1":"775","2":"-0.6441687271"},{"1":"776","2":"-0.5588199356"},{"1":"777","2":"0.1763386438"},{"1":"778","2":"0.0457982438"},{"1":"779","2":"-0.2227222657"},{"1":"780","2":"0.2159808002"},{"1":"781","2":"0.0679599234"},{"1":"782","2":"0.6595912864"},{"1":"783","2":"-0.2561579385"},{"1":"784","2":"0.2518207609"},{"1":"785","2":"0.0938271028"},{"1":"786","2":"0.0790120818"},{"1":"787","2":"-0.0943584991"},{"1":"788","2":"0.4177476573"},{"1":"789","2":"0.2506480720"},{"1":"790","2":"-0.9246268162"},{"1":"791","2":"-0.3494049582"},{"1":"792","2":"-0.4054020859"},{"1":"793","2":"0.1900198752"},{"1":"794","2":"-0.4642222295"},{"1":"795","2":"-0.5502730017"},{"1":"796","2":"0.7646672863"},{"1":"797","2":"-0.6382167570"},{"1":"798","2":"0.0646982912"},{"1":"799","2":"0.1824116409"},{"1":"800","2":"-0.3847068920"},{"1":"801","2":"-0.2657597654"},{"1":"802","2":"-0.0578743374"},{"1":"803","2":"-0.3845676528"},{"1":"804","2":"-0.4464640014"},{"1":"805","2":"0.3566687440"},{"1":"806","2":"0.6359177293"},{"1":"807","2":"0.4166519176"},{"1":"808","2":"-0.0064140954"},{"1":"809","2":"0.2091652449"},{"1":"810","2":"0.1126447540"},{"1":"811","2":"0.0590965601"},{"1":"812","2":"-0.1454431204"},{"1":"813","2":"0.7148743801"},{"1":"814","2":"-1.1578956851"},{"1":"815","2":"0.0031760109"},{"1":"816","2":"0.0117546849"},{"1":"817","2":"-0.1468220534"},{"1":"818","2":"-0.3737010548"},{"1":"819","2":"-0.2678264772"},{"1":"820","2":"0.0359171329"},{"1":"821","2":"-0.0661784250"},{"1":"822","2":"0.4133180998"},{"1":"823","2":"-0.1196136111"},{"1":"824","2":"-0.4500890218"},{"1":"825","2":"-0.6832816496"},{"1":"826","2":"-0.0697738433"},{"1":"827","2":"0.2784089210"},{"1":"828","2":"-0.1156418170"},{"1":"829","2":"-0.3863117351"},{"1":"830","2":"0.7426794750"},{"1":"831","2":"-0.0744001199"},{"1":"832","2":"1.2701395784"},{"1":"833","2":"1.6481463600"},{"1":"834","2":"-0.1907364977"},{"1":"835","2":"-0.6765805115"},{"1":"836","2":"-0.5206072042"},{"1":"837","2":"0.1604431028"},{"1":"838","2":"-0.2721937512"},{"1":"839","2":"-0.0944107398"},{"1":"840","2":"0.5394680923"},{"1":"841","2":"-0.3559739405"},{"1":"842","2":"0.8404265282"},{"1":"843","2":"0.2423855833"},{"1":"844","2":"-0.4864638312"},{"1":"845","2":"-0.1007063716"},{"1":"846","2":"0.0796126033"},{"1":"847","2":"0.9922827207"},{"1":"848","2":"-0.8372199480"},{"1":"849","2":"0.9025143497"},{"1":"850","2":"-0.3840596974"},{"1":"851","2":"0.3330052732"},{"1":"852","2":"-0.2404929683"},{"1":"853","2":"0.0228017880"},{"1":"854","2":"0.6717280815"},{"1":"855","2":"-0.6894324179"},{"1":"856","2":"0.3790731388"},{"1":"857","2":"0.3612092859"},{"1":"858","2":"-0.0209141688"},{"1":"859","2":"-0.4436371540"},{"1":"860","2":"-0.3102431724"},{"1":"861","2":"-0.5860342872"},{"1":"862","2":"-0.3923710612"},{"1":"863","2":"-0.7387534993"},{"1":"864","2":"-0.6345683336"},{"1":"865","2":"-0.0390253467"},{"1":"866","2":"-0.1056642717"},{"1":"867","2":"-0.9996349048"},{"1":"868","2":"-0.3828183848"},{"1":"869","2":"-0.4478331294"},{"1":"870","2":"0.2222957363"},{"1":"871","2":"-0.9226174633"},{"1":"872","2":"0.7350496039"},{"1":"873","2":"0.0050029917"},{"1":"874","2":"-0.3577701061"},{"1":"875","2":"-0.0810706153"},{"1":"876","2":"-0.4894124623"},{"1":"877","2":"0.3532883778"},{"1":"878","2":"-0.6309114036"},{"1":"879","2":"0.0980338566"},{"1":"880","2":"-0.3341343918"},{"1":"881","2":"0.1230539575"},{"1":"882","2":"-0.2374226497"},{"1":"883","2":"0.4006303265"},{"1":"884","2":"0.7271720142"},{"1":"885","2":"-0.6226563719"},{"1":"886","2":"0.0704490770"},{"1":"887","2":"0.6689417563"},{"1":"888","2":"0.0884434435"},{"1":"889","2":"-0.5725138847"},{"1":"890","2":"-0.6323066737"},{"1":"891","2":"0.0614792393"},{"1":"892","2":"-0.0448705501"},{"1":"893","2":"-0.4452185690"},{"1":"894","2":"0.6728085316"},{"1":"895","2":"0.1845959233"},{"1":"896","2":"0.6094503625"},{"1":"897","2":"0.1405095580"},{"1":"898","2":"0.0286810943"},{"1":"899","2":"-0.5083879737"},{"1":"900","2":"0.2917080282"},{"1":"901","2":"0.7741975330"},{"1":"902","2":"-0.0589374164"},{"1":"903","2":"-0.7713520238"},{"1":"904","2":"-0.0659111660"},{"1":"905","2":"-0.8291358437"},{"1":"906","2":"-0.1068516336"},{"1":"907","2":"-0.0008544647"},{"1":"908","2":"-0.7798745742"},{"1":"909","2":"-0.1567817187"},{"1":"910","2":"-0.0458421055"},{"1":"911","2":"0.9613481252"},{"1":"912","2":"-0.0295746229"},{"1":"913","2":"0.2556826223"},{"1":"914","2":"0.0827633631"},{"1":"915","2":"0.1312654249"},{"1":"916","2":"0.6287809097"},{"1":"917","2":"0.1017895537"},{"1":"918","2":"-1.4299704818"},{"1":"919","2":"0.2686193577"},{"1":"920","2":"-0.1870281605"},{"1":"921","2":"-0.8048225766"},{"1":"922","2":"0.0927608079"},{"1":"923","2":"1.0367911765"},{"1":"924","2":"-0.0489760505"},{"1":"925","2":"0.4896128015"},{"1":"926","2":"0.0011489860"},{"1":"927","2":"0.1624780393"},{"1":"928","2":"0.4247493294"},{"1":"929","2":"-0.4783373922"},{"1":"930","2":"-0.4311071583"},{"1":"931","2":"0.6427574432"},{"1":"932","2":"-0.0741338981"},{"1":"933","2":"0.3428752700"},{"1":"934","2":"0.0534182873"},{"1":"935","2":"0.4152940594"},{"1":"936","2":"-0.9274865930"},{"1":"937","2":"0.0185090540"},{"1":"938","2":"0.5300132525"},{"1":"939","2":"0.5265690910"},{"1":"940","2":"0.3756997518"},{"1":"941","2":"0.1197970845"},{"1":"942","2":"-0.1589345558"},{"1":"943","2":"-0.4475552837"},{"1":"944","2":"0.1719228517"},{"1":"945","2":"0.0115143022"},{"1":"946","2":"0.1110573493"},{"1":"947","2":"-0.7696167347"},{"1":"948","2":"0.7564665113"},{"1":"949","2":"0.1115921515"},{"1":"950","2":"0.1992756517"},{"1":"951","2":"0.4296724886"},{"1":"952","2":"-0.1308737702"},{"1":"953","2":"0.5038520409"},{"1":"954","2":"-0.3996217492"},{"1":"955","2":"0.0755067515"},{"1":"956","2":"-0.6193805940"},{"1":"957","2":"0.5458997510"},{"1":"958","2":"0.0832619243"},{"1":"959","2":"-1.2026275134"},{"1":"960","2":"-0.2082382890"},{"1":"961","2":"0.2423658532"},{"1":"962","2":"-0.3534150484"},{"1":"963","2":"0.9980140276"},{"1":"964","2":"-1.4711066528"},{"1":"965","2":"0.2197053598"},{"1":"966","2":"0.0867052695"},{"1":"967","2":"0.4027259598"},{"1":"968","2":"0.3373016540"},{"1":"969","2":"0.5474074562"},{"1":"970","2":"-0.3621867893"},{"1":"971","2":"0.8151176947"},{"1":"972","2":"0.2203761203"},{"1":"973","2":"0.2452045878"},{"1":"974","2":"-0.0599658925"},{"1":"975","2":"0.4067243173"},{"1":"976","2":"-0.5981470896"},{"1":"977","2":"-0.9640306120"},{"1":"978","2":"0.9242517381"},{"1":"979","2":"0.2533093835"},{"1":"980","2":"-0.6419364302"},{"1":"981","2":"-0.6463949041"},{"1":"982","2":"-1.0108162246"},{"1":"983","2":"-0.1477698500"},{"1":"984","2":"0.0080129743"},{"1":"985","2":"-0.1897083601"},{"1":"986","2":"0.5536491359"},{"1":"987","2":"0.4073544884"},{"1":"988","2":"0.0926848213"},{"1":"989","2":"-0.0977259830"},{"1":"990","2":"0.0813297835"},{"1":"991","2":"0.1883796870"},{"1":"992","2":"-0.5639697056"},{"1":"993","2":"-0.5684833077"},{"1":"994","2":"-0.0845116951"},{"1":"995","2":"-0.2367769410"},{"1":"996","2":"-0.5609299846"},{"1":"997","2":"-0.7309392958"},{"1":"998","2":"0.4356661800"},{"1":"999","2":"-0.6263488769"},{"1":"1000","2":"-0.2483740477"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] --- # Prepping the *infer* Pipeline - Before I show you how to do this, let's first save our estimated slope from our *actual* sample - We'll want this later! ```r # save as obs_slope sample_slope <- school_reg_tidy %>% # this is the regression tidied with broom's tidy() filter(term=="str") %>% pull(estimate) # confirm what it is sample_slope ``` ``` ## [1] -2.279808 ``` --- # The *infer* Pipeline: Specify .center[  ] --- # The *infer* Pipeline: Specify .left-code[ ### Specify ```r data %>% specify(y ~ x) ``` ] .right-plot[ .smallest[ - Take our data and pipe it into the `specify()` function, which is essentially a `lm()` function for regression (for our purposes) ```r CASchool %>% specify(testscr ~ str) ``` <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["testscr"],"name":[1],"type":["dbl"],"align":["right"]},{"label":["str"],"name":[2],"type":["dbl"],"align":["right"]}],"data":[{"1":"690.8","2":"17.88991"},{"1":"661.2","2":"21.52466"},{"1":"643.6","2":"18.69723"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] - Note nothing happens yet ] --- # The *infer* Pipeline: Hypothesize .center[  ] --- # The *infer* Pipeline: Hypothesize .left-code[ ### Specify ### Hypothesize .quitesmall[ `%>% hypothesize(null = "independence")` ] ] .right-plot[ .smallest[ - Describe what the null hypothesis is here - In `infer`'s language, we are hypothesizing that `str` and `testscr` are `independent` `\((\beta_1=0)\)`<sup>.magenta[†]</sup> ```r CASchool %>% specify(testscr ~ str) %>% * hypothesize(null = "independence") ``` <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["testscr"],"name":[1],"type":["dbl"],"align":["right"]},{"label":["str"],"name":[2],"type":["dbl"],"align":["right"]}],"data":[{"1":"690.8","2":"17.88991"},{"1":"661.2","2":"21.52466"},{"1":"643.6","2":"18.69723"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] .tiny[<sup>.magenta[†]</sup> See more [here](https://moderndive.netlify.com/9-hypothesis-testing.html) about what other hypotheses you can test with `infer`.] ] --- # The *infer* Pipeline: Generate I .center[  ] --- # The *infer* Pipeline: Generate I .left-code[ ### Specify ### Hypothesize ### Generate .quitesmall[ `%>% generate(reps = n, type = "permute")` ] ] .right-plot[ - Now the magic starts, as we run a number of simulated samples - Set the number of `reps` and set the `type` equal to `"permute"` - we want `permutation` (not `bootstrap`!) because we are simulating a world where `\(\beta_1=0\)` by construction! ```r CASchool %>% specify(testscr ~ str) %>% hypothesize(null = "independence") %>% * generate(reps = 1000, * type = "permute") ``` ] --- # The *infer* Pipeline: Generate II .left-code[ ### Specify ### Hypothesize ### Generate .quitesmall[ `%>% generate(reps = n, type = "permute")` ] ] .right-plot[ .quitesmall[ <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["testscr"],"name":[1],"type":["dbl"],"align":["right"]},{"label":["str"],"name":[2],"type":["dbl"],"align":["right"]},{"label":["replicate"],"name":[3],"type":["int"],"align":["right"]}],"data":[{"1":"693.95","2":"17.88991","3":"1"},{"1":"642.40","2":"21.52466","3":"1"},{"1":"680.45","2":"18.69723","3":"1"},{"1":"672.70","2":"17.35714","3":"1"},{"1":"666.45","2":"18.67133","3":"1"},{"1":"654.20","2":"21.40625","3":"1"},{"1":"671.95","2":"19.50000","3":"1"},{"1":"671.75","2":"20.89412","3":"1"},{"1":"624.55","2":"19.94737","3":"1"},{"1":"699.10","2":"20.80556","3":"1"},{"1":"681.60","2":"21.23809","3":"1"},{"1":"664.95","2":"21.00000","3":"1"},{"1":"629.80","2":"20.60000","3":"1"},{"1":"671.90","2":"20.00822","3":"1"},{"1":"649.50","2":"18.02778","3":"1"},{"1":"682.45","2":"20.25196","3":"1"},{"1":"650.65","2":"16.97787","3":"1"},{"1":"655.55","2":"16.50980","3":"1"},{"1":"661.45","2":"22.70402","3":"1"},{"1":"655.05","2":"19.91111","3":"1"},{"1":"659.35","2":"18.33333","3":"1"},{"1":"624.95","2":"22.61905","3":"1"},{"1":"634.70","2":"19.44828","3":"1"},{"1":"643.05","2":"25.05263","3":"1"},{"1":"627.10","2":"20.67544","3":"1"},{"1":"664.95","2":"18.68235","3":"1"},{"1":"684.35","2":"22.84553","3":"1"},{"1":"639.30","2":"19.26667","3":"1"},{"1":"643.70","2":"19.25000","3":"1"},{"1":"670.00","2":"20.54545","3":"1"},{"1":"666.65","2":"20.60697","3":"1"},{"1":"679.75","2":"21.07268","3":"1"},{"1":"641.55","2":"21.53581","3":"1"},{"1":"647.60","2":"19.90400","3":"1"},{"1":"655.35","2":"21.19407","3":"1"},{"1":"663.85","2":"21.86535","3":"1"},{"1":"669.35","2":"18.32965","3":"1"},{"1":"634.10","2":"16.22857","3":"1"},{"1":"617.35","2":"19.17857","3":"1"},{"1":"663.35","2":"20.27737","3":"1"},{"1":"657.55","2":"22.98614","3":"1"},{"1":"661.90","2":"20.44444","3":"1"},{"1":"677.25","2":"19.82085","3":"1"},{"1":"668.00","2":"23.20522","3":"1"},{"1":"616.30","2":"19.26697","3":"1"},{"1":"691.05","2":"23.30189","3":"1"},{"1":"655.05","2":"21.18829","3":"1"},{"1":"698.20","2":"20.87180","3":"1"},{"1":"662.45","2":"19.01749","3":"1"},{"1":"646.55","2":"21.91938","3":"1"},{"1":"678.80","2":"20.10124","3":"1"},{"1":"623.45","2":"21.47651","3":"1"},{"1":"669.35","2":"20.06579","3":"1"},{"1":"657.40","2":"20.37510","3":"1"},{"1":"656.55","2":"22.44648","3":"1"},{"1":"641.45","2":"22.89524","3":"1"},{"1":"656.50","2":"20.49797","3":"1"},{"1":"631.05","2":"20.00000","3":"1"},{"1":"668.80","2":"22.25658","3":"1"},{"1":"660.75","2":"21.56436","3":"1"},{"1":"628.65","2":"19.47737","3":"1"},{"1":"698.25","2":"17.67002","3":"1"},{"1":"655.75","2":"21.94756","3":"1"},{"1":"645.25","2":"21.78339","3":"1"},{"1":"656.80","2":"19.14000","3":"1"},{"1":"665.90","2":"18.11050","3":"1"},{"1":"637.95","2":"20.68242","3":"1"},{"1":"630.55","2":"22.62361","3":"1"},{"1":"636.10","2":"21.78650","3":"1"},{"1":"645.60","2":"18.58293","3":"1"},{"1":"676.60","2":"21.54545","3":"1"},{"1":"654.10","2":"21.15289","3":"1"},{"1":"628.25","2":"16.63333","3":"1"},{"1":"676.95","2":"21.14438","3":"1"},{"1":"646.35","2":"19.78182","3":"1"},{"1":"661.35","2":"18.98373","3":"1"},{"1":"655.70","2":"17.66767","3":"1"},{"1":"668.60","2":"17.75499","3":"1"},{"1":"652.10","2":"15.27273","3":"1"},{"1":"654.20","2":"14.00000","3":"1"},{"1":"635.60","2":"20.59613","3":"1"},{"1":"657.80","2":"16.31169","3":"1"},{"1":"644.20","2":"21.12796","3":"1"},{"1":"647.60","2":"17.48801","3":"1"},{"1":"650.45","2":"17.88679","3":"1"},{"1":"612.65","2":"19.30676","3":"1"},{"1":"660.10","2":"20.89231","3":"1"},{"1":"655.35","2":"21.28684","3":"1"},{"1":"662.55","2":"20.19560","3":"1"},{"1":"661.60","2":"24.95000","3":"1"},{"1":"618.05","2":"18.13043","3":"1"},{"1":"647.70","2":"20.00000","3":"1"},{"1":"673.05","2":"18.72951","3":"1"},{"1":"675.40","2":"18.25000","3":"1"},{"1":"634.90","2":"18.99257","3":"1"},{"1":"663.45","2":"19.88764","3":"1"},{"1":"687.55","2":"19.37895","3":"1"},{"1":"627.30","2":"20.46259","3":"1"},{"1":"653.10","2":"22.29157","3":"1"},{"1":"646.40","2":"20.70474","3":"1"},{"1":"672.55","2":"19.06005","3":"1"},{"1":"635.95","2":"20.23247","3":"1"},{"1":"695.20","2":"19.69012","3":"1"},{"1":"626.80","2":"20.36254","3":"1"},{"1":"643.70","2":"19.75422","3":"1"},{"1":"652.50","2":"19.37977","3":"1"},{"1":"626.10","2":"22.92351","3":"1"},{"1":"666.15","2":"19.37340","3":"1"},{"1":"636.50","2":"19.15516","3":"1"},{"1":"661.20","2":"21.30000","3":"1"},{"1":"686.05","2":"18.30357","3":"1"},{"1":"641.10","2":"21.07926","3":"1"},{"1":"626.90","2":"18.79121","3":"1"},{"1":"645.75","2":"19.62662","3":"1"},{"1":"676.85","2":"19.59016","3":"1"},{"1":"665.20","2":"20.87187","3":"1"},{"1":"667.15","2":"21.11500","3":"1"},{"1":"678.05","2":"20.08452","3":"1"},{"1":"646.70","2":"19.91049","3":"1"},{"1":"671.25","2":"17.81285","3":"1"},{"1":"643.25","2":"18.13333","3":"1"},{"1":"690.80","2":"19.22221","3":"1"},{"1":"673.90","2":"18.66072","3":"1"},{"1":"662.50","2":"19.60000","3":"1"},{"1":"634.55","2":"19.28384","3":"1"},{"1":"666.85","2":"22.81818","3":"1"},{"1":"682.15","2":"18.80922","3":"1"},{"1":"649.15","2":"21.37363","3":"1"},{"1":"684.95","2":"20.02041","3":"1"},{"1":"651.35","2":"21.49862","3":"1"},{"1":"653.50","2":"15.42857","3":"1"},{"1":"660.20","2":"22.40000","3":"1"},{"1":"663.50","2":"20.12709","3":"1"},{"1":"648.25","2":"19.03798","3":"1"},{"1":"664.45","2":"17.34216","3":"1"},{"1":"657.15","2":"17.01863","3":"1"},{"1":"644.40","2":"20.80000","3":"1"},{"1":"643.50","2":"21.15385","3":"1"},{"1":"632.20","2":"18.45833","3":"1"},{"1":"636.70","2":"19.14082","3":"1"},{"1":"664.00","2":"19.40766","3":"1"},{"1":"612.50","2":"19.56896","3":"1"},{"1":"627.25","2":"21.50120","3":"1"},{"1":"651.85","2":"17.52941","3":"1"},{"1":"646.95","2":"16.43017","3":"1"},{"1":"656.70","2":"19.79654","3":"1"},{"1":"638.70","2":"17.18613","3":"1"},{"1":"657.90","2":"17.61589","3":"1"},{"1":"696.55","2":"20.12537","3":"1"},{"1":"661.60","2":"22.16667","3":"1"},{"1":"647.30","2":"19.96154","3":"1"},{"1":"664.70","2":"19.03945","3":"1"},{"1":"667.60","2":"15.22436","3":"1"},{"1":"648.95","2":"21.14475","3":"1"},{"1":"672.30","2":"19.64390","3":"1"},{"1":"616.30","2":"21.04869","3":"1"},{"1":"640.75","2":"20.17544","3":"1"},{"1":"678.40","2":"21.39130","3":"1"},{"1":"620.50","2":"20.00833","3":"1"},{"1":"653.55","2":"20.29137","3":"1"},{"1":"638.55","2":"17.66667","3":"1"},{"1":"700.30","2":"18.22055","3":"1"},{"1":"670.70","2":"20.27100","3":"1"},{"1":"644.50","2":"20.19895","3":"1"},{"1":"622.05","2":"21.38424","3":"1"},{"1":"666.00","2":"20.97368","3":"1"},{"1":"645.10","2":"20.00000","3":"1"},{"1":"638.35","2":"17.15328","3":"1"},{"1":"650.60","2":"22.34977","3":"1"},{"1":"606.75","2":"22.17007","3":"1"},{"1":"653.80","2":"18.18182","3":"1"},{"1":"667.45","2":"18.95714","3":"1"},{"1":"654.60","2":"19.74533","3":"1"},{"1":"661.85","2":"16.42623","3":"1"},{"1":"631.90","2":"16.62540","3":"1"},{"1":"651.80","2":"16.38177","3":"1"},{"1":"660.95","2":"20.07416","3":"1"},{"1":"618.30","2":"17.99544","3":"1"},{"1":"684.30","2":"19.39130","3":"1"},{"1":"654.85","2":"16.42857","3":"1"},{"1":"635.05","2":"16.72949","3":"1"},{"1":"659.90","2":"24.41345","3":"1"},{"1":"674.25","2":"18.26415","3":"1"},{"1":"650.90","2":"18.95504","3":"1"},{"1":"628.75","2":"21.03896","3":"1"},{"1":"698.45","2":"20.74074","3":"1"},{"1":"647.05","2":"18.10000","3":"1"},{"1":"655.80","2":"19.84615","3":"1"},{"1":"643.40","2":"21.60000","3":"1"},{"1":"642.75","2":"22.44242","3":"1"},{"1":"686.70","2":"23.01438","3":"1"},{"1":"652.35","2":"17.74892","3":"1"},{"1":"664.15","2":"18.28664","3":"1"},{"1":"640.50","2":"19.26544","3":"1"},{"1":"643.20","2":"22.66667","3":"1"},{"1":"660.30","2":"19.29412","3":"1"},{"1":"662.40","2":"17.36364","3":"1"},{"1":"694.25","2":"19.82143","3":"1"},{"1":"642.20","2":"20.43378","3":"1"},{"1":"656.40","2":"21.03721","3":"1"},{"1":"651.20","2":"19.92462","3":"1"},{"1":"643.40","2":"19.00986","3":"1"},{"1":"694.80","2":"23.82222","3":"1"},{"1":"673.25","2":"19.36909","3":"1"},{"1":"665.95","2":"19.82857","3":"1"},{"1":"632.95","2":"15.25885","3":"1"},{"1":"631.85","2":"17.16129","3":"1"},{"1":"675.70","2":"21.81333","3":"1"},{"1":"676.15","2":"19.07471","3":"1"},{"1":"657.00","2":"25.78512","3":"1"},{"1":"646.50","2":"18.21261","3":"1"},{"1":"644.45","2":"18.16606","3":"1"},{"1":"648.70","2":"16.97297","3":"1"},{"1":"644.55","2":"21.50087","3":"1"},{"1":"671.65","2":"20.60000","3":"1"},{"1":"671.70","2":"16.99029","3":"1"},{"1":"645.55","2":"20.77954","3":"1"},{"1":"659.40","2":"15.51247","3":"1"},{"1":"639.50","2":"19.88506","3":"1"},{"1":"630.35","2":"21.39882","3":"1"},{"1":"664.00","2":"20.49751","3":"1"},{"1":"642.20","2":"19.36376","3":"1"},{"1":"673.55","2":"17.65957","3":"1"},{"1":"655.05","2":"21.01796","3":"1"},{"1":"632.00","2":"19.05565","3":"1"},{"1":"634.20","2":"22.53846","3":"1"},{"1":"689.10","2":"21.10787","3":"1"},{"1":"664.15","2":"20.05135","3":"1"},{"1":"706.75","2":"14.20176","3":"1"},{"1":"669.95","2":"18.47687","3":"1"},{"1":"634.40","2":"18.63542","3":"1"},{"1":"636.60","2":"20.94595","3":"1"},{"1":"679.65","2":"21.08548","3":"1"},{"1":"666.15","2":"18.69288","3":"1"},{"1":"643.50","2":"20.86808","3":"1"},{"1":"664.30","2":"19.82558","3":"1"},{"1":"671.90","2":"19.75000","3":"1"},{"1":"639.35","2":"19.50000","3":"1"},{"1":"639.90","2":"18.39080","3":"1"},{"1":"635.75","2":"18.78676","3":"1"},{"1":"645.00","2":"19.77018","3":"1"},{"1":"668.40","2":"19.33333","3":"1"},{"1":"625.85","2":"21.46392","3":"1"},{"1":"662.90","2":"23.08492","3":"1"},{"1":"658.35","2":"21.06299","3":"1"},{"1":"659.05","2":"18.68687","3":"1"},{"1":"663.90","2":"20.77024","3":"1"},{"1":"649.85","2":"19.30556","3":"1"},{"1":"664.40","2":"20.13280","3":"1"},{"1":"658.60","2":"20.66964","3":"1"},{"1":"671.60","2":"22.28155","3":"1"},{"1":"637.65","2":"20.60027","3":"1"},{"1":"668.95","2":"20.82734","3":"1"},{"1":"616.30","2":"19.22492","3":"1"},{"1":"643.60","2":"17.65477","3":"1"},{"1":"652.30","2":"17.00000","3":"1"},{"1":"653.55","2":"16.49773","3":"1"},{"1":"605.55","2":"19.78261","3":"1"},{"1":"676.55","2":"22.30216","3":"1"},{"1":"666.05","2":"17.73077","3":"1"},{"1":"648.35","2":"20.44836","3":"1"},{"1":"669.80","2":"20.37169","3":"1"},{"1":"615.75","2":"20.16479","3":"1"},{"1":"632.85","2":"21.61538","3":"1"},{"1":"662.40","2":"20.56143","3":"1"},{"1":"667.45","2":"19.95551","3":"1"},{"1":"683.40","2":"21.18387","3":"1"},{"1":"681.30","2":"18.81042","3":"1"},{"1":"640.10","2":"20.57838","3":"1"},{"1":"648.20","2":"18.32461","3":"1"},{"1":"654.90","2":"18.82063","3":"1"},{"1":"644.70","2":"20.81633","3":"1"},{"1":"661.95","2":"20.00000","3":"1"},{"1":"644.20","2":"19.68182","3":"1"},{"1":"651.15","2":"19.39018","3":"1"},{"1":"663.85","2":"20.92732","3":"1"},{"1":"665.10","2":"19.94437","3":"1"},{"1":"655.30","2":"20.79109","3":"1"},{"1":"654.85","2":"19.20354","3":"1"},{"1":"672.20","2":"19.02439","3":"1"},{"1":"622.60","2":"17.62058","3":"1"},{"1":"679.40","2":"20.23715","3":"1"},{"1":"646.90","2":"19.29374","3":"1"},{"1":"632.45","2":"18.82998","3":"1"},{"1":"664.75","2":"20.33949","3":"1"},{"1":"683.35","2":"19.22900","3":"1"},{"1":"691.90","2":"17.89130","3":"1"},{"1":"662.75","2":"19.51881","3":"1"},{"1":"666.65","2":"19.08451","3":"1"},{"1":"651.45","2":"19.93548","3":"1"},{"1":"644.95","2":"18.87326","3":"1"},{"1":"662.70","2":"20.14178","3":"1"},{"1":"649.70","2":"23.55637","3":"1"},{"1":"646.00","2":"21.46479","3":"1"},{"1":"650.90","2":"19.19101","3":"1"},{"1":"652.85","2":"20.13080","3":"1"},{"1":"668.65","2":"25.80000","3":"1"},{"1":"669.30","2":"18.77774","3":"1"},{"1":"691.35","2":"19.10982","3":"1"},{"1":"640.90","2":"19.70109","3":"1"},{"1":"704.30","2":"18.61594","3":"1"},{"1":"635.45","2":"20.99721","3":"1"},{"1":"661.90","2":"20.00000","3":"1"},{"1":"662.65","2":"20.98325","3":"1"},{"1":"623.20","2":"21.64262","3":"1"},{"1":"630.55","2":"20.02967","3":"1"},{"1":"633.05","2":"19.81140","3":"1"},{"1":"655.40","2":"18.00000","3":"1"},{"1":"695.30","2":"19.35811","3":"1"},{"1":"637.35","2":"20.17912","3":"1"},{"1":"640.85","2":"21.11986","3":"1"},{"1":"628.55","2":"23.38974","3":"1"},{"1":"681.30","2":"22.18182","3":"1"},{"1":"634.95","2":"19.94283","3":"1"},{"1":"638.30","2":"17.78826","3":"1"},{"1":"631.95","2":"14.70588","3":"1"},{"1":"638.20","2":"19.04077","3":"1"},{"1":"666.10","2":"20.89195","3":"1"},{"1":"672.45","2":"19.83851","3":"1"},{"1":"661.85","2":"19.52191","3":"1"},{"1":"652.30","2":"20.68622","3":"1"},{"1":"652.40","2":"18.18182","3":"1"},{"1":"652.40","2":"18.89224","3":"1"},{"1":"659.15","2":"24.88889","3":"1"},{"1":"654.30","2":"18.58064","3":"1"},{"1":"665.65","2":"18.04000","3":"1"},{"1":"634.00","2":"17.73399","3":"1"},{"1":"652.00","2":"21.45455","3":"1"},{"1":"660.05","2":"19.92343","3":"1"},{"1":"633.90","2":"20.33942","3":"1"},{"1":"635.20","2":"22.54608","3":"1"},{"1":"621.75","2":"21.10344","3":"1"},{"1":"640.15","2":"18.19743","3":"1"},{"1":"644.45","2":"20.10768","3":"1"},{"1":"666.55","2":"19.15984","3":"1"},{"1":"661.85","2":"19.54545","3":"1"},{"1":"669.10","2":"20.88889","3":"1"},{"1":"658.80","2":"18.39150","3":"1"},{"1":"681.90","2":"19.17990","3":"1"},{"1":"671.30","2":"19.39771","3":"1"},{"1":"638.00","2":"21.67827","3":"1"},{"1":"662.55","2":"19.28889","3":"1"},{"1":"659.80","2":"20.34927","3":"1"},{"1":"652.10","2":"20.96416","3":"1"},{"1":"636.90","2":"19.46039","3":"1"},{"1":"625.30","2":"19.28572","3":"1"},{"1":"659.40","2":"20.91979","3":"1"},{"1":"677.95","2":"20.90021","3":"1"},{"1":"645.75","2":"20.59575","3":"1"},{"1":"609.00","2":"19.37500","3":"1"},{"1":"684.80","2":"19.95122","3":"1"},{"1":"616.45","2":"18.84973","3":"1"},{"1":"641.45","2":"18.11787","3":"1"},{"1":"656.80","2":"19.18341","3":"1"},{"1":"623.60","2":"22.00000","3":"1"},{"1":"619.80","2":"21.58416","3":"1"},{"1":"671.60","2":"20.38889","3":"1"},{"1":"673.30","2":"16.29310","3":"1"},{"1":"672.05","2":"18.27778","3":"1"},{"1":"634.05","2":"19.37472","3":"1"},{"1":"648.00","2":"18.90909","3":"1"},{"1":"673.55","2":"16.40693","3":"1"},{"1":"665.35","2":"15.59140","3":"1"},{"1":"653.95","2":"18.70694","3":"1"},{"1":"632.25","2":"18.32985","3":"1"},{"1":"682.65","2":"17.90235","3":"1"},{"1":"655.85","2":"18.91157","3":"1"},{"1":"672.05","2":"20.32497","3":"1"},{"1":"624.15","2":"20.02457","3":"1"},{"1":"666.85","2":"24.00000","3":"1"},{"1":"679.80","2":"17.60784","3":"1"},{"1":"634.15","2":"19.34853","3":"1"},{"1":"653.70","2":"19.67846","3":"1"},{"1":"621.40","2":"18.72861","3":"1"},{"1":"668.90","2":"15.88235","3":"1"},{"1":"637.10","2":"20.05491","3":"1"},{"1":"653.85","2":"17.98825","3":"1"},{"1":"666.70","2":"16.96629","3":"1"},{"1":"658.00","2":"19.23937","3":"1"},{"1":"668.10","2":"19.19586","3":"1"},{"1":"672.35","2":"19.59906","3":"1"},{"1":"637.95","2":"20.54348","3":"1"},{"1":"669.30","2":"18.58848","3":"1"},{"1":"657.75","2":"15.60419","3":"1"},{"1":"669.85","2":"15.29304","3":"1"},{"1":"641.80","2":"17.65537","3":"1"},{"1":"628.40","2":"17.57976","3":"1"},{"1":"639.85","2":"22.33333","3":"1"},{"1":"633.15","2":"18.75000","3":"1"},{"1":"647.25","2":"18.10241","3":"1"},{"1":"633.65","2":"20.25641","3":"1"},{"1":"653.40","2":"18.80207","3":"1"},{"1":"637.00","2":"18.77230","3":"1"},{"1":"656.65","2":"20.40521","3":"1"},{"1":"666.60","2":"18.65079","3":"1"},{"1":"638.30","2":"20.70707","3":"1"},{"1":"667.20","2":"22.00000","3":"1"},{"1":"631.40","2":"17.69978","3":"1"},{"1":"682.55","2":"21.48329","3":"1"},{"1":"679.50","2":"16.70103","3":"1"},{"1":"630.40","2":"19.57567","3":"1"},{"1":"651.40","2":"17.25806","3":"1"},{"1":"651.90","2":"17.37526","3":"1"},{"1":"657.65","2":"17.34931","3":"1"},{"1":"620.30","2":"16.26229","3":"1"},{"1":"636.95","2":"17.70045","3":"1"},{"1":"639.80","2":"20.12881","3":"1"},{"1":"646.20","2":"18.26539","3":"1"},{"1":"680.05","2":"14.54214","3":"1"},{"1":"634.10","2":"19.15261","3":"1"},{"1":"639.25","2":"17.36574","3":"1"},{"1":"655.20","2":"15.13898","3":"1"},{"1":"657.50","2":"17.84266","3":"1"},{"1":"639.75","2":"15.40704","3":"1"},{"1":"645.55","2":"18.86534","3":"1"},{"1":"657.00","2":"16.47413","3":"1"},{"1":"635.60","2":"17.86263","3":"1"},{"1":"649.30","2":"21.88586","3":"1"},{"1":"650.55","2":"20.20000","3":"1"},{"1":"623.10","2":"19.03640","3":"1"},{"1":"641.45","2":"17.88991","3":"2"},{"1":"663.35","2":"21.52466","3":"2"},{"1":"661.85","2":"18.69723","3":"2"},{"1":"669.30","2":"17.35714","3":"2"},{"1":"652.10","2":"18.67133","3":"2"},{"1":"635.20","2":"21.40625","3":"2"},{"1":"682.45","2":"19.50000","3":"2"},{"1":"649.85","2":"20.89412","3":"2"},{"1":"638.00","2":"19.94737","3":"2"},{"1":"669.30","2":"20.80556","3":"2"},{"1":"659.05","2":"21.23809","3":"2"},{"1":"659.35","2":"21.00000","3":"2"},{"1":"664.15","2":"20.60000","3":"2"},{"1":"671.30","2":"20.00822","3":"2"},{"1":"652.30","2":"18.02778","3":"2"},{"1":"666.55","2":"20.25196","3":"2"},{"1":"668.80","2":"16.97787","3":"2"},{"1":"642.20","2":"16.50980","3":"2"},{"1":"652.30","2":"22.70402","3":"2"},{"1":"646.95","2":"19.91111","3":"2"},{"1":"646.50","2":"18.33333","3":"2"},{"1":"653.55","2":"22.61905","3":"2"},{"1":"684.95","2":"19.44828","3":"2"},{"1":"639.50","2":"25.05263","3":"2"},{"1":"681.90","2":"20.67544","3":"2"},{"1":"636.95","2":"18.68235","3":"2"},{"1":"644.45","2":"22.84553","3":"2"},{"1":"682.65","2":"19.26667","3":"2"},{"1":"651.85","2":"19.25000","3":"2"},{"1":"630.55","2":"20.54545","3":"2"},{"1":"655.30","2":"20.60697","3":"2"},{"1":"666.70","2":"21.07268","3":"2"},{"1":"638.70","2":"21.53581","3":"2"},{"1":"665.10","2":"19.90400","3":"2"},{"1":"664.00","2":"21.19407","3":"2"},{"1":"668.10","2":"21.86535","3":"2"},{"1":"634.55","2":"18.32965","3":"2"},{"1":"646.40","2":"16.22857","3":"2"},{"1":"645.75","2":"19.17857","3":"2"},{"1":"644.20","2":"20.27737","3":"2"},{"1":"636.70","2":"22.98614","3":"2"},{"1":"682.55","2":"20.44444","3":"2"},{"1":"666.05","2":"19.82085","3":"2"},{"1":"642.75","2":"23.20522","3":"2"},{"1":"684.30","2":"19.26697","3":"2"},{"1":"628.65","2":"23.30189","3":"2"},{"1":"698.45","2":"21.18829","3":"2"},{"1":"666.85","2":"20.87180","3":"2"},{"1":"648.35","2":"19.01749","3":"2"},{"1":"634.10","2":"21.91938","3":"2"},{"1":"618.30","2":"20.10124","3":"2"},{"1":"644.40","2":"21.47651","3":"2"},{"1":"672.20","2":"20.06579","3":"2"},{"1":"679.75","2":"20.37510","3":"2"},{"1":"668.95","2":"22.44648","3":"2"},{"1":"667.45","2":"22.89524","3":"2"},{"1":"634.05","2":"20.49797","3":"2"},{"1":"666.85","2":"20.00000","3":"2"},{"1":"668.60","2":"22.25658","3":"2"},{"1":"646.00","2":"21.56436","3":"2"},{"1":"681.30","2":"19.47737","3":"2"},{"1":"666.60","2":"17.67002","3":"2"},{"1":"672.05","2":"21.94756","3":"2"},{"1":"626.90","2":"21.78339","3":"2"},{"1":"666.65","2":"19.14000","3":"2"},{"1":"655.40","2":"18.11050","3":"2"},{"1":"660.20","2":"20.68242","3":"2"},{"1":"612.50","2":"22.62361","3":"2"},{"1":"681.30","2":"21.78650","3":"2"},{"1":"663.85","2":"18.58293","3":"2"},{"1":"666.15","2":"21.54545","3":"2"},{"1":"665.35","2":"21.15289","3":"2"},{"1":"639.90","2":"16.63333","3":"2"},{"1":"657.50","2":"21.14438","3":"2"},{"1":"652.10","2":"19.78182","3":"2"},{"1":"643.20","2":"18.98373","3":"2"},{"1":"668.90","2":"17.66767","3":"2"},{"1":"676.60","2":"17.75499","3":"2"},{"1":"638.35","2":"15.27273","3":"2"},{"1":"653.40","2":"14.00000","3":"2"},{"1":"662.55","2":"20.59613","3":"2"},{"1":"678.80","2":"16.31169","3":"2"},{"1":"699.10","2":"21.12796","3":"2"},{"1":"661.85","2":"17.48801","3":"2"},{"1":"639.30","2":"17.88679","3":"2"},{"1":"639.85","2":"19.30676","3":"2"},{"1":"671.75","2":"20.89231","3":"2"},{"1":"621.75","2":"21.28684","3":"2"},{"1":"641.80","2":"20.19560","3":"2"},{"1":"666.45","2":"24.95000","3":"2"},{"1":"669.35","2":"18.13043","3":"2"},{"1":"649.30","2":"20.00000","3":"2"},{"1":"647.60","2":"18.72951","3":"2"},{"1":"655.05","2":"18.25000","3":"2"},{"1":"652.00","2":"18.99257","3":"2"},{"1":"635.60","2":"19.88764","3":"2"},{"1":"630.35","2":"19.37895","3":"2"},{"1":"639.25","2":"20.46259","3":"2"},{"1":"696.55","2":"22.29157","3":"2"},{"1":"643.70","2":"20.70474","3":"2"},{"1":"664.15","2":"19.06005","3":"2"},{"1":"628.55","2":"20.23247","3":"2"},{"1":"665.90","2":"19.69012","3":"2"},{"1":"695.30","2":"20.36254","3":"2"},{"1":"652.40","2":"19.75422","3":"2"},{"1":"653.80","2":"19.37977","3":"2"},{"1":"657.90","2":"22.92351","3":"2"},{"1":"663.90","2":"19.37340","3":"2"},{"1":"662.55","2":"19.15516","3":"2"},{"1":"664.00","2":"21.30000","3":"2"},{"1":"643.05","2":"18.30357","3":"2"},{"1":"643.50","2":"21.07926","3":"2"},{"1":"643.70","2":"18.79121","3":"2"},{"1":"676.55","2":"19.62662","3":"2"},{"1":"644.70","2":"19.59016","3":"2"},{"1":"651.90","2":"20.87187","3":"2"},{"1":"657.40","2":"21.11500","3":"2"},{"1":"648.00","2":"20.08452","3":"2"},{"1":"635.45","2":"19.91049","3":"2"},{"1":"694.80","2":"17.81285","3":"2"},{"1":"664.70","2":"18.13333","3":"2"},{"1":"665.20","2":"19.22221","3":"2"},{"1":"629.80","2":"18.66072","3":"2"},{"1":"632.45","2":"19.60000","3":"2"},{"1":"634.15","2":"19.28384","3":"2"},{"1":"632.25","2":"22.81818","3":"2"},{"1":"628.75","2":"18.80922","3":"2"},{"1":"623.45","2":"21.37363","3":"2"},{"1":"654.90","2":"20.02041","3":"2"},{"1":"623.60","2":"21.49862","3":"2"},{"1":"637.95","2":"15.42857","3":"2"},{"1":"659.40","2":"22.40000","3":"2"},{"1":"679.40","2":"20.12709","3":"2"},{"1":"656.50","2":"19.03798","3":"2"},{"1":"634.40","2":"17.34216","3":"2"},{"1":"646.90","2":"17.01863","3":"2"},{"1":"645.25","2":"20.80000","3":"2"},{"1":"654.20","2":"21.15385","3":"2"},{"1":"632.95","2":"18.45833","3":"2"},{"1":"621.40","2":"19.14082","3":"2"},{"1":"655.80","2":"19.40766","3":"2"},{"1":"647.25","2":"19.56896","3":"2"},{"1":"655.05","2":"21.50120","3":"2"},{"1":"632.00","2":"17.52941","3":"2"},{"1":"637.95","2":"16.43017","3":"2"},{"1":"652.40","2":"19.79654","3":"2"},{"1":"657.75","2":"17.18613","3":"2"},{"1":"668.65","2":"17.61589","3":"2"},{"1":"644.95","2":"20.12537","3":"2"},{"1":"662.50","2":"22.16667","3":"2"},{"1":"657.65","2":"19.96154","3":"2"},{"1":"678.40","2":"19.03945","3":"2"},{"1":"640.75","2":"15.22436","3":"2"},{"1":"615.75","2":"21.14475","3":"2"},{"1":"662.75","2":"19.64390","3":"2"},{"1":"626.10","2":"21.04869","3":"2"},{"1":"635.05","2":"20.17544","3":"2"},{"1":"640.85","2":"21.39130","3":"2"},{"1":"671.65","2":"20.00833","3":"2"},{"1":"670.70","2":"20.29137","3":"2"},{"1":"656.80","2":"17.66667","3":"2"},{"1":"616.30","2":"18.22055","3":"2"},{"1":"639.80","2":"20.27100","3":"2"},{"1":"638.20","2":"20.19895","3":"2"},{"1":"672.35","2":"21.38424","3":"2"},{"1":"674.25","2":"20.97368","3":"2"},{"1":"624.95","2":"20.00000","3":"2"},{"1":"684.80","2":"17.15328","3":"2"},{"1":"643.50","2":"22.34977","3":"2"},{"1":"662.45","2":"22.17007","3":"2"},{"1":"606.75","2":"18.18182","3":"2"},{"1":"616.45","2":"18.95714","3":"2"},{"1":"631.40","2":"19.74533","3":"2"},{"1":"667.60","2":"16.42623","3":"2"},{"1":"656.65","2":"16.62540","3":"2"},{"1":"654.20","2":"16.38177","3":"2"},{"1":"700.30","2":"20.07416","3":"2"},{"1":"659.90","2":"17.99544","3":"2"},{"1":"635.95","2":"19.39130","3":"2"},{"1":"650.60","2":"16.42857","3":"2"},{"1":"654.85","2":"16.72949","3":"2"},{"1":"638.30","2":"24.41345","3":"2"},{"1":"631.95","2":"18.26415","3":"2"},{"1":"656.55","2":"18.95504","3":"2"},{"1":"642.40","2":"21.03896","3":"2"},{"1":"660.95","2":"20.74074","3":"2"},{"1":"665.65","2":"18.10000","3":"2"},{"1":"650.65","2":"19.84615","3":"2"},{"1":"644.20","2":"21.60000","3":"2"},{"1":"691.90","2":"22.44242","3":"2"},{"1":"673.90","2":"23.01438","3":"2"},{"1":"672.70","2":"17.74892","3":"2"},{"1":"668.00","2":"18.28664","3":"2"},{"1":"672.05","2":"19.26544","3":"2"},{"1":"653.10","2":"22.66667","3":"2"},{"1":"659.40","2":"19.29412","3":"2"},{"1":"663.45","2":"17.36364","3":"2"},{"1":"637.10","2":"19.82143","3":"2"},{"1":"605.55","2":"20.43378","3":"2"},{"1":"677.25","2":"21.03721","3":"2"},{"1":"638.55","2":"19.92462","3":"2"},{"1":"617.35","2":"19.00986","3":"2"},{"1":"676.85","2":"23.82222","3":"2"},{"1":"618.05","2":"19.36909","3":"2"},{"1":"636.60","2":"19.82857","3":"2"},{"1":"619.80","2":"15.25885","3":"2"},{"1":"640.10","2":"17.16129","3":"2"},{"1":"658.60","2":"21.81333","3":"2"},{"1":"672.45","2":"19.07471","3":"2"},{"1":"695.20","2":"25.78512","3":"2"},{"1":"609.00","2":"18.21261","3":"2"},{"1":"646.70","2":"18.16606","3":"2"},{"1":"641.10","2":"16.97297","3":"2"},{"1":"662.65","2":"21.50087","3":"2"},{"1":"679.65","2":"20.60000","3":"2"},{"1":"665.95","2":"16.99029","3":"2"},{"1":"651.45","2":"20.77954","3":"2"},{"1":"634.95","2":"15.51247","3":"2"},{"1":"662.70","2":"19.88506","3":"2"},{"1":"622.60","2":"21.39882","3":"2"},{"1":"646.55","2":"20.49751","3":"2"},{"1":"686.70","2":"19.36376","3":"2"},{"1":"646.20","2":"17.65957","3":"2"},{"1":"661.95","2":"21.01796","3":"2"},{"1":"673.05","2":"19.05565","3":"2"},{"1":"671.90","2":"22.53846","3":"2"},{"1":"650.45","2":"21.10787","3":"2"},{"1":"655.55","2":"20.05135","3":"2"},{"1":"644.45","2":"14.20176","3":"2"},{"1":"680.05","2":"18.47687","3":"2"},{"1":"660.10","2":"18.63542","3":"2"},{"1":"672.55","2":"20.94595","3":"2"},{"1":"634.70","2":"21.08548","3":"2"},{"1":"658.35","2":"18.69288","3":"2"},{"1":"627.30","2":"20.86808","3":"2"},{"1":"645.75","2":"19.82558","3":"2"},{"1":"643.25","2":"19.75000","3":"2"},{"1":"647.70","2":"19.50000","3":"2"},{"1":"679.80","2":"18.39080","3":"2"},{"1":"689.10","2":"18.78676","3":"2"},{"1":"625.30","2":"19.77018","3":"2"},{"1":"645.55","2":"19.33333","3":"2"},{"1":"671.95","2":"21.46392","3":"2"},{"1":"656.40","2":"23.08492","3":"2"},{"1":"664.95","2":"21.06299","3":"2"},{"1":"669.85","2":"18.68687","3":"2"},{"1":"636.50","2":"20.77024","3":"2"},{"1":"667.20","2":"19.30556","3":"2"},{"1":"653.95","2":"20.13280","3":"2"},{"1":"616.30","2":"20.66964","3":"2"},{"1":"650.90","2":"22.28155","3":"2"},{"1":"638.30","2":"20.60027","3":"2"},{"1":"651.35","2":"20.82734","3":"2"},{"1":"640.50","2":"19.22492","3":"2"},{"1":"661.90","2":"17.65477","3":"2"},{"1":"630.40","2":"17.00000","3":"2"},{"1":"645.55","2":"16.49773","3":"2"},{"1":"667.15","2":"19.78261","3":"2"},{"1":"639.35","2":"22.30216","3":"2"},{"1":"659.15","2":"17.73077","3":"2"},{"1":"670.00","2":"20.44836","3":"2"},{"1":"651.15","2":"20.37169","3":"2"},{"1":"679.50","2":"20.16479","3":"2"},{"1":"663.50","2":"21.61538","3":"2"},{"1":"661.20","2":"20.56143","3":"2"},{"1":"627.25","2":"19.95551","3":"2"},{"1":"657.55","2":"21.18387","3":"2"},{"1":"691.05","2":"18.81042","3":"2"},{"1":"654.10","2":"20.57838","3":"2"},{"1":"655.75","2":"18.32461","3":"2"},{"1":"666.00","2":"18.82063","3":"2"},{"1":"648.95","2":"20.81633","3":"2"},{"1":"644.55","2":"20.00000","3":"2"},{"1":"645.10","2":"19.68182","3":"2"},{"1":"636.90","2":"19.39018","3":"2"},{"1":"640.90","2":"20.92732","3":"2"},{"1":"641.45","2":"19.94437","3":"2"},{"1":"653.50","2":"20.79109","3":"2"},{"1":"662.40","2":"19.20354","3":"2"},{"1":"661.45","2":"19.02439","3":"2"},{"1":"664.40","2":"17.62058","3":"2"},{"1":"662.90","2":"20.23715","3":"2"},{"1":"643.40","2":"19.29374","3":"2"},{"1":"623.20","2":"18.82998","3":"2"},{"1":"672.30","2":"20.33949","3":"2"},{"1":"630.55","2":"19.22900","3":"2"},{"1":"664.30","2":"17.89130","3":"2"},{"1":"673.30","2":"19.51881","3":"2"},{"1":"634.10","2":"19.08451","3":"2"},{"1":"652.85","2":"19.93548","3":"2"},{"1":"639.75","2":"18.87326","3":"2"},{"1":"647.60","2":"20.14178","3":"2"},{"1":"612.65","2":"23.55637","3":"2"},{"1":"637.35","2":"21.46479","3":"2"},{"1":"675.40","2":"19.19101","3":"2"},{"1":"648.20","2":"20.13080","3":"2"},{"1":"698.20","2":"25.80000","3":"2"},{"1":"666.10","2":"18.77774","3":"2"},{"1":"640.15","2":"19.10982","3":"2"},{"1":"669.95","2":"19.70109","3":"2"},{"1":"636.10","2":"18.61594","3":"2"},{"1":"664.95","2":"20.99721","3":"2"},{"1":"633.15","2":"20.00000","3":"2"},{"1":"620.50","2":"20.98325","3":"2"},{"1":"653.55","2":"21.64262","3":"2"},{"1":"655.70","2":"20.02967","3":"2"},{"1":"678.05","2":"19.81140","3":"2"},{"1":"634.00","2":"18.00000","3":"2"},{"1":"654.30","2":"19.35811","3":"2"},{"1":"631.90","2":"20.17912","3":"2"},{"1":"661.60","2":"21.11986","3":"2"},{"1":"622.05","2":"23.38974","3":"2"},{"1":"684.35","2":"22.18182","3":"2"},{"1":"706.75","2":"19.94283","3":"2"},{"1":"655.85","2":"17.78826","3":"2"},{"1":"643.40","2":"14.70588","3":"2"},{"1":"657.00","2":"19.04077","3":"2"},{"1":"666.15","2":"20.89195","3":"2"},{"1":"649.50","2":"19.83851","3":"2"},{"1":"648.70","2":"19.52191","3":"2"},{"1":"657.00","2":"20.68622","3":"2"},{"1":"637.00","2":"18.18182","3":"2"},{"1":"667.45","2":"18.89224","3":"2"},{"1":"686.05","2":"24.88889","3":"2"},{"1":"658.00","2":"18.58064","3":"2"},{"1":"680.45","2":"18.04000","3":"2"},{"1":"664.75","2":"17.73399","3":"2"},{"1":"645.60","2":"21.45455","3":"2"},{"1":"651.80","2":"19.92343","3":"2"},{"1":"671.60","2":"20.33942","3":"2"},{"1":"634.20","2":"22.54608","3":"2"},{"1":"655.05","2":"21.10344","3":"2"},{"1":"661.60","2":"18.19743","3":"2"},{"1":"676.95","2":"20.10768","3":"2"},{"1":"698.25","2":"19.15984","3":"2"},{"1":"693.95","2":"19.54545","3":"2"},{"1":"623.10","2":"20.88889","3":"2"},{"1":"663.85","2":"18.39150","3":"2"},{"1":"675.70","2":"19.17990","3":"2"},{"1":"632.20","2":"19.39771","3":"2"},{"1":"677.95","2":"21.67827","3":"2"},{"1":"644.50","2":"19.28889","3":"2"},{"1":"654.85","2":"20.34927","3":"2"},{"1":"671.90","2":"20.96416","3":"2"},{"1":"616.30","2":"19.46039","3":"2"},{"1":"669.80","2":"19.28572","3":"2"},{"1":"666.65","2":"20.91979","3":"2"},{"1":"633.90","2":"20.90021","3":"2"},{"1":"651.20","2":"20.59575","3":"2"},{"1":"624.15","2":"19.37500","3":"2"},{"1":"660.75","2":"19.95122","3":"2"},{"1":"653.70","2":"18.84973","3":"2"},{"1":"647.05","2":"18.11787","3":"2"},{"1":"628.25","2":"19.18341","3":"2"},{"1":"655.35","2":"22.00000","3":"2"},{"1":"658.80","2":"21.58416","3":"2"},{"1":"634.90","2":"20.38889","3":"2"},{"1":"646.35","2":"16.29310","3":"2"},{"1":"673.55","2":"18.27778","3":"2"},{"1":"651.40","2":"19.37472","3":"2"},{"1":"647.30","2":"18.90909","3":"2"},{"1":"631.85","2":"16.40693","3":"2"},{"1":"620.30","2":"15.59140","3":"2"},{"1":"642.20","2":"18.70694","3":"2"},{"1":"631.05","2":"18.32985","3":"2"},{"1":"668.40","2":"17.90235","3":"2"},{"1":"654.60","2":"18.91157","3":"2"},{"1":"682.15","2":"20.32497","3":"2"},{"1":"659.80","2":"20.02457","3":"2"},{"1":"649.70","2":"24.00000","3":"2"},{"1":"671.60","2":"17.60784","3":"2"},{"1":"650.90","2":"19.34853","3":"2"},{"1":"649.15","2":"19.67846","3":"2"},{"1":"691.35","2":"18.72861","3":"2"},{"1":"673.25","2":"15.88235","3":"2"},{"1":"624.55","2":"20.05491","3":"2"},{"1":"687.55","2":"17.98825","3":"2"},{"1":"632.85","2":"16.96629","3":"2"},{"1":"633.65","2":"19.23937","3":"2"},{"1":"641.55","2":"19.19586","3":"2"},{"1":"662.40","2":"19.59906","3":"2"},{"1":"664.45","2":"20.54348","3":"2"},{"1":"653.85","2":"18.58848","3":"2"},{"1":"652.50","2":"15.60419","3":"2"},{"1":"681.60","2":"15.29304","3":"2"},{"1":"655.20","2":"17.65537","3":"2"},{"1":"643.60","2":"17.57976","3":"2"},{"1":"650.55","2":"22.33333","3":"2"},{"1":"627.10","2":"18.75000","3":"2"},{"1":"683.35","2":"18.10241","3":"2"},{"1":"645.00","2":"20.25641","3":"2"},{"1":"656.70","2":"18.80207","3":"2"},{"1":"635.60","2":"18.77230","3":"2"},{"1":"648.25","2":"20.40521","3":"2"},{"1":"671.25","2":"18.65079","3":"2"},{"1":"657.15","2":"20.70707","3":"2"},{"1":"661.85","2":"22.00000","3":"2"},{"1":"656.80","2":"17.69978","3":"2"},{"1":"661.90","2":"21.48329","3":"2"},{"1":"671.70","2":"16.70103","3":"2"},{"1":"669.35","2":"19.57567","3":"2"},{"1":"660.05","2":"17.25806","3":"2"},{"1":"625.85","2":"17.37526","3":"2"},{"1":"694.25","2":"17.34931","3":"2"},{"1":"652.35","2":"16.26229","3":"2"},{"1":"673.55","2":"17.70045","3":"2"},{"1":"626.80","2":"20.12881","3":"2"},{"1":"704.30","2":"18.26539","3":"2"},{"1":"660.30","2":"14.54214","3":"2"},{"1":"683.40","2":"19.15261","3":"2"},{"1":"637.65","2":"17.36574","3":"2"},{"1":"635.75","2":"15.13898","3":"2"},{"1":"661.35","2":"17.84266","3":"2"},{"1":"676.15","2":"15.40704","3":"2"},{"1":"690.80","2":"18.86534","3":"2"},{"1":"655.35","2":"16.47413","3":"2"},{"1":"657.80","2":"17.86263","3":"2"},{"1":"633.05","2":"21.88586","3":"2"},{"1":"669.10","2":"20.20000","3":"2"},{"1":"628.40","2":"19.03640","3":"2"},{"1":"651.90","2":"17.88991","3":"3"},{"1":"612.50","2":"21.52466","3":"3"},{"1":"654.20","2":"18.69723","3":"3"},{"1":"672.35","2":"17.35714","3":"3"},{"1":"667.45","2":"18.67133","3":"3"},{"1":"662.70","2":"21.40625","3":"3"},{"1":"650.65","2":"19.50000","3":"3"},{"1":"631.40","2":"20.89412","3":"3"},{"1":"682.15","2":"19.94737","3":"3"},{"1":"657.75","2":"20.80556","3":"3"},{"1":"672.20","2":"21.23809","3":"3"},{"1":"628.40","2":"21.00000","3":"3"},{"1":"652.30","2":"20.60000","3":"3"},{"1":"676.60","2":"20.00822","3":"3"},{"1":"626.80","2":"18.02778","3":"3"},{"1":"666.85","2":"20.25196","3":"3"},{"1":"653.10","2":"16.97787","3":"3"},{"1":"639.35","2":"16.50980","3":"3"},{"1":"622.60","2":"22.70402","3":"3"},{"1":"643.60","2":"19.91111","3":"3"},{"1":"680.45","2":"18.33333","3":"3"},{"1":"648.25","2":"22.61905","3":"3"},{"1":"619.80","2":"19.44828","3":"3"},{"1":"662.90","2":"25.05263","3":"3"},{"1":"655.05","2":"20.67544","3":"3"},{"1":"641.55","2":"18.68235","3":"3"},{"1":"655.20","2":"22.84553","3":"3"},{"1":"630.55","2":"19.26667","3":"3"},{"1":"655.75","2":"19.25000","3":"3"},{"1":"651.40","2":"20.54545","3":"3"},{"1":"646.90","2":"20.60697","3":"3"},{"1":"644.20","2":"21.07268","3":"3"},{"1":"662.45","2":"21.53581","3":"3"},{"1":"642.75","2":"19.90400","3":"3"},{"1":"666.45","2":"21.19407","3":"3"},{"1":"638.30","2":"21.86535","3":"3"},{"1":"654.10","2":"18.32965","3":"3"},{"1":"677.25","2":"16.22857","3":"3"},{"1":"634.90","2":"19.17857","3":"3"},{"1":"676.95","2":"20.27737","3":"3"},{"1":"662.55","2":"22.98614","3":"3"},{"1":"655.80","2":"20.44444","3":"3"},{"1":"659.90","2":"19.82085","3":"3"},{"1":"660.75","2":"23.20522","3":"3"},{"1":"632.45","2":"19.26697","3":"3"},{"1":"640.10","2":"23.30189","3":"3"},{"1":"624.15","2":"21.18829","3":"3"},{"1":"669.30","2":"20.87180","3":"3"},{"1":"679.50","2":"19.01749","3":"3"},{"1":"645.10","2":"21.91938","3":"3"},{"1":"653.85","2":"20.10124","3":"3"},{"1":"652.85","2":"21.47651","3":"3"},{"1":"695.20","2":"20.06579","3":"3"},{"1":"655.85","2":"20.37510","3":"3"},{"1":"664.40","2":"22.44648","3":"3"},{"1":"638.20","2":"22.89524","3":"3"},{"1":"651.15","2":"20.49797","3":"3"},{"1":"654.30","2":"20.00000","3":"3"},{"1":"654.85","2":"22.25658","3":"3"},{"1":"647.60","2":"21.56436","3":"3"},{"1":"637.10","2":"19.47737","3":"3"},{"1":"636.60","2":"17.67002","3":"3"},{"1":"633.15","2":"21.94756","3":"3"},{"1":"684.95","2":"21.78339","3":"3"},{"1":"666.85","2":"19.14000","3":"3"},{"1":"664.00","2":"18.11050","3":"3"},{"1":"648.95","2":"20.68242","3":"3"},{"1":"648.00","2":"22.62361","3":"3"},{"1":"661.45","2":"21.78650","3":"3"},{"1":"656.50","2":"18.58293","3":"3"},{"1":"673.55","2":"21.54545","3":"3"},{"1":"661.95","2":"21.15289","3":"3"},{"1":"638.35","2":"16.63333","3":"3"},{"1":"659.35","2":"21.14438","3":"3"},{"1":"676.85","2":"19.78182","3":"3"},{"1":"642.40","2":"18.98373","3":"3"},{"1":"663.90","2":"17.66767","3":"3"},{"1":"655.55","2":"17.75499","3":"3"},{"1":"647.25","2":"15.27273","3":"3"},{"1":"645.55","2":"14.00000","3":"3"},{"1":"645.55","2":"20.59613","3":"3"},{"1":"650.55","2":"16.31169","3":"3"},{"1":"674.25","2":"21.12796","3":"3"},{"1":"691.05","2":"17.48801","3":"3"},{"1":"636.95","2":"17.88679","3":"3"},{"1":"637.35","2":"19.30676","3":"3"},{"1":"668.90","2":"20.89231","3":"3"},{"1":"675.70","2":"21.28684","3":"3"},{"1":"691.90","2":"20.19560","3":"3"},{"1":"671.25","2":"24.95000","3":"3"},{"1":"657.90","2":"18.13043","3":"3"},{"1":"635.05","2":"20.00000","3":"3"},{"1":"675.40","2":"18.72951","3":"3"},{"1":"662.75","2":"18.25000","3":"3"},{"1":"668.65","2":"18.99257","3":"3"},{"1":"671.30","2":"19.88764","3":"3"},{"1":"632.00","2":"19.37895","3":"3"},{"1":"636.90","2":"20.46259","3":"3"},{"1":"704.30","2":"22.29157","3":"3"},{"1":"669.35","2":"20.70474","3":"3"},{"1":"656.55","2":"19.06005","3":"3"},{"1":"664.70","2":"20.23247","3":"3"},{"1":"647.60","2":"19.69012","3":"3"},{"1":"676.55","2":"20.36254","3":"3"},{"1":"658.00","2":"19.75422","3":"3"},{"1":"671.90","2":"19.37977","3":"3"},{"1":"661.85","2":"22.92351","3":"3"},{"1":"655.35","2":"19.37340","3":"3"},{"1":"681.90","2":"19.15516","3":"3"},{"1":"672.05","2":"21.30000","3":"3"},{"1":"639.90","2":"18.30357","3":"3"},{"1":"632.95","2":"21.07926","3":"3"},{"1":"668.60","2":"18.79121","3":"3"},{"1":"684.35","2":"19.62662","3":"3"},{"1":"681.30","2":"19.59016","3":"3"},{"1":"657.00","2":"20.87187","3":"3"},{"1":"630.35","2":"21.11500","3":"3"},{"1":"647.30","2":"20.08452","3":"3"},{"1":"668.10","2":"19.91049","3":"3"},{"1":"634.05","2":"17.81285","3":"3"},{"1":"646.55","2":"18.13333","3":"3"},{"1":"623.10","2":"19.22221","3":"3"},{"1":"645.00","2":"18.66072","3":"3"},{"1":"671.90","2":"19.60000","3":"3"},{"1":"644.40","2":"19.28384","3":"3"},{"1":"653.95","2":"22.81818","3":"3"},{"1":"666.05","2":"18.80922","3":"3"},{"1":"687.55","2":"21.37363","3":"3"},{"1":"628.75","2":"20.02041","3":"3"},{"1":"657.40","2":"21.49862","3":"3"},{"1":"683.35","2":"15.42857","3":"3"},{"1":"659.40","2":"22.40000","3":"3"},{"1":"666.15","2":"20.12709","3":"3"},{"1":"664.45","2":"19.03798","3":"3"},{"1":"655.35","2":"17.34216","3":"3"},{"1":"653.50","2":"17.01863","3":"3"},{"1":"693.95","2":"20.80000","3":"3"},{"1":"623.20","2":"21.15385","3":"3"},{"1":"616.30","2":"18.45833","3":"3"},{"1":"695.30","2":"19.14082","3":"3"},{"1":"667.45","2":"19.40766","3":"3"},{"1":"651.45","2":"19.56896","3":"3"},{"1":"639.30","2":"21.50120","3":"3"},{"1":"626.90","2":"17.52941","3":"3"},{"1":"634.15","2":"16.43017","3":"3"},{"1":"636.10","2":"19.79654","3":"3"},{"1":"672.45","2":"17.18613","3":"3"},{"1":"667.20","2":"17.61589","3":"3"},{"1":"672.30","2":"20.12537","3":"3"},{"1":"646.50","2":"22.16667","3":"3"},{"1":"691.35","2":"19.96154","3":"3"},{"1":"643.05","2":"19.03945","3":"3"},{"1":"641.45","2":"15.22436","3":"3"},{"1":"663.35","2":"21.14475","3":"3"},{"1":"646.40","2":"19.64390","3":"3"},{"1":"643.20","2":"21.04869","3":"3"},{"1":"633.65","2":"20.17544","3":"3"},{"1":"652.00","2":"21.39130","3":"3"},{"1":"649.15","2":"20.00833","3":"3"},{"1":"671.75","2":"20.29137","3":"3"},{"1":"639.80","2":"17.66667","3":"3"},{"1":"673.90","2":"18.22055","3":"3"},{"1":"621.40","2":"20.27100","3":"3"},{"1":"609.00","2":"20.19895","3":"3"},{"1":"645.75","2":"21.38424","3":"3"},{"1":"652.50","2":"20.97368","3":"3"},{"1":"605.55","2":"20.00000","3":"3"},{"1":"637.65","2":"17.15328","3":"3"},{"1":"645.75","2":"22.34977","3":"3"},{"1":"652.30","2":"22.17007","3":"3"},{"1":"673.30","2":"18.18182","3":"3"},{"1":"632.25","2":"18.95714","3":"3"},{"1":"635.95","2":"19.74533","3":"3"},{"1":"669.95","2":"16.42623","3":"3"},{"1":"654.90","2":"16.62540","3":"3"},{"1":"635.20","2":"16.38177","3":"3"},{"1":"634.40","2":"20.07416","3":"3"},{"1":"632.20","2":"17.99544","3":"3"},{"1":"661.60","2":"19.39130","3":"3"},{"1":"671.70","2":"16.42857","3":"3"},{"1":"647.05","2":"16.72949","3":"3"},{"1":"606.75","2":"24.41345","3":"3"},{"1":"664.15","2":"18.26415","3":"3"},{"1":"669.35","2":"18.95504","3":"3"},{"1":"668.00","2":"21.03896","3":"3"},{"1":"659.15","2":"20.74074","3":"3"},{"1":"616.30","2":"18.10000","3":"3"},{"1":"694.80","2":"19.84615","3":"3"},{"1":"634.95","2":"21.60000","3":"3"},{"1":"620.50","2":"22.44242","3":"3"},{"1":"650.45","2":"23.01438","3":"3"},{"1":"649.70","2":"17.74892","3":"3"},{"1":"631.95","2":"18.28664","3":"3"},{"1":"664.30","2":"19.26544","3":"3"},{"1":"671.65","2":"22.66667","3":"3"},{"1":"664.00","2":"19.29412","3":"3"},{"1":"634.20","2":"17.36364","3":"3"},{"1":"663.85","2":"19.82143","3":"3"},{"1":"657.15","2":"20.43378","3":"3"},{"1":"661.90","2":"21.03721","3":"3"},{"1":"655.05","2":"19.92462","3":"3"},{"1":"627.30","2":"19.00986","3":"3"},{"1":"660.05","2":"23.82222","3":"3"},{"1":"672.55","2":"19.36909","3":"3"},{"1":"620.30","2":"19.82857","3":"3"},{"1":"665.90","2":"15.25885","3":"3"},{"1":"673.05","2":"17.16129","3":"3"},{"1":"676.15","2":"21.81333","3":"3"},{"1":"616.45","2":"19.07471","3":"3"},{"1":"640.85","2":"25.78512","3":"3"},{"1":"658.60","2":"18.21261","3":"3"},{"1":"652.10","2":"18.16606","3":"3"},{"1":"655.05","2":"16.97297","3":"3"},{"1":"655.40","2":"21.50087","3":"3"},{"1":"699.10","2":"20.60000","3":"3"},{"1":"646.00","2":"16.99029","3":"3"},{"1":"656.65","2":"20.77954","3":"3"},{"1":"662.50","2":"15.51247","3":"3"},{"1":"659.40","2":"19.88506","3":"3"},{"1":"681.60","2":"21.39882","3":"3"},{"1":"644.50","2":"20.49751","3":"3"},{"1":"627.10","2":"19.36376","3":"3"},{"1":"625.30","2":"17.65957","3":"3"},{"1":"666.00","2":"21.01796","3":"3"},{"1":"643.50","2":"19.05565","3":"3"},{"1":"666.60","2":"22.53846","3":"3"},{"1":"660.95","2":"21.10787","3":"3"},{"1":"631.05","2":"20.05135","3":"3"},{"1":"635.60","2":"14.20176","3":"3"},{"1":"663.45","2":"18.47687","3":"3"},{"1":"628.65","2":"18.63542","3":"3"},{"1":"636.50","2":"20.94595","3":"3"},{"1":"624.55","2":"21.08548","3":"3"},{"1":"679.40","2":"18.69288","3":"3"},{"1":"656.80","2":"20.86808","3":"3"},{"1":"635.60","2":"19.82558","3":"3"},{"1":"637.95","2":"19.75000","3":"3"},{"1":"672.70","2":"19.50000","3":"3"},{"1":"706.75","2":"18.39080","3":"3"},{"1":"659.80","2":"18.78676","3":"3"},{"1":"679.75","2":"19.77018","3":"3"},{"1":"631.85","2":"19.33333","3":"3"},{"1":"679.65","2":"21.46392","3":"3"},{"1":"638.00","2":"23.08492","3":"3"},{"1":"628.25","2":"21.06299","3":"3"},{"1":"659.05","2":"18.68687","3":"3"},{"1":"642.20","2":"20.77024","3":"3"},{"1":"643.25","2":"19.30556","3":"3"},{"1":"638.70","2":"20.13280","3":"3"},{"1":"618.05","2":"20.66964","3":"3"},{"1":"647.70","2":"22.28155","3":"3"},{"1":"670.00","2":"20.60027","3":"3"},{"1":"655.30","2":"20.82734","3":"3"},{"1":"658.80","2":"19.22492","3":"3"},{"1":"672.05","2":"17.65477","3":"3"},{"1":"669.85","2":"17.00000","3":"3"},{"1":"634.10","2":"16.49773","3":"3"},{"1":"657.55","2":"19.78261","3":"3"},{"1":"626.10","2":"22.30216","3":"3"},{"1":"669.10","2":"17.73077","3":"3"},{"1":"623.45","2":"20.44836","3":"3"},{"1":"646.70","2":"20.37169","3":"3"},{"1":"678.80","2":"20.16479","3":"3"},{"1":"648.70","2":"21.61538","3":"3"},{"1":"700.30","2":"20.56143","3":"3"},{"1":"640.50","2":"19.95551","3":"3"},{"1":"666.10","2":"21.18387","3":"3"},{"1":"696.55","2":"18.81042","3":"3"},{"1":"662.55","2":"20.57838","3":"3"},{"1":"643.40","2":"18.32461","3":"3"},{"1":"615.75","2":"18.82063","3":"3"},{"1":"642.20","2":"20.81633","3":"3"},{"1":"657.65","2":"20.00000","3":"3"},{"1":"652.40","2":"19.68182","3":"3"},{"1":"657.80","2":"19.39018","3":"3"},{"1":"621.75","2":"20.92732","3":"3"},{"1":"652.35","2":"19.94437","3":"3"},{"1":"689.10","2":"20.79109","3":"3"},{"1":"666.15","2":"19.20354","3":"3"},{"1":"648.20","2":"19.02439","3":"3"},{"1":"631.90","2":"17.62058","3":"3"},{"1":"669.80","2":"20.23715","3":"3"},{"1":"643.40","2":"19.29374","3":"3"},{"1":"664.95","2":"18.82998","3":"3"},{"1":"648.35","2":"20.33949","3":"3"},{"1":"653.70","2":"19.22900","3":"3"},{"1":"635.75","2":"17.89130","3":"3"},{"1":"668.80","2":"19.51881","3":"3"},{"1":"652.40","2":"19.08451","3":"3"},{"1":"644.45","2":"19.93548","3":"3"},{"1":"644.70","2":"18.87326","3":"3"},{"1":"671.95","2":"20.14178","3":"3"},{"1":"668.40","2":"23.55637","3":"3"},{"1":"686.70","2":"21.46479","3":"3"},{"1":"635.45","2":"19.19101","3":"3"},{"1":"660.10","2":"20.13080","3":"3"},{"1":"664.75","2":"25.80000","3":"3"},{"1":"679.80","2":"18.77774","3":"3"},{"1":"638.30","2":"19.10982","3":"3"},{"1":"646.35","2":"19.70109","3":"3"},{"1":"684.80","2":"18.61594","3":"3"},{"1":"649.30","2":"20.99721","3":"3"},{"1":"662.65","2":"20.00000","3":"3"},{"1":"683.40","2":"20.98325","3":"3"},{"1":"646.95","2":"21.64262","3":"3"},{"1":"668.95","2":"20.02967","3":"3"},{"1":"650.60","2":"19.81140","3":"3"},{"1":"661.60","2":"18.00000","3":"3"},{"1":"633.90","2":"19.35811","3":"3"},{"1":"664.95","2":"20.17912","3":"3"},{"1":"658.35","2":"21.11986","3":"3"},{"1":"643.70","2":"23.38974","3":"3"},{"1":"665.95","2":"22.18182","3":"3"},{"1":"622.05","2":"19.94283","3":"3"},{"1":"690.80","2":"17.78826","3":"3"},{"1":"641.45","2":"14.70588","3":"3"},{"1":"630.40","2":"19.04077","3":"3"},{"1":"634.55","2":"20.89195","3":"3"},{"1":"660.20","2":"19.83851","3":"3"},{"1":"638.55","2":"19.52191","3":"3"},{"1":"678.40","2":"20.68622","3":"3"},{"1":"665.65","2":"18.18182","3":"3"},{"1":"681.30","2":"18.89224","3":"3"},{"1":"673.25","2":"24.88889","3":"3"},{"1":"653.55","2":"18.58064","3":"3"},{"1":"665.35","2":"18.04000","3":"3"},{"1":"624.95","2":"17.73399","3":"3"},{"1":"671.60","2":"21.45455","3":"3"},{"1":"654.20","2":"19.92343","3":"3"},{"1":"617.35","2":"20.33942","3":"3"},{"1":"645.25","2":"22.54608","3":"3"},{"1":"666.70","2":"21.10344","3":"3"},{"1":"666.65","2":"18.19743","3":"3"},{"1":"644.95","2":"20.10768","3":"3"},{"1":"627.25","2":"19.15984","3":"3"},{"1":"651.85","2":"19.54545","3":"3"},{"1":"698.20","2":"20.88889","3":"3"},{"1":"667.15","2":"18.39150","3":"3"},{"1":"673.55","2":"19.17990","3":"3"},{"1":"677.95","2":"19.39771","3":"3"},{"1":"666.65","2":"21.67827","3":"3"},{"1":"654.60","2":"19.28889","3":"3"},{"1":"652.10","2":"20.34927","3":"3"},{"1":"630.55","2":"20.96416","3":"3"},{"1":"661.85","2":"19.46039","3":"3"},{"1":"654.85","2":"19.28572","3":"3"},{"1":"641.10","2":"20.91979","3":"3"},{"1":"655.70","2":"20.90021","3":"3"},{"1":"645.60","2":"20.59575","3":"3"},{"1":"643.50","2":"19.37500","3":"3"},{"1":"661.35","2":"19.95122","3":"3"},{"1":"665.20","2":"18.84973","3":"3"},{"1":"660.30","2":"18.11787","3":"3"},{"1":"644.55","2":"19.18341","3":"3"},{"1":"682.45","2":"22.00000","3":"3"},{"1":"664.15","2":"21.58416","3":"3"},{"1":"661.85","2":"20.38889","3":"3"},{"1":"682.65","2":"16.29310","3":"3"},{"1":"698.25","2":"18.27778","3":"3"},{"1":"663.85","2":"19.37472","3":"3"},{"1":"640.15","2":"18.90909","3":"3"},{"1":"651.35","2":"16.40693","3":"3"},{"1":"670.70","2":"15.59140","3":"3"},{"1":"633.05","2":"18.70694","3":"3"},{"1":"662.40","2":"18.32985","3":"3"},{"1":"643.70","2":"17.90235","3":"3"},{"1":"649.50","2":"18.91157","3":"3"},{"1":"656.80","2":"20.32497","3":"3"},{"1":"686.05","2":"20.02457","3":"3"},{"1":"680.05","2":"24.00000","3":"3"},{"1":"644.20","2":"17.60784","3":"3"},{"1":"665.10","2":"19.34853","3":"3"},{"1":"632.85","2":"19.67846","3":"3"},{"1":"639.25","2":"18.72861","3":"3"},{"1":"653.40","2":"15.88235","3":"3"},{"1":"694.25","2":"20.05491","3":"3"},{"1":"640.90","2":"17.98825","3":"3"},{"1":"637.95","2":"16.96629","3":"3"},{"1":"661.90","2":"19.23937","3":"3"},{"1":"650.90","2":"19.19586","3":"3"},{"1":"653.80","2":"19.59906","3":"3"},{"1":"625.85","2":"20.54348","3":"3"},{"1":"651.20","2":"18.58848","3":"3"},{"1":"634.00","2":"15.60419","3":"3"},{"1":"682.55","2":"15.29304","3":"3"},{"1":"639.85","2":"17.65537","3":"3"},{"1":"657.50","2":"17.57976","3":"3"},{"1":"649.85","2":"22.33333","3":"3"},{"1":"612.65","2":"18.75000","3":"3"},{"1":"636.70","2":"18.10241","3":"3"},{"1":"623.60","2":"20.25641","3":"3"},{"1":"661.20","2":"18.80207","3":"3"},{"1":"678.05","2":"18.77230","3":"3"},{"1":"637.00","2":"20.40521","3":"3"},{"1":"669.30","2":"18.65079","3":"3"},{"1":"628.55","2":"20.70707","3":"3"},{"1":"666.55","2":"22.00000","3":"3"},{"1":"656.70","2":"17.69978","3":"3"},{"1":"646.20","2":"21.48329","3":"3"},{"1":"684.30","2":"16.70103","3":"3"},{"1":"644.45","2":"19.57567","3":"3"},{"1":"650.90","2":"17.25806","3":"3"},{"1":"634.10","2":"17.37526","3":"3"},{"1":"651.80","2":"17.34931","3":"3"},{"1":"662.40","2":"16.26229","3":"3"},{"1":"639.75","2":"17.70045","3":"3"},{"1":"640.75","2":"20.12881","3":"3"},{"1":"639.50","2":"18.26539","3":"3"},{"1":"656.40","2":"14.54214","3":"3"},{"1":"634.70","2":"19.15261","3":"3"},{"1":"616.30","2":"17.36574","3":"3"},{"1":"667.60","2":"15.13898","3":"3"},{"1":"663.50","2":"17.84266","3":"3"},{"1":"618.30","2":"15.40704","3":"3"},{"1":"653.55","2":"18.86534","3":"3"},{"1":"657.00","2":"16.47413","3":"3"},{"1":"641.80","2":"17.86263","3":"3"},{"1":"629.80","2":"21.88586","3":"3"},{"1":"698.45","2":"20.20000","3":"3"},{"1":"671.60","2":"19.03640","3":"3"},{"1":"625.30","2":"17.88991","3":"4"},{"1":"668.90","2":"21.52466","3":"4"},{"1":"640.10","2":"18.69723","3":"4"},{"1":"657.00","2":"17.35714","3":"4"},{"1":"689.10","2":"18.67133","3":"4"},{"1":"652.40","2":"21.40625","3":"4"},{"1":"683.35","2":"19.50000","3":"4"},{"1":"631.95","2":"20.89412","3":"4"},{"1":"672.35","2":"19.94737","3":"4"},{"1":"661.35","2":"20.80556","3":"4"},{"1":"651.35","2":"21.23809","3":"4"},{"1":"694.80","2":"21.00000","3":"4"},{"1":"654.85","2":"20.60000","3":"4"},{"1":"682.15","2":"20.00822","3":"4"},{"1":"669.10","2":"18.02778","3":"4"},{"1":"694.25","2":"20.25196","3":"4"},{"1":"660.20","2":"16.97787","3":"4"},{"1":"646.35","2":"16.50980","3":"4"},{"1":"634.40","2":"22.70402","3":"4"},{"1":"666.85","2":"19.91111","3":"4"},{"1":"699.10","2":"18.33333","3":"4"},{"1":"645.60","2":"22.61905","3":"4"},{"1":"642.40","2":"19.44828","3":"4"},{"1":"673.25","2":"25.05263","3":"4"},{"1":"657.50","2":"20.67544","3":"4"},{"1":"637.10","2":"18.68235","3":"4"},{"1":"659.15","2":"22.84553","3":"4"},{"1":"657.55","2":"19.26667","3":"4"},{"1":"627.25","2":"19.25000","3":"4"},{"1":"648.70","2":"20.54545","3":"4"},{"1":"651.15","2":"20.60697","3":"4"},{"1":"671.60","2":"21.07268","3":"4"},{"1":"670.70","2":"21.53581","3":"4"},{"1":"619.80","2":"19.90400","3":"4"},{"1":"623.20","2":"21.19407","3":"4"},{"1":"653.50","2":"21.86535","3":"4"},{"1":"658.00","2":"18.32965","3":"4"},{"1":"662.90","2":"16.22857","3":"4"},{"1":"651.85","2":"19.17857","3":"4"},{"1":"625.85","2":"20.27737","3":"4"},{"1":"633.65","2":"22.98614","3":"4"},{"1":"676.60","2":"20.44444","3":"4"},{"1":"681.90","2":"19.82085","3":"4"},{"1":"664.00","2":"23.20522","3":"4"},{"1":"691.05","2":"19.26697","3":"4"},{"1":"657.00","2":"23.30189","3":"4"},{"1":"677.95","2":"21.18829","3":"4"},{"1":"631.90","2":"20.87180","3":"4"},{"1":"683.40","2":"19.01749","3":"4"},{"1":"661.85","2":"21.91938","3":"4"},{"1":"627.10","2":"20.10124","3":"4"},{"1":"655.80","2":"21.47651","3":"4"},{"1":"643.40","2":"20.06579","3":"4"},{"1":"654.85","2":"20.37510","3":"4"},{"1":"635.20","2":"22.44648","3":"4"},{"1":"641.45","2":"22.89524","3":"4"},{"1":"623.60","2":"20.49797","3":"4"},{"1":"645.55","2":"20.00000","3":"4"},{"1":"665.65","2":"22.25658","3":"4"},{"1":"635.60","2":"21.56436","3":"4"},{"1":"616.45","2":"19.47737","3":"4"},{"1":"616.30","2":"17.67002","3":"4"},{"1":"666.60","2":"21.94756","3":"4"},{"1":"675.40","2":"21.78339","3":"4"},{"1":"640.75","2":"19.14000","3":"4"},{"1":"634.00","2":"18.11050","3":"4"},{"1":"652.30","2":"20.68242","3":"4"},{"1":"644.20","2":"22.62361","3":"4"},{"1":"618.05","2":"21.78650","3":"4"},{"1":"665.90","2":"18.58293","3":"4"},{"1":"656.40","2":"21.54545","3":"4"},{"1":"661.45","2":"21.15289","3":"4"},{"1":"667.60","2":"16.63333","3":"4"},{"1":"698.45","2":"21.14438","3":"4"},{"1":"643.50","2":"19.78182","3":"4"},{"1":"663.90","2":"18.98373","3":"4"},{"1":"665.35","2":"17.66767","3":"4"},{"1":"669.30","2":"17.75499","3":"4"},{"1":"646.50","2":"15.27273","3":"4"},{"1":"621.75","2":"14.00000","3":"4"},{"1":"655.35","2":"20.59613","3":"4"},{"1":"635.95","2":"16.31169","3":"4"},{"1":"664.95","2":"21.12796","3":"4"},{"1":"644.40","2":"17.48801","3":"4"},{"1":"669.35","2":"17.88679","3":"4"},{"1":"637.00","2":"19.30676","3":"4"},{"1":"658.35","2":"20.89231","3":"4"},{"1":"638.35","2":"21.28684","3":"4"},{"1":"612.50","2":"20.19560","3":"4"},{"1":"669.80","2":"24.95000","3":"4"},{"1":"666.05","2":"18.13043","3":"4"},{"1":"645.00","2":"20.00000","3":"4"},{"1":"662.40","2":"18.72951","3":"4"},{"1":"647.25","2":"18.25000","3":"4"},{"1":"669.85","2":"18.99257","3":"4"},{"1":"678.40","2":"19.88764","3":"4"},{"1":"634.10","2":"19.37895","3":"4"},{"1":"653.70","2":"20.46259","3":"4"},{"1":"668.10","2":"22.29157","3":"4"},{"1":"620.30","2":"20.70474","3":"4"},{"1":"663.50","2":"19.06005","3":"4"},{"1":"668.95","2":"20.23247","3":"4"},{"1":"639.30","2":"19.69012","3":"4"},{"1":"657.40","2":"20.36254","3":"4"},{"1":"647.70","2":"19.75422","3":"4"},{"1":"648.20","2":"19.37977","3":"4"},{"1":"640.15","2":"22.92351","3":"4"},{"1":"639.80","2":"19.37340","3":"4"},{"1":"666.55","2":"19.15516","3":"4"},{"1":"650.65","2":"21.30000","3":"4"},{"1":"634.15","2":"18.30357","3":"4"},{"1":"655.75","2":"21.07926","3":"4"},{"1":"706.75","2":"18.79121","3":"4"},{"1":"653.10","2":"19.62662","3":"4"},{"1":"663.35","2":"19.59016","3":"4"},{"1":"654.20","2":"20.87187","3":"4"},{"1":"664.45","2":"21.11500","3":"4"},{"1":"653.55","2":"20.08452","3":"4"},{"1":"644.50","2":"19.91049","3":"4"},{"1":"646.95","2":"17.81285","3":"4"},{"1":"636.10","2":"18.13333","3":"4"},{"1":"650.90","2":"19.22221","3":"4"},{"1":"659.40","2":"18.66072","3":"4"},{"1":"684.35","2":"19.60000","3":"4"},{"1":"659.35","2":"19.28384","3":"4"},{"1":"657.80","2":"22.81818","3":"4"},{"1":"636.70","2":"18.80922","3":"4"},{"1":"656.70","2":"21.37363","3":"4"},{"1":"632.20","2":"20.02041","3":"4"},{"1":"671.25","2":"21.49862","3":"4"},{"1":"655.05","2":"15.42857","3":"4"},{"1":"691.35","2":"22.40000","3":"4"},{"1":"667.45","2":"20.12709","3":"4"},{"1":"646.90","2":"19.03798","3":"4"},{"1":"665.10","2":"17.34216","3":"4"},{"1":"663.85","2":"17.01863","3":"4"},{"1":"668.60","2":"20.80000","3":"4"},{"1":"679.65","2":"21.15385","3":"4"},{"1":"628.25","2":"18.45833","3":"4"},{"1":"682.65","2":"19.14082","3":"4"},{"1":"664.15","2":"19.40766","3":"4"},{"1":"626.80","2":"19.56896","3":"4"},{"1":"673.05","2":"21.50120","3":"4"},{"1":"639.75","2":"17.52941","3":"4"},{"1":"672.05","2":"16.43017","3":"4"},{"1":"630.55","2":"19.79654","3":"4"},{"1":"647.05","2":"17.18613","3":"4"},{"1":"646.55","2":"17.61589","3":"4"},{"1":"679.40","2":"20.12537","3":"4"},{"1":"663.45","2":"22.16667","3":"4"},{"1":"637.35","2":"19.96154","3":"4"},{"1":"671.95","2":"19.03945","3":"4"},{"1":"662.55","2":"15.22436","3":"4"},{"1":"624.55","2":"21.14475","3":"4"},{"1":"655.40","2":"19.64390","3":"4"},{"1":"671.70","2":"21.04869","3":"4"},{"1":"654.30","2":"20.17544","3":"4"},{"1":"690.80","2":"21.39130","3":"4"},{"1":"638.00","2":"20.00833","3":"4"},{"1":"661.60","2":"20.29137","3":"4"},{"1":"646.20","2":"17.66667","3":"4"},{"1":"620.50","2":"18.22055","3":"4"},{"1":"668.65","2":"20.27100","3":"4"},{"1":"664.75","2":"20.19895","3":"4"},{"1":"641.80","2":"21.38424","3":"4"},{"1":"661.90","2":"20.97368","3":"4"},{"1":"664.95","2":"20.00000","3":"4"},{"1":"698.20","2":"17.15328","3":"4"},{"1":"679.75","2":"22.34977","3":"4"},{"1":"649.70","2":"22.17007","3":"4"},{"1":"651.80","2":"18.18182","3":"4"},{"1":"664.70","2":"18.95714","3":"4"},{"1":"648.95","2":"19.74533","3":"4"},{"1":"646.40","2":"16.42623","3":"4"},{"1":"658.80","2":"16.62540","3":"4"},{"1":"666.85","2":"16.38177","3":"4"},{"1":"666.15","2":"20.07416","3":"4"},{"1":"656.50","2":"17.99544","3":"4"},{"1":"643.25","2":"19.39130","3":"4"},{"1":"660.30","2":"16.42857","3":"4"},{"1":"679.80","2":"16.72949","3":"4"},{"1":"662.65","2":"24.41345","3":"4"},{"1":"646.70","2":"18.26415","3":"4"},{"1":"628.40","2":"18.95504","3":"4"},{"1":"700.30","2":"21.03896","3":"4"},{"1":"668.40","2":"20.74074","3":"4"},{"1":"655.55","2":"18.10000","3":"4"},{"1":"606.75","2":"19.84615","3":"4"},{"1":"632.85","2":"21.60000","3":"4"},{"1":"673.55","2":"22.44242","3":"4"},{"1":"659.40","2":"23.01438","3":"4"},{"1":"639.25","2":"17.74892","3":"4"},{"1":"704.30","2":"18.28664","3":"4"},{"1":"630.40","2":"19.26544","3":"4"},{"1":"640.50","2":"22.66667","3":"4"},{"1":"632.95","2":"19.29412","3":"4"},{"1":"634.05","2":"17.36364","3":"4"},{"1":"656.55","2":"19.82143","3":"4"},{"1":"652.10","2":"20.43378","3":"4"},{"1":"635.05","2":"21.03721","3":"4"},{"1":"666.15","2":"19.92462","3":"4"},{"1":"661.95","2":"19.00986","3":"4"},{"1":"634.90","2":"23.82222","3":"4"},{"1":"665.20","2":"19.36909","3":"4"},{"1":"686.05","2":"19.82857","3":"4"},{"1":"695.20","2":"15.25885","3":"4"},{"1":"653.55","2":"17.16129","3":"4"},{"1":"652.30","2":"21.81333","3":"4"},{"1":"644.95","2":"19.07471","3":"4"},{"1":"622.60","2":"25.78512","3":"4"},{"1":"647.30","2":"18.21261","3":"4"},{"1":"639.35","2":"18.16606","3":"4"},{"1":"634.70","2":"16.97297","3":"4"},{"1":"645.55","2":"21.50087","3":"4"},{"1":"642.75","2":"20.60000","3":"4"},{"1":"668.80","2":"16.99029","3":"4"},{"1":"629.80","2":"20.77954","3":"4"},{"1":"693.95","2":"15.51247","3":"4"},{"1":"639.85","2":"19.88506","3":"4"},{"1":"632.00","2":"21.39882","3":"4"},{"1":"672.20","2":"20.49751","3":"4"},{"1":"667.15","2":"19.36376","3":"4"},{"1":"665.95","2":"17.65957","3":"4"},{"1":"662.45","2":"21.01796","3":"4"},{"1":"605.55","2":"19.05565","3":"4"},{"1":"634.55","2":"22.53846","3":"4"},{"1":"645.75","2":"21.10787","3":"4"},{"1":"661.90","2":"20.05135","3":"4"},{"1":"653.40","2":"14.20176","3":"4"},{"1":"662.70","2":"18.47687","3":"4"},{"1":"630.55","2":"18.63542","3":"4"},{"1":"653.80","2":"20.94595","3":"4"},{"1":"658.60","2":"21.08548","3":"4"},{"1":"680.05","2":"18.69288","3":"4"},{"1":"678.80","2":"20.86808","3":"4"},{"1":"660.05","2":"19.82558","3":"4"},{"1":"664.00","2":"19.75000","3":"4"},{"1":"628.55","2":"19.50000","3":"4"},{"1":"655.05","2":"18.39080","3":"4"},{"1":"621.40","2":"18.78676","3":"4"},{"1":"649.50","2":"19.77018","3":"4"},{"1":"618.30","2":"19.33333","3":"4"},{"1":"640.90","2":"21.46392","3":"4"},{"1":"664.15","2":"23.08492","3":"4"},{"1":"672.55","2":"21.06299","3":"4"},{"1":"676.85","2":"18.68687","3":"4"},{"1":"634.20","2":"20.77024","3":"4"},{"1":"644.70","2":"19.30556","3":"4"},{"1":"670.00","2":"20.13280","3":"4"},{"1":"649.30","2":"20.66964","3":"4"},{"1":"652.00","2":"22.28155","3":"4"},{"1":"645.10","2":"20.60027","3":"4"},{"1":"657.75","2":"20.82734","3":"4"},{"1":"668.00","2":"19.22492","3":"4"},{"1":"662.55","2":"17.65477","3":"4"},{"1":"671.60","2":"17.00000","3":"4"},{"1":"655.05","2":"16.49773","3":"4"},{"1":"645.75","2":"19.78261","3":"4"},{"1":"644.45","2":"22.30216","3":"4"},{"1":"641.55","2":"17.73077","3":"4"},{"1":"657.90","2":"20.44836","3":"4"},{"1":"659.80","2":"20.37169","3":"4"},{"1":"672.45","2":"20.16479","3":"4"},{"1":"657.65","2":"21.61538","3":"4"},{"1":"632.25","2":"20.56143","3":"4"},{"1":"638.30","2":"19.95551","3":"4"},{"1":"660.95","2":"21.18387","3":"4"},{"1":"652.50","2":"18.81042","3":"4"},{"1":"633.15","2":"20.57838","3":"4"},{"1":"666.70","2":"18.32461","3":"4"},{"1":"661.85","2":"18.82063","3":"4"},{"1":"642.20","2":"20.81633","3":"4"},{"1":"643.70","2":"20.00000","3":"4"},{"1":"672.05","2":"19.68182","3":"4"},{"1":"672.70","2":"19.39018","3":"4"},{"1":"673.90","2":"20.92732","3":"4"},{"1":"636.60","2":"19.94437","3":"4"},{"1":"684.30","2":"20.79109","3":"4"},{"1":"696.55","2":"19.20354","3":"4"},{"1":"691.90","2":"19.02439","3":"4"},{"1":"635.75","2":"17.62058","3":"4"},{"1":"647.60","2":"20.23715","3":"4"},{"1":"612.65","2":"19.29374","3":"4"},{"1":"652.10","2":"18.82998","3":"4"},{"1":"649.15","2":"20.33949","3":"4"},{"1":"648.25","2":"19.22900","3":"4"},{"1":"626.90","2":"17.89130","3":"4"},{"1":"639.90","2":"19.51881","3":"4"},{"1":"659.05","2":"19.08451","3":"4"},{"1":"656.65","2":"19.93548","3":"4"},{"1":"671.90","2":"18.87326","3":"4"},{"1":"650.55","2":"20.14178","3":"4"},{"1":"661.60","2":"23.55637","3":"4"},{"1":"669.95","2":"21.46479","3":"4"},{"1":"652.40","2":"19.19101","3":"4"},{"1":"651.90","2":"20.13080","3":"4"},{"1":"609.00","2":"25.80000","3":"4"},{"1":"673.55","2":"18.77774","3":"4"},{"1":"656.80","2":"19.10982","3":"4"},{"1":"636.95","2":"19.70109","3":"4"},{"1":"638.55","2":"18.61594","3":"4"},{"1":"680.45","2":"20.99721","3":"4"},{"1":"654.10","2":"20.00000","3":"4"},{"1":"657.15","2":"20.98325","3":"4"},{"1":"664.40","2":"21.64262","3":"4"},{"1":"652.35","2":"20.02967","3":"4"},{"1":"671.75","2":"19.81140","3":"4"},{"1":"655.20","2":"18.00000","3":"4"},{"1":"617.35","2":"19.35811","3":"4"},{"1":"633.05","2":"20.17912","3":"4"},{"1":"646.00","2":"21.11986","3":"4"},{"1":"661.20","2":"23.38974","3":"4"},{"1":"648.00","2":"22.18182","3":"4"},{"1":"638.70","2":"19.94283","3":"4"},{"1":"640.85","2":"17.78826","3":"4"},{"1":"673.30","2":"14.70588","3":"4"},{"1":"638.30","2":"19.04077","3":"4"},{"1":"637.65","2":"20.89195","3":"4"},{"1":"659.90","2":"19.83851","3":"4"},{"1":"669.30","2":"19.52191","3":"4"},{"1":"643.05","2":"20.68622","3":"4"},{"1":"655.85","2":"18.18182","3":"4"},{"1":"655.35","2":"18.89224","3":"4"},{"1":"650.90","2":"24.88889","3":"4"},{"1":"636.90","2":"18.58064","3":"4"},{"1":"648.35","2":"18.04000","3":"4"},{"1":"675.70","2":"17.73399","3":"4"},{"1":"654.60","2":"21.45455","3":"4"},{"1":"645.25","2":"19.92343","3":"4"},{"1":"661.85","2":"20.33942","3":"4"},{"1":"662.40","2":"22.54608","3":"4"},{"1":"698.25","2":"21.10344","3":"4"},{"1":"667.20","2":"18.19743","3":"4"},{"1":"631.05","2":"20.10768","3":"4"},{"1":"669.35","2":"19.15984","3":"4"},{"1":"681.30","2":"19.54545","3":"4"},{"1":"662.75","2":"20.88889","3":"4"},{"1":"650.60","2":"18.39150","3":"4"},{"1":"643.40","2":"19.17990","3":"4"},{"1":"679.50","2":"19.39771","3":"4"},{"1":"623.10","2":"21.67827","3":"4"},{"1":"667.45","2":"19.28889","3":"4"},{"1":"636.50","2":"20.34927","3":"4"},{"1":"660.10","2":"20.96416","3":"4"},{"1":"676.15","2":"19.46039","3":"4"},{"1":"695.30","2":"19.28572","3":"4"},{"1":"666.00","2":"20.91979","3":"4"},{"1":"651.40","2":"20.90021","3":"4"},{"1":"647.60","2":"20.59575","3":"4"},{"1":"652.85","2":"19.37500","3":"4"},{"1":"664.30","2":"19.95122","3":"4"},{"1":"654.20","2":"18.84973","3":"4"},{"1":"642.20","2":"18.11787","3":"4"},{"1":"639.50","2":"19.18341","3":"4"},{"1":"634.10","2":"22.00000","3":"4"},{"1":"631.85","2":"21.58416","3":"4"},{"1":"643.70","2":"20.38889","3":"4"},{"1":"637.95","2":"16.29310","3":"4"},{"1":"633.90","2":"18.27778","3":"4"},{"1":"637.95","2":"19.37472","3":"4"},{"1":"676.55","2":"18.90909","3":"4"},{"1":"676.95","2":"16.40693","3":"4"},{"1":"634.95","2":"15.59140","3":"4"},{"1":"626.10","2":"18.70694","3":"4"},{"1":"666.65","2":"18.32985","3":"4"},{"1":"624.95","2":"17.90235","3":"4"},{"1":"656.80","2":"18.91157","3":"4"},{"1":"663.85","2":"20.32497","3":"4"},{"1":"651.45","2":"20.02457","3":"4"},{"1":"616.30","2":"24.00000","3":"4"},{"1":"649.85","2":"17.60784","3":"4"},{"1":"641.45","2":"19.34853","3":"4"},{"1":"671.65","2":"19.67846","3":"4"},{"1":"643.60","2":"18.72861","3":"4"},{"1":"616.30","2":"15.88235","3":"4"},{"1":"672.30","2":"20.05491","3":"4"},{"1":"644.45","2":"17.98825","3":"4"},{"1":"687.55","2":"16.96629","3":"4"},{"1":"615.75","2":"19.23937","3":"4"},{"1":"655.30","2":"19.19586","3":"4"},{"1":"631.40","2":"19.59906","3":"4"},{"1":"632.45","2":"20.54348","3":"4"},{"1":"635.60","2":"18.58848","3":"4"},{"1":"643.50","2":"15.60419","3":"4"},{"1":"653.85","2":"15.29304","3":"4"},{"1":"638.20","2":"17.65537","3":"4"},{"1":"678.05","2":"17.57976","3":"4"},{"1":"644.55","2":"22.33333","3":"4"},{"1":"686.70","2":"18.75000","3":"4"},{"1":"627.30","2":"18.10241","3":"4"},{"1":"653.95","2":"20.25641","3":"4"},{"1":"681.60","2":"18.80207","3":"4"},{"1":"660.75","2":"18.77230","3":"4"},{"1":"682.45","2":"20.40521","3":"4"},{"1":"674.25","2":"18.65079","3":"4"},{"1":"671.90","2":"20.70707","3":"4"},{"1":"622.05","2":"22.00000","3":"4"},{"1":"630.35","2":"17.69978","3":"4"},{"1":"628.75","2":"21.48329","3":"4"},{"1":"624.15","2":"16.70103","3":"4"},{"1":"644.20","2":"19.57567","3":"4"},{"1":"671.30","2":"17.25806","3":"4"},{"1":"623.45","2":"17.37526","3":"4"},{"1":"666.10","2":"17.34931","3":"4"},{"1":"635.45","2":"16.26229","3":"4"},{"1":"681.30","2":"17.70045","3":"4"},{"1":"682.55","2":"20.12881","3":"4"},{"1":"684.80","2":"18.26539","3":"4"},{"1":"677.25","2":"14.54214","3":"4"},{"1":"666.65","2":"19.15261","3":"4"},{"1":"662.50","2":"17.36574","3":"4"},{"1":"654.90","2":"15.13898","3":"4"},{"1":"666.45","2":"17.84266","3":"4"},{"1":"650.45","2":"15.40704","3":"4"},{"1":"641.10","2":"18.86534","3":"4"},{"1":"628.65","2":"16.47413","3":"4"},{"1":"655.70","2":"17.86263","3":"4"},{"1":"651.20","2":"21.88586","3":"4"},{"1":"684.95","2":"20.20000","3":"4"},{"1":"643.20","2":"19.03640","3":"4"},{"1":"671.95","2":"17.88991","3":"5"},{"1":"681.90","2":"21.52466","3":"5"},{"1":"643.60","2":"18.69723","3":"5"},{"1":"642.40","2":"17.35714","3":"5"},{"1":"632.85","2":"18.67133","3":"5"},{"1":"635.75","2":"21.40625","3":"5"},{"1":"631.85","2":"19.50000","3":"5"},{"1":"648.20","2":"20.89412","3":"5"},{"1":"647.60","2":"19.94737","3":"5"},{"1":"672.20","2":"20.80556","3":"5"},{"1":"623.20","2":"21.23809","3":"5"},{"1":"639.30","2":"21.00000","3":"5"},{"1":"655.30","2":"20.60000","3":"5"},{"1":"668.40","2":"20.00822","3":"5"},{"1":"631.05","2":"18.02778","3":"5"},{"1":"649.85","2":"20.25196","3":"5"},{"1":"653.50","2":"16.97787","3":"5"},{"1":"650.60","2":"16.50980","3":"5"},{"1":"643.40","2":"22.70402","3":"5"},{"1":"612.65","2":"19.91111","3":"5"},{"1":"661.85","2":"18.33333","3":"5"},{"1":"668.65","2":"22.61905","3":"5"},{"1":"661.90","2":"19.44828","3":"5"},{"1":"643.05","2":"25.05263","3":"5"},{"1":"656.65","2":"20.67544","3":"5"},{"1":"645.75","2":"18.68235","3":"5"},{"1":"667.45","2":"22.84553","3":"5"},{"1":"635.45","2":"19.26667","3":"5"},{"1":"670.00","2":"19.25000","3":"5"},{"1":"660.95","2":"20.54545","3":"5"},{"1":"684.35","2":"20.60697","3":"5"},{"1":"667.60","2":"21.07268","3":"5"},{"1":"644.55","2":"21.53581","3":"5"},{"1":"638.70","2":"19.90400","3":"5"},{"1":"654.85","2":"21.19407","3":"5"},{"1":"637.95","2":"21.86535","3":"5"},{"1":"652.30","2":"18.32965","3":"5"},{"1":"633.05","2":"16.22857","3":"5"},{"1":"642.20","2":"19.17857","3":"5"},{"1":"672.05","2":"20.27737","3":"5"},{"1":"634.70","2":"22.98614","3":"5"},{"1":"624.55","2":"20.44444","3":"5"},{"1":"641.80","2":"19.82085","3":"5"},{"1":"660.20","2":"23.20522","3":"5"},{"1":"646.00","2":"19.26697","3":"5"},{"1":"652.50","2":"23.30189","3":"5"},{"1":"666.45","2":"21.18829","3":"5"},{"1":"682.65","2":"20.87180","3":"5"},{"1":"664.15","2":"19.01749","3":"5"},{"1":"664.45","2":"21.91938","3":"5"},{"1":"632.45","2":"20.10124","3":"5"},{"1":"643.20","2":"21.47651","3":"5"},{"1":"691.90","2":"20.06579","3":"5"},{"1":"675.40","2":"20.37510","3":"5"},{"1":"672.70","2":"22.44648","3":"5"},{"1":"679.75","2":"22.89524","3":"5"},{"1":"679.80","2":"20.49797","3":"5"},{"1":"668.90","2":"20.00000","3":"5"},{"1":"669.10","2":"22.25658","3":"5"},{"1":"652.30","2":"21.56436","3":"5"},{"1":"663.85","2":"19.47737","3":"5"},{"1":"648.35","2":"17.67002","3":"5"},{"1":"645.10","2":"21.94756","3":"5"},{"1":"664.95","2":"21.78339","3":"5"},{"1":"652.40","2":"19.14000","3":"5"},{"1":"653.85","2":"18.11050","3":"5"},{"1":"678.80","2":"20.68242","3":"5"},{"1":"627.25","2":"22.62361","3":"5"},{"1":"649.30","2":"21.78650","3":"5"},{"1":"673.55","2":"18.58293","3":"5"},{"1":"627.30","2":"21.54545","3":"5"},{"1":"678.40","2":"21.15289","3":"5"},{"1":"658.60","2":"16.63333","3":"5"},{"1":"634.15","2":"21.14438","3":"5"},{"1":"638.35","2":"19.78182","3":"5"},{"1":"676.60","2":"18.98373","3":"5"},{"1":"634.10","2":"17.66767","3":"5"},{"1":"641.10","2":"17.75499","3":"5"},{"1":"680.05","2":"15.27273","3":"5"},{"1":"648.25","2":"14.00000","3":"5"},{"1":"657.75","2":"20.59613","3":"5"},{"1":"681.30","2":"16.31169","3":"5"},{"1":"659.40","2":"21.12796","3":"5"},{"1":"659.80","2":"17.48801","3":"5"},{"1":"626.90","2":"17.88679","3":"5"},{"1":"698.20","2":"19.30676","3":"5"},{"1":"643.70","2":"20.89231","3":"5"},{"1":"661.90","2":"21.28684","3":"5"},{"1":"656.80","2":"20.19560","3":"5"},{"1":"639.25","2":"24.95000","3":"5"},{"1":"679.65","2":"18.13043","3":"5"},{"1":"628.25","2":"20.00000","3":"5"},{"1":"667.45","2":"18.72951","3":"5"},{"1":"642.20","2":"18.25000","3":"5"},{"1":"634.95","2":"18.99257","3":"5"},{"1":"704.30","2":"19.88764","3":"5"},{"1":"630.55","2":"19.37895","3":"5"},{"1":"694.25","2":"20.46259","3":"5"},{"1":"666.05","2":"22.29157","3":"5"},{"1":"678.05","2":"20.70474","3":"5"},{"1":"659.35","2":"19.06005","3":"5"},{"1":"664.30","2":"20.23247","3":"5"},{"1":"634.10","2":"19.69012","3":"5"},{"1":"665.95","2":"20.36254","3":"5"},{"1":"654.20","2":"19.75422","3":"5"},{"1":"655.05","2":"19.37977","3":"5"},{"1":"659.40","2":"22.92351","3":"5"},{"1":"654.60","2":"19.37340","3":"5"},{"1":"645.60","2":"19.15516","3":"5"},{"1":"646.95","2":"21.30000","3":"5"},{"1":"662.65","2":"18.30357","3":"5"},{"1":"622.05","2":"21.07926","3":"5"},{"1":"636.50","2":"18.79121","3":"5"},{"1":"632.25","2":"19.62662","3":"5"},{"1":"706.75","2":"19.59016","3":"5"},{"1":"623.45","2":"20.87187","3":"5"},{"1":"620.30","2":"21.11500","3":"5"},{"1":"646.90","2":"20.08452","3":"5"},{"1":"673.05","2":"19.91049","3":"5"},{"1":"672.05","2":"17.81285","3":"5"},{"1":"630.55","2":"18.13333","3":"5"},{"1":"644.40","2":"19.22221","3":"5"},{"1":"665.20","2":"18.66072","3":"5"},{"1":"637.65","2":"19.60000","3":"5"},{"1":"696.55","2":"19.28384","3":"5"},{"1":"645.00","2":"22.81818","3":"5"},{"1":"633.90","2":"18.80922","3":"5"},{"1":"684.80","2":"21.37363","3":"5"},{"1":"638.30","2":"20.02041","3":"5"},{"1":"676.15","2":"21.49862","3":"5"},{"1":"677.95","2":"15.42857","3":"5"},{"1":"656.55","2":"22.40000","3":"5"},{"1":"682.45","2":"20.12709","3":"5"},{"1":"660.05","2":"19.03798","3":"5"},{"1":"618.05","2":"17.34216","3":"5"},{"1":"636.60","2":"17.01863","3":"5"},{"1":"656.80","2":"20.80000","3":"5"},{"1":"663.85","2":"21.15385","3":"5"},{"1":"676.95","2":"18.45833","3":"5"},{"1":"665.10","2":"19.14082","3":"5"},{"1":"675.70","2":"19.40766","3":"5"},{"1":"634.40","2":"19.56896","3":"5"},{"1":"698.45","2":"21.50120","3":"5"},{"1":"657.90","2":"17.52941","3":"5"},{"1":"656.40","2":"16.43017","3":"5"},{"1":"658.80","2":"19.79654","3":"5"},{"1":"636.70","2":"17.18613","3":"5"},{"1":"661.85","2":"17.61589","3":"5"},{"1":"655.70","2":"20.12537","3":"5"},{"1":"653.80","2":"22.16667","3":"5"},{"1":"641.45","2":"19.96154","3":"5"},{"1":"670.70","2":"19.03945","3":"5"},{"1":"640.15","2":"15.22436","3":"5"},{"1":"664.75","2":"21.14475","3":"5"},{"1":"623.10","2":"19.64390","3":"5"},{"1":"649.70","2":"21.04869","3":"5"},{"1":"679.40","2":"20.17544","3":"5"},{"1":"654.20","2":"21.39130","3":"5"},{"1":"673.55","2":"20.00833","3":"5"},{"1":"624.15","2":"20.29137","3":"5"},{"1":"669.30","2":"17.66667","3":"5"},{"1":"659.90","2":"18.22055","3":"5"},{"1":"616.30","2":"20.27100","3":"5"},{"1":"640.50","2":"20.19895","3":"5"},{"1":"661.95","2":"21.38424","3":"5"},{"1":"664.15","2":"20.97368","3":"5"},{"1":"651.45","2":"20.00000","3":"5"},{"1":"635.95","2":"17.15328","3":"5"},{"1":"618.30","2":"22.34977","3":"5"},{"1":"673.30","2":"22.17007","3":"5"},{"1":"644.45","2":"18.18182","3":"5"},{"1":"664.95","2":"18.95714","3":"5"},{"1":"672.45","2":"19.74533","3":"5"},{"1":"657.65","2":"16.42623","3":"5"},{"1":"652.00","2":"16.62540","3":"5"},{"1":"655.40","2":"16.38177","3":"5"},{"1":"646.20","2":"20.07416","3":"5"},{"1":"652.35","2":"17.99544","3":"5"},{"1":"655.85","2":"19.39130","3":"5"},{"1":"657.00","2":"16.42857","3":"5"},{"1":"682.15","2":"16.72949","3":"5"},{"1":"671.65","2":"24.41345","3":"5"},{"1":"633.65","2":"18.26415","3":"5"},{"1":"631.95","2":"18.95504","3":"5"},{"1":"643.70","2":"21.03896","3":"5"},{"1":"683.40","2":"20.74074","3":"5"},{"1":"661.85","2":"18.10000","3":"5"},{"1":"647.70","2":"19.84615","3":"5"},{"1":"672.35","2":"21.60000","3":"5"},{"1":"667.20","2":"22.44242","3":"5"},{"1":"681.60","2":"23.01438","3":"5"},{"1":"660.75","2":"17.74892","3":"5"},{"1":"638.20","2":"18.28664","3":"5"},{"1":"643.50","2":"19.26544","3":"5"},{"1":"659.05","2":"22.66667","3":"5"},{"1":"641.55","2":"19.29412","3":"5"},{"1":"619.80","2":"17.36364","3":"5"},{"1":"625.30","2":"19.82143","3":"5"},{"1":"634.90","2":"20.43378","3":"5"},{"1":"635.20","2":"21.03721","3":"5"},{"1":"644.70","2":"19.92462","3":"5"},{"1":"649.15","2":"19.00986","3":"5"},{"1":"665.35","2":"23.82222","3":"5"},{"1":"636.90","2":"19.36909","3":"5"},{"1":"686.05","2":"19.82857","3":"5"},{"1":"691.05","2":"15.25885","3":"5"},{"1":"652.10","2":"17.16129","3":"5"},{"1":"639.85","2":"21.81333","3":"5"},{"1":"657.50","2":"19.07471","3":"5"},{"1":"661.35","2":"25.78512","3":"5"},{"1":"655.55","2":"18.21261","3":"5"},{"1":"639.75","2":"18.16606","3":"5"},{"1":"628.55","2":"16.97297","3":"5"},{"1":"650.55","2":"21.50087","3":"5"},{"1":"699.10","2":"20.60000","3":"5"},{"1":"666.85","2":"16.99029","3":"5"},{"1":"634.20","2":"20.77954","3":"5"},{"1":"634.55","2":"15.51247","3":"5"},{"1":"654.10","2":"19.88506","3":"5"},{"1":"612.50","2":"21.39882","3":"5"},{"1":"649.50","2":"20.49751","3":"5"},{"1":"662.70","2":"19.36376","3":"5"},{"1":"637.35","2":"17.65957","3":"5"},{"1":"615.75","2":"21.01796","3":"5"},{"1":"657.55","2":"19.05565","3":"5"},{"1":"648.70","2":"22.53846","3":"5"},{"1":"657.00","2":"21.10787","3":"5"},{"1":"694.80","2":"20.05135","3":"5"},{"1":"661.60","2":"14.20176","3":"5"},{"1":"660.30","2":"18.47687","3":"5"},{"1":"639.35","2":"18.63542","3":"5"},{"1":"671.60","2":"20.94595","3":"5"},{"1":"647.60","2":"21.08548","3":"5"},{"1":"605.55","2":"18.69288","3":"5"},{"1":"640.90","2":"20.86808","3":"5"},{"1":"657.15","2":"19.82558","3":"5"},{"1":"666.00","2":"19.75000","3":"5"},{"1":"654.90","2":"19.50000","3":"5"},{"1":"653.95","2":"18.39080","3":"5"},{"1":"673.25","2":"18.78676","3":"5"},{"1":"662.55","2":"19.77018","3":"5"},{"1":"669.95","2":"19.33333","3":"5"},{"1":"630.35","2":"21.46392","3":"5"},{"1":"677.25","2":"23.08492","3":"5"},{"1":"666.55","2":"21.06299","3":"5"},{"1":"651.20","2":"18.68687","3":"5"},{"1":"668.80","2":"20.77024","3":"5"},{"1":"689.10","2":"19.30556","3":"5"},{"1":"625.85","2":"20.13280","3":"5"},{"1":"671.25","2":"20.66964","3":"5"},{"1":"653.55","2":"22.28155","3":"5"},{"1":"633.15","2":"20.60027","3":"5"},{"1":"623.60","2":"20.82734","3":"5"},{"1":"643.25","2":"19.22492","3":"5"},{"1":"668.60","2":"17.65477","3":"5"},{"1":"695.20","2":"17.00000","3":"5"},{"1":"661.45","2":"16.49773","3":"5"},{"1":"655.05","2":"19.78261","3":"5"},{"1":"671.60","2":"22.30216","3":"5"},{"1":"632.00","2":"17.73077","3":"5"},{"1":"621.40","2":"20.44836","3":"5"},{"1":"624.95","2":"20.37169","3":"5"},{"1":"667.15","2":"20.16479","3":"5"},{"1":"646.40","2":"21.61538","3":"5"},{"1":"666.65","2":"20.56143","3":"5"},{"1":"687.55","2":"19.95551","3":"5"},{"1":"662.50","2":"21.18387","3":"5"},{"1":"673.90","2":"18.81042","3":"5"},{"1":"645.55","2":"20.57838","3":"5"},{"1":"666.85","2":"18.32461","3":"5"},{"1":"646.55","2":"18.82063","3":"5"},{"1":"648.00","2":"20.81633","3":"5"},{"1":"654.85","2":"20.00000","3":"5"},{"1":"684.95","2":"19.68182","3":"5"},{"1":"698.25","2":"19.39018","3":"5"},{"1":"637.00","2":"20.92732","3":"5"},{"1":"616.45","2":"19.94437","3":"5"},{"1":"666.65","2":"20.79109","3":"5"},{"1":"651.80","2":"19.20354","3":"5"},{"1":"635.60","2":"19.02439","3":"5"},{"1":"663.45","2":"17.62058","3":"5"},{"1":"632.20","2":"20.23715","3":"5"},{"1":"628.75","2":"19.29374","3":"5"},{"1":"650.90","2":"18.82998","3":"5"},{"1":"622.60","2":"20.33949","3":"5"},{"1":"671.90","2":"19.22900","3":"5"},{"1":"671.75","2":"17.89130","3":"5"},{"1":"646.50","2":"19.51881","3":"5"},{"1":"669.30","2":"19.08451","3":"5"},{"1":"626.10","2":"19.93548","3":"5"},{"1":"616.30","2":"18.87326","3":"5"},{"1":"666.10","2":"20.14178","3":"5"},{"1":"645.25","2":"23.55637","3":"5"},{"1":"681.30","2":"21.46479","3":"5"},{"1":"655.80","2":"19.19101","3":"5"},{"1":"660.10","2":"20.13080","3":"5"},{"1":"643.40","2":"25.80000","3":"5"},{"1":"651.35","2":"18.77774","3":"5"},{"1":"658.00","2":"19.10982","3":"5"},{"1":"669.35","2":"19.70109","3":"5"},{"1":"651.90","2":"18.61594","3":"5"},{"1":"662.40","2":"20.99721","3":"5"},{"1":"662.45","2":"20.00000","3":"5"},{"1":"655.75","2":"20.98325","3":"5"},{"1":"655.05","2":"21.64262","3":"5"},{"1":"646.70","2":"20.02967","3":"5"},{"1":"658.35","2":"19.81140","3":"5"},{"1":"671.90","2":"18.00000","3":"5"},{"1":"668.00","2":"19.35811","3":"5"},{"1":"650.65","2":"20.17912","3":"5"},{"1":"683.35","2":"21.11986","3":"5"},{"1":"627.10","2":"23.38974","3":"5"},{"1":"662.75","2":"22.18182","3":"5"},{"1":"682.55","2":"19.94283","3":"5"},{"1":"631.90","2":"17.78826","3":"5"},{"1":"662.55","2":"14.70588","3":"5"},{"1":"647.05","2":"19.04077","3":"5"},{"1":"645.75","2":"20.89195","3":"5"},{"1":"663.90","2":"19.83851","3":"5"},{"1":"661.60","2":"19.52191","3":"5"},{"1":"686.70","2":"20.68622","3":"5"},{"1":"645.55","2":"18.18182","3":"5"},{"1":"647.25","2":"18.89224","3":"5"},{"1":"636.95","2":"24.88889","3":"5"},{"1":"657.80","2":"18.58064","3":"5"},{"1":"609.00","2":"18.04000","3":"5"},{"1":"674.25","2":"17.73399","3":"5"},{"1":"657.40","2":"21.45455","3":"5"},{"1":"661.20","2":"19.92343","3":"5"},{"1":"653.10","2":"20.33942","3":"5"},{"1":"662.90","2":"22.54608","3":"5"},{"1":"634.05","2":"21.10344","3":"5"},{"1":"651.15","2":"18.19743","3":"5"},{"1":"669.80","2":"20.10768","3":"5"},{"1":"638.30","2":"19.15984","3":"5"},{"1":"638.55","2":"19.54545","3":"5"},{"1":"665.90","2":"20.88889","3":"5"},{"1":"690.80","2":"18.39150","3":"5"},{"1":"662.40","2":"19.17990","3":"5"},{"1":"639.90","2":"19.39771","3":"5"},{"1":"632.95","2":"21.67827","3":"5"},{"1":"668.95","2":"19.28889","3":"5"},{"1":"644.20","2":"20.34927","3":"5"},{"1":"659.15","2":"20.96416","3":"5"},{"1":"663.35","2":"19.46039","3":"5"},{"1":"664.40","2":"19.28572","3":"5"},{"1":"668.10","2":"20.91979","3":"5"},{"1":"640.75","2":"20.90021","3":"5"},{"1":"638.00","2":"20.59575","3":"5"},{"1":"669.35","2":"19.37500","3":"5"},{"1":"637.10","2":"19.95122","3":"5"},{"1":"652.85","2":"18.84973","3":"5"},{"1":"616.30","2":"18.11787","3":"5"},{"1":"693.95","2":"19.18341","3":"5"},{"1":"635.60","2":"22.00000","3":"5"},{"1":"628.40","2":"21.58416","3":"5"},{"1":"648.95","2":"20.38889","3":"5"},{"1":"628.65","2":"16.29310","3":"5"},{"1":"621.75","2":"18.27778","3":"5"},{"1":"630.40","2":"19.37472","3":"5"},{"1":"639.50","2":"18.90909","3":"5"},{"1":"679.50","2":"16.40693","3":"5"},{"1":"647.30","2":"15.59140","3":"5"},{"1":"617.35","2":"18.70694","3":"5"},{"1":"626.80","2":"18.32985","3":"5"},{"1":"650.45","2":"17.90235","3":"5"},{"1":"663.50","2":"18.91157","3":"5"},{"1":"680.45","2":"20.32497","3":"5"},{"1":"666.70","2":"20.02457","3":"5"},{"1":"644.95","2":"24.00000","3":"5"},{"1":"643.50","2":"17.60784","3":"5"},{"1":"669.85","2":"19.34853","3":"5"},{"1":"676.85","2":"19.67846","3":"5"},{"1":"672.30","2":"18.72861","3":"5"},{"1":"684.30","2":"15.88235","3":"5"},{"1":"653.40","2":"20.05491","3":"5"},{"1":"644.45","2":"17.98825","3":"5"},{"1":"650.90","2":"16.96629","3":"5"},{"1":"636.10","2":"19.23937","3":"5"},{"1":"653.70","2":"19.19586","3":"5"},{"1":"664.00","2":"19.59906","3":"5"},{"1":"629.80","2":"20.54348","3":"5"},{"1":"695.30","2":"18.58848","3":"5"},{"1":"640.85","2":"15.60419","3":"5"},{"1":"656.50","2":"15.29304","3":"5"},{"1":"654.30","2":"17.65537","3":"5"},{"1":"664.00","2":"17.57976","3":"5"},{"1":"655.35","2":"22.33333","3":"5"},{"1":"646.35","2":"18.75000","3":"5"},{"1":"642.75","2":"18.10241","3":"5"},{"1":"700.30","2":"20.25641","3":"5"},{"1":"651.85","2":"18.80207","3":"5"},{"1":"652.10","2":"18.77230","3":"5"},{"1":"672.55","2":"20.40521","3":"5"},{"1":"634.00","2":"18.65079","3":"5"},{"1":"665.65","2":"20.70707","3":"5"},{"1":"635.05","2":"22.00000","3":"5"},{"1":"676.55","2":"17.69978","3":"5"},{"1":"651.40","2":"21.48329","3":"5"},{"1":"655.35","2":"16.70103","3":"5"},{"1":"666.15","2":"19.57567","3":"5"},{"1":"639.80","2":"17.25806","3":"5"},{"1":"666.15","2":"17.37526","3":"5"},{"1":"637.95","2":"17.34931","3":"5"},{"1":"653.55","2":"16.26229","3":"5"},{"1":"644.50","2":"17.70045","3":"5"},{"1":"644.20","2":"20.12881","3":"5"},{"1":"664.70","2":"18.26539","3":"5"},{"1":"671.70","2":"14.54214","3":"5"},{"1":"620.50","2":"19.15261","3":"5"},{"1":"640.10","2":"17.36574","3":"5"},{"1":"655.20","2":"15.13898","3":"5"},{"1":"691.35","2":"17.84266","3":"5"},{"1":"652.40","2":"15.40704","3":"5"},{"1":"631.40","2":"18.86534","3":"5"},{"1":"656.70","2":"16.47413","3":"5"},{"1":"606.75","2":"17.86263","3":"5"},{"1":"641.45","2":"21.88586","3":"5"},{"1":"671.30","2":"20.20000","3":"5"},{"1":"666.60","2":"19.03640","3":"5"},{"1":"657.80","2":"17.88991","3":"6"},{"1":"616.30","2":"21.52466","3":"6"},{"1":"660.20","2":"18.69723","3":"6"},{"1":"663.45","2":"17.35714","3":"6"},{"1":"635.95","2":"18.67133","3":"6"},{"1":"605.55","2":"21.40625","3":"6"},{"1":"606.75","2":"19.50000","3":"6"},{"1":"656.40","2":"20.89412","3":"6"},{"1":"661.95","2":"19.94737","3":"6"},{"1":"653.85","2":"20.80556","3":"6"},{"1":"637.95","2":"21.23809","3":"6"},{"1":"645.75","2":"21.00000","3":"6"},{"1":"623.60","2":"20.60000","3":"6"},{"1":"639.90","2":"20.00822","3":"6"},{"1":"643.40","2":"18.02778","3":"6"},{"1":"655.85","2":"20.25196","3":"6"},{"1":"678.05","2":"16.97787","3":"6"},{"1":"639.25","2":"16.50980","3":"6"},{"1":"680.45","2":"22.70402","3":"6"},{"1":"652.30","2":"19.91111","3":"6"},{"1":"637.10","2":"18.33333","3":"6"},{"1":"655.55","2":"22.61905","3":"6"},{"1":"658.00","2":"19.44828","3":"6"},{"1":"657.15","2":"25.05263","3":"6"},{"1":"662.55","2":"20.67544","3":"6"},{"1":"647.25","2":"18.68235","3":"6"},{"1":"686.05","2":"22.84553","3":"6"},{"1":"609.00","2":"19.26667","3":"6"},{"1":"659.40","2":"19.25000","3":"6"},{"1":"652.85","2":"20.54545","3":"6"},{"1":"620.30","2":"20.60697","3":"6"},{"1":"661.85","2":"21.07268","3":"6"},{"1":"646.35","2":"21.53581","3":"6"},{"1":"654.10","2":"19.90400","3":"6"},{"1":"644.20","2":"21.19407","3":"6"},{"1":"649.70","2":"21.86535","3":"6"},{"1":"698.45","2":"18.32965","3":"6"},{"1":"618.30","2":"16.22857","3":"6"},{"1":"644.50","2":"19.17857","3":"6"},{"1":"646.95","2":"20.27737","3":"6"},{"1":"641.80","2":"22.98614","3":"6"},{"1":"672.05","2":"20.44444","3":"6"},{"1":"637.35","2":"19.82085","3":"6"},{"1":"662.55","2":"23.20522","3":"6"},{"1":"664.45","2":"19.26697","3":"6"},{"1":"666.45","2":"23.30189","3":"6"},{"1":"626.90","2":"21.18829","3":"6"},{"1":"633.15","2":"20.87180","3":"6"},{"1":"660.05","2":"19.01749","3":"6"},{"1":"676.85","2":"21.91938","3":"6"},{"1":"636.10","2":"20.10124","3":"6"},{"1":"651.90","2":"21.47651","3":"6"},{"1":"669.95","2":"20.06579","3":"6"},{"1":"645.00","2":"20.37510","3":"6"},{"1":"643.70","2":"22.44648","3":"6"},{"1":"664.30","2":"22.89524","3":"6"},{"1":"616.30","2":"20.49797","3":"6"},{"1":"634.70","2":"20.00000","3":"6"},{"1":"655.05","2":"22.25658","3":"6"},{"1":"655.35","2":"21.56436","3":"6"},{"1":"662.40","2":"19.47737","3":"6"},{"1":"626.80","2":"17.67002","3":"6"},{"1":"679.50","2":"21.94756","3":"6"},{"1":"638.30","2":"21.78339","3":"6"},{"1":"691.35","2":"19.14000","3":"6"},{"1":"623.20","2":"18.11050","3":"6"},{"1":"653.95","2":"20.68242","3":"6"},{"1":"640.90","2":"22.62361","3":"6"},{"1":"656.65","2":"21.78650","3":"6"},{"1":"656.70","2":"18.58293","3":"6"},{"1":"652.10","2":"21.54545","3":"6"},{"1":"704.30","2":"21.15289","3":"6"},{"1":"652.00","2":"16.63333","3":"6"},{"1":"698.25","2":"21.14438","3":"6"},{"1":"678.40","2":"19.78182","3":"6"},{"1":"654.30","2":"18.98373","3":"6"},{"1":"665.35","2":"17.66767","3":"6"},{"1":"655.35","2":"17.75499","3":"6"},{"1":"630.40","2":"15.27273","3":"6"},{"1":"635.60","2":"14.00000","3":"6"},{"1":"630.55","2":"20.59613","3":"6"},{"1":"669.35","2":"16.31169","3":"6"},{"1":"671.95","2":"21.12796","3":"6"},{"1":"643.05","2":"17.48801","3":"6"},{"1":"679.80","2":"17.88679","3":"6"},{"1":"615.75","2":"19.30676","3":"6"},{"1":"659.40","2":"20.89231","3":"6"},{"1":"635.60","2":"21.28684","3":"6"},{"1":"664.00","2":"20.19560","3":"6"},{"1":"644.45","2":"24.95000","3":"6"},{"1":"663.85","2":"18.13043","3":"6"},{"1":"654.20","2":"20.00000","3":"6"},{"1":"656.80","2":"18.72951","3":"6"},{"1":"673.55","2":"18.25000","3":"6"},{"1":"640.50","2":"18.99257","3":"6"},{"1":"638.30","2":"19.88764","3":"6"},{"1":"624.95","2":"19.37895","3":"6"},{"1":"671.25","2":"20.46259","3":"6"},{"1":"636.50","2":"22.29157","3":"6"},{"1":"677.95","2":"20.70474","3":"6"},{"1":"695.20","2":"19.06005","3":"6"},{"1":"663.35","2":"20.23247","3":"6"},{"1":"630.35","2":"19.69012","3":"6"},{"1":"624.15","2":"20.36254","3":"6"},{"1":"681.30","2":"19.75422","3":"6"},{"1":"632.00","2":"19.37977","3":"6"},{"1":"672.35","2":"22.92351","3":"6"},{"1":"677.25","2":"19.37340","3":"6"},{"1":"673.25","2":"19.15516","3":"6"},{"1":"676.15","2":"21.30000","3":"6"},{"1":"619.80","2":"18.30357","3":"6"},{"1":"664.75","2":"21.07926","3":"6"},{"1":"625.85","2":"18.79121","3":"6"},{"1":"648.25","2":"19.62662","3":"6"},{"1":"661.60","2":"19.59016","3":"6"},{"1":"655.05","2":"20.87187","3":"6"},{"1":"623.10","2":"21.11500","3":"6"},{"1":"669.85","2":"20.08452","3":"6"},{"1":"658.80","2":"19.91049","3":"6"},{"1":"646.90","2":"17.81285","3":"6"},{"1":"634.95","2":"18.13333","3":"6"},{"1":"661.85","2":"19.22221","3":"6"},{"1":"631.05","2":"18.66072","3":"6"},{"1":"657.50","2":"19.60000","3":"6"},{"1":"649.50","2":"19.28384","3":"6"},{"1":"668.95","2":"22.81818","3":"6"},{"1":"644.40","2":"18.80922","3":"6"},{"1":"634.10","2":"21.37363","3":"6"},{"1":"668.40","2":"20.02041","3":"6"},{"1":"650.60","2":"21.49862","3":"6"},{"1":"661.35","2":"15.42857","3":"6"},{"1":"638.20","2":"22.40000","3":"6"},{"1":"638.70","2":"20.12709","3":"6"},{"1":"696.55","2":"19.03798","3":"6"},{"1":"676.60","2":"17.34216","3":"6"},{"1":"645.25","2":"17.01863","3":"6"},{"1":"628.65","2":"20.80000","3":"6"},{"1":"652.40","2":"21.15385","3":"6"},{"1":"634.55","2":"18.45833","3":"6"},{"1":"691.05","2":"19.14082","3":"6"},{"1":"666.10","2":"19.40766","3":"6"},{"1":"668.80","2":"19.56896","3":"6"},{"1":"687.55","2":"21.50120","3":"6"},{"1":"645.55","2":"17.52941","3":"6"},{"1":"661.85","2":"16.43017","3":"6"},{"1":"644.70","2":"19.79654","3":"6"},{"1":"650.55","2":"17.18613","3":"6"},{"1":"654.85","2":"17.61589","3":"6"},{"1":"653.40","2":"20.12537","3":"6"},{"1":"666.85","2":"22.16667","3":"6"},{"1":"642.40","2":"19.96154","3":"6"},{"1":"664.15","2":"19.03945","3":"6"},{"1":"621.40","2":"15.22436","3":"6"},{"1":"665.10","2":"21.14475","3":"6"},{"1":"617.35","2":"19.64390","3":"6"},{"1":"682.45","2":"21.04869","3":"6"},{"1":"637.65","2":"20.17544","3":"6"},{"1":"666.60","2":"21.39130","3":"6"},{"1":"651.35","2":"20.00833","3":"6"},{"1":"644.55","2":"20.29137","3":"6"},{"1":"683.35","2":"17.66667","3":"6"},{"1":"642.75","2":"18.22055","3":"6"},{"1":"672.45","2":"20.27100","3":"6"},{"1":"637.00","2":"20.19895","3":"6"},{"1":"669.30","2":"21.38424","3":"6"},{"1":"631.90","2":"20.97368","3":"6"},{"1":"634.90","2":"20.00000","3":"6"},{"1":"646.50","2":"17.15328","3":"6"},{"1":"634.40","2":"22.34977","3":"6"},{"1":"661.90","2":"22.17007","3":"6"},{"1":"635.45","2":"18.18182","3":"6"},{"1":"662.45","2":"18.95714","3":"6"},{"1":"672.70","2":"19.74533","3":"6"},{"1":"639.30","2":"16.42623","3":"6"},{"1":"641.45","2":"16.62540","3":"6"},{"1":"620.50","2":"16.38177","3":"6"},{"1":"679.40","2":"20.07416","3":"6"},{"1":"663.90","2":"17.99544","3":"6"},{"1":"628.75","2":"19.39130","3":"6"},{"1":"658.60","2":"16.42857","3":"6"},{"1":"655.20","2":"16.72949","3":"6"},{"1":"676.55","2":"24.41345","3":"6"},{"1":"656.80","2":"18.26415","3":"6"},{"1":"671.30","2":"18.95504","3":"6"},{"1":"638.55","2":"21.03896","3":"6"},{"1":"661.45","2":"20.74074","3":"6"},{"1":"671.65","2":"18.10000","3":"6"},{"1":"616.45","2":"19.84615","3":"6"},{"1":"622.05","2":"21.60000","3":"6"},{"1":"652.50","2":"22.44242","3":"6"},{"1":"669.80","2":"23.01438","3":"6"},{"1":"634.00","2":"17.74892","3":"6"},{"1":"632.85","2":"18.28664","3":"6"},{"1":"657.00","2":"19.26544","3":"6"},{"1":"627.25","2":"22.66667","3":"6"},{"1":"668.00","2":"19.29412","3":"6"},{"1":"647.30","2":"17.36364","3":"6"},{"1":"624.55","2":"19.82143","3":"6"},{"1":"636.70","2":"20.43378","3":"6"},{"1":"643.20","2":"21.03721","3":"6"},{"1":"686.70","2":"19.92462","3":"6"},{"1":"682.55","2":"19.00986","3":"6"},{"1":"662.70","2":"23.82222","3":"6"},{"1":"673.55","2":"19.36909","3":"6"},{"1":"698.20","2":"19.82857","3":"6"},{"1":"644.95","2":"15.25885","3":"6"},{"1":"627.10","2":"17.16129","3":"6"},{"1":"643.50","2":"21.81333","3":"6"},{"1":"671.70","2":"19.07471","3":"6"},{"1":"664.95","2":"25.78512","3":"6"},{"1":"667.15","2":"18.21261","3":"6"},{"1":"642.20","2":"18.16606","3":"6"},{"1":"612.65","2":"16.97297","3":"6"},{"1":"650.65","2":"21.50087","3":"6"},{"1":"683.40","2":"20.60000","3":"6"},{"1":"632.95","2":"16.99029","3":"6"},{"1":"671.60","2":"20.77954","3":"6"},{"1":"634.05","2":"15.51247","3":"6"},{"1":"648.20","2":"19.88506","3":"6"},{"1":"682.65","2":"21.39882","3":"6"},{"1":"684.80","2":"20.49751","3":"6"},{"1":"663.50","2":"19.36376","3":"6"},{"1":"639.35","2":"17.65957","3":"6"},{"1":"684.95","2":"21.01796","3":"6"},{"1":"643.70","2":"19.05565","3":"6"},{"1":"670.70","2":"22.53846","3":"6"},{"1":"668.65","2":"21.10787","3":"6"},{"1":"628.40","2":"20.05135","3":"6"},{"1":"668.60","2":"14.20176","3":"6"},{"1":"657.55","2":"18.47687","3":"6"},{"1":"664.70","2":"18.63542","3":"6"},{"1":"660.95","2":"20.94595","3":"6"},{"1":"637.95","2":"21.08548","3":"6"},{"1":"650.90","2":"18.69288","3":"6"},{"1":"673.30","2":"20.86808","3":"6"},{"1":"644.20","2":"19.82558","3":"6"},{"1":"657.40","2":"19.75000","3":"6"},{"1":"646.40","2":"19.50000","3":"6"},{"1":"662.65","2":"18.39080","3":"6"},{"1":"653.55","2":"18.78676","3":"6"},{"1":"649.85","2":"19.77018","3":"6"},{"1":"666.55","2":"19.33333","3":"6"},{"1":"657.65","2":"21.46392","3":"6"},{"1":"648.00","2":"23.08492","3":"6"},{"1":"643.60","2":"21.06299","3":"6"},{"1":"660.75","2":"18.68687","3":"6"},{"1":"668.10","2":"20.77024","3":"6"},{"1":"645.10","2":"19.30556","3":"6"},{"1":"651.45","2":"20.13280","3":"6"},{"1":"653.70","2":"20.66964","3":"6"},{"1":"673.90","2":"22.28155","3":"6"},{"1":"690.80","2":"20.60027","3":"6"},{"1":"636.95","2":"20.82734","3":"6"},{"1":"661.20","2":"19.22492","3":"6"},{"1":"665.20","2":"17.65477","3":"6"},{"1":"665.95","2":"17.00000","3":"6"},{"1":"645.55","2":"16.49773","3":"6"},{"1":"641.10","2":"19.78261","3":"6"},{"1":"684.35","2":"22.30216","3":"6"},{"1":"664.40","2":"17.73077","3":"6"},{"1":"652.10","2":"20.44836","3":"6"},{"1":"635.05","2":"20.37169","3":"6"},{"1":"671.90","2":"20.16479","3":"6"},{"1":"647.70","2":"21.61538","3":"6"},{"1":"666.15","2":"20.56143","3":"6"},{"1":"667.20","2":"19.95551","3":"6"},{"1":"653.10","2":"21.18387","3":"6"},{"1":"641.45","2":"18.81042","3":"6"},{"1":"670.00","2":"20.57838","3":"6"},{"1":"666.65","2":"18.32461","3":"6"},{"1":"665.65","2":"18.82063","3":"6"},{"1":"633.65","2":"20.81633","3":"6"},{"1":"694.25","2":"20.00000","3":"6"},{"1":"666.85","2":"19.68182","3":"6"},{"1":"680.05","2":"19.39018","3":"6"},{"1":"639.85","2":"20.92732","3":"6"},{"1":"634.15","2":"19.94437","3":"6"},{"1":"628.25","2":"20.79109","3":"6"},{"1":"653.80","2":"19.20354","3":"6"},{"1":"660.30","2":"19.02439","3":"6"},{"1":"689.10","2":"17.62058","3":"6"},{"1":"640.15","2":"20.23715","3":"6"},{"1":"669.10","2":"19.29374","3":"6"},{"1":"655.40","2":"18.82998","3":"6"},{"1":"649.30","2":"20.33949","3":"6"},{"1":"678.80","2":"19.22900","3":"6"},{"1":"674.25","2":"17.89130","3":"6"},{"1":"662.90","2":"19.51881","3":"6"},{"1":"628.55","2":"19.08451","3":"6"},{"1":"656.55","2":"19.93548","3":"6"},{"1":"675.40","2":"18.87326","3":"6"},{"1":"655.30","2":"20.14178","3":"6"},{"1":"671.75","2":"23.55637","3":"6"},{"1":"673.05","2":"21.46479","3":"6"},{"1":"659.80","2":"19.19101","3":"6"},{"1":"700.30","2":"20.13080","3":"6"},{"1":"635.20","2":"25.80000","3":"6"},{"1":"642.20","2":"18.77774","3":"6"},{"1":"632.20","2":"19.10982","3":"6"},{"1":"667.45","2":"19.70109","3":"6"},{"1":"679.65","2":"18.61594","3":"6"},{"1":"651.15","2":"20.99721","3":"6"},{"1":"659.15","2":"20.00000","3":"6"},{"1":"653.50","2":"20.98325","3":"6"},{"1":"682.15","2":"21.64262","3":"6"},{"1":"654.85","2":"20.02967","3":"6"},{"1":"638.00","2":"19.81140","3":"6"},{"1":"664.95","2":"18.00000","3":"6"},{"1":"699.10","2":"19.35811","3":"6"},{"1":"666.15","2":"20.17912","3":"6"},{"1":"657.75","2":"21.11986","3":"6"},{"1":"662.40","2":"23.38974","3":"6"},{"1":"693.95","2":"22.18182","3":"6"},{"1":"657.00","2":"19.94283","3":"6"},{"1":"662.75","2":"17.78826","3":"6"},{"1":"650.90","2":"14.70588","3":"6"},{"1":"653.55","2":"19.04077","3":"6"},{"1":"672.20","2":"20.89195","3":"6"},{"1":"645.75","2":"19.83851","3":"6"},{"1":"636.60","2":"19.52191","3":"6"},{"1":"650.45","2":"20.68622","3":"6"},{"1":"636.90","2":"18.18182","3":"6"},{"1":"681.90","2":"18.89224","3":"6"},{"1":"641.55","2":"24.88889","3":"6"},{"1":"644.45","2":"18.58064","3":"6"},{"1":"666.70","2":"18.04000","3":"6"},{"1":"651.85","2":"17.73399","3":"6"},{"1":"633.05","2":"21.45455","3":"6"},{"1":"694.80","2":"19.92343","3":"6"},{"1":"632.25","2":"20.33942","3":"6"},{"1":"661.90","2":"22.54608","3":"6"},{"1":"654.90","2":"21.10344","3":"6"},{"1":"652.40","2":"18.19743","3":"6"},{"1":"648.35","2":"20.10768","3":"6"},{"1":"639.75","2":"19.15984","3":"6"},{"1":"659.35","2":"19.54545","3":"6"},{"1":"652.35","2":"20.88889","3":"6"},{"1":"663.85","2":"18.39150","3":"6"},{"1":"672.30","2":"19.17990","3":"6"},{"1":"621.75","2":"19.39771","3":"6"},{"1":"659.90","2":"21.67827","3":"6"},{"1":"662.50","2":"19.28889","3":"6"},{"1":"660.10","2":"20.34927","3":"6"},{"1":"664.15","2":"20.96416","3":"6"},{"1":"631.95","2":"19.46039","3":"6"},{"1":"616.30","2":"19.28572","3":"6"},{"1":"655.05","2":"20.91979","3":"6"},{"1":"676.95","2":"20.90021","3":"6"},{"1":"646.00","2":"20.59575","3":"6"},{"1":"669.30","2":"19.37500","3":"6"},{"1":"651.40","2":"19.95122","3":"6"},{"1":"645.60","2":"18.84973","3":"6"},{"1":"626.10","2":"18.11787","3":"6"},{"1":"658.35","2":"19.18341","3":"6"},{"1":"612.50","2":"22.00000","3":"6"},{"1":"681.60","2":"21.58416","3":"6"},{"1":"706.75","2":"20.38889","3":"6"},{"1":"684.30","2":"16.29310","3":"6"},{"1":"691.90","2":"18.27778","3":"6"},{"1":"640.75","2":"19.37472","3":"6"},{"1":"634.10","2":"18.90909","3":"6"},{"1":"651.20","2":"16.40693","3":"6"},{"1":"655.75","2":"15.59140","3":"6"},{"1":"666.65","2":"18.70694","3":"6"},{"1":"649.15","2":"18.32985","3":"6"},{"1":"672.55","2":"17.90235","3":"6"},{"1":"652.30","2":"18.91157","3":"6"},{"1":"622.60","2":"20.32497","3":"6"},{"1":"631.40","2":"20.02457","3":"6"},{"1":"625.30","2":"24.00000","3":"6"},{"1":"656.50","2":"17.60784","3":"6"},{"1":"629.80","2":"19.34853","3":"6"},{"1":"618.05","2":"19.67846","3":"6"},{"1":"643.40","2":"18.72861","3":"6"},{"1":"679.75","2":"15.88235","3":"6"},{"1":"695.30","2":"20.05491","3":"6"},{"1":"623.45","2":"17.98825","3":"6"},{"1":"654.60","2":"16.96629","3":"6"},{"1":"671.90","2":"19.23937","3":"6"},{"1":"675.70","2":"19.19586","3":"6"},{"1":"655.70","2":"19.59906","3":"6"},{"1":"671.60","2":"20.54348","3":"6"},{"1":"640.10","2":"18.58848","3":"6"},{"1":"643.50","2":"15.60419","3":"6"},{"1":"672.05","2":"15.29304","3":"6"},{"1":"648.95","2":"17.65537","3":"6"},{"1":"651.80","2":"17.57976","3":"6"},{"1":"639.50","2":"22.33333","3":"6"},{"1":"635.75","2":"18.75000","3":"6"},{"1":"643.25","2":"18.10241","3":"6"},{"1":"646.55","2":"20.25641","3":"6"},{"1":"681.30","2":"18.80207","3":"6"},{"1":"634.20","2":"18.77230","3":"6"},{"1":"666.05","2":"20.40521","3":"6"},{"1":"655.80","2":"18.65079","3":"6"},{"1":"666.00","2":"20.70707","3":"6"},{"1":"638.35","2":"22.00000","3":"6"},{"1":"667.60","2":"17.69978","3":"6"},{"1":"648.70","2":"21.48329","3":"6"},{"1":"667.45","2":"16.70103","3":"6"},{"1":"647.60","2":"19.57567","3":"6"},{"1":"669.35","2":"17.25806","3":"6"},{"1":"661.60","2":"17.37526","3":"6"},{"1":"630.55","2":"17.34931","3":"6"},{"1":"668.90","2":"16.26229","3":"6"},{"1":"640.85","2":"17.70045","3":"6"},{"1":"627.30","2":"20.12881","3":"6"},{"1":"633.90","2":"18.26539","3":"6"},{"1":"657.90","2":"14.54214","3":"6"},{"1":"646.70","2":"19.15261","3":"6"},{"1":"665.90","2":"17.36574","3":"6"},{"1":"659.05","2":"15.13898","3":"6"},{"1":"639.80","2":"17.84266","3":"6"},{"1":"646.20","2":"15.40704","3":"6"},{"1":"654.20","2":"18.86534","3":"6"},{"1":"632.45","2":"16.47413","3":"6"},{"1":"631.85","2":"17.86263","3":"6"},{"1":"647.60","2":"21.88586","3":"6"},{"1":"647.05","2":"20.20000","3":"6"},{"1":"664.00","2":"19.03640","3":"6"},{"1":"652.30","2":"17.88991","3":"7"},{"1":"632.45","2":"21.52466","3":"7"},{"1":"656.65","2":"18.69723","3":"7"},{"1":"653.55","2":"17.35714","3":"7"},{"1":"668.80","2":"18.67133","3":"7"},{"1":"633.15","2":"21.40625","3":"7"},{"1":"621.75","2":"19.50000","3":"7"},{"1":"648.25","2":"20.89412","3":"7"},{"1":"654.30","2":"19.94737","3":"7"},{"1":"662.65","2":"20.80556","3":"7"},{"1":"638.35","2":"21.23809","3":"7"},{"1":"631.95","2":"21.00000","3":"7"},{"1":"654.60","2":"20.60000","3":"7"},{"1":"627.30","2":"20.00822","3":"7"},{"1":"635.60","2":"18.02778","3":"7"},{"1":"649.70","2":"20.25196","3":"7"},{"1":"656.70","2":"16.97787","3":"7"},{"1":"672.45","2":"16.50980","3":"7"},{"1":"662.75","2":"22.70402","3":"7"},{"1":"660.95","2":"19.91111","3":"7"},{"1":"656.80","2":"18.33333","3":"7"},{"1":"634.95","2":"22.61905","3":"7"},{"1":"660.75","2":"19.44828","3":"7"},{"1":"638.30","2":"25.05263","3":"7"},{"1":"650.90","2":"20.67544","3":"7"},{"1":"676.85","2":"18.68235","3":"7"},{"1":"629.80","2":"22.84553","3":"7"},{"1":"669.80","2":"19.26667","3":"7"},{"1":"655.30","2":"19.25000","3":"7"},{"1":"617.35","2":"20.54545","3":"7"},{"1":"696.55","2":"20.60697","3":"7"},{"1":"682.65","2":"21.07268","3":"7"},{"1":"632.85","2":"21.53581","3":"7"},{"1":"647.70","2":"19.90400","3":"7"},{"1":"664.00","2":"21.19407","3":"7"},{"1":"678.40","2":"21.86535","3":"7"},{"1":"630.55","2":"18.32965","3":"7"},{"1":"657.00","2":"16.22857","3":"7"},{"1":"672.30","2":"19.17857","3":"7"},{"1":"643.40","2":"20.27737","3":"7"},{"1":"658.00","2":"22.98614","3":"7"},{"1":"654.85","2":"20.44444","3":"7"},{"1":"657.90","2":"19.82085","3":"7"},{"1":"662.50","2":"23.20522","3":"7"},{"1":"666.10","2":"19.26697","3":"7"},{"1":"630.40","2":"23.30189","3":"7"},{"1":"699.10","2":"21.18829","3":"7"},{"1":"653.50","2":"20.87180","3":"7"},{"1":"628.75","2":"19.01749","3":"7"},{"1":"632.00","2":"21.91938","3":"7"},{"1":"618.05","2":"20.10124","3":"7"},{"1":"694.80","2":"21.47651","3":"7"},{"1":"634.70","2":"20.06579","3":"7"},{"1":"669.30","2":"20.37510","3":"7"},{"1":"637.00","2":"22.44648","3":"7"},{"1":"620.30","2":"22.89524","3":"7"},{"1":"628.55","2":"20.49797","3":"7"},{"1":"684.30","2":"20.00000","3":"7"},{"1":"637.35","2":"22.25658","3":"7"},{"1":"666.65","2":"21.56436","3":"7"},{"1":"670.00","2":"19.47737","3":"7"},{"1":"623.20","2":"17.67002","3":"7"},{"1":"654.85","2":"21.94756","3":"7"},{"1":"648.95","2":"21.78339","3":"7"},{"1":"639.85","2":"19.14000","3":"7"},{"1":"661.85","2":"18.11050","3":"7"},{"1":"667.20","2":"20.68242","3":"7"},{"1":"671.65","2":"22.62361","3":"7"},{"1":"648.20","2":"21.78650","3":"7"},{"1":"646.95","2":"18.58293","3":"7"},{"1":"646.20","2":"21.54545","3":"7"},{"1":"630.55","2":"21.15289","3":"7"},{"1":"631.05","2":"16.63333","3":"7"},{"1":"664.45","2":"21.14438","3":"7"},{"1":"661.90","2":"19.78182","3":"7"},{"1":"676.15","2":"18.98373","3":"7"},{"1":"663.85","2":"17.66767","3":"7"},{"1":"636.95","2":"17.75499","3":"7"},{"1":"683.35","2":"15.27273","3":"7"},{"1":"631.90","2":"14.00000","3":"7"},{"1":"656.40","2":"20.59613","3":"7"},{"1":"648.70","2":"16.31169","3":"7"},{"1":"638.00","2":"21.12796","3":"7"},{"1":"652.40","2":"17.48801","3":"7"},{"1":"641.10","2":"17.88679","3":"7"},{"1":"679.40","2":"19.30676","3":"7"},{"1":"645.55","2":"20.89231","3":"7"},{"1":"684.80","2":"21.28684","3":"7"},{"1":"658.60","2":"20.19560","3":"7"},{"1":"664.30","2":"24.95000","3":"7"},{"1":"612.65","2":"18.13043","3":"7"},{"1":"639.90","2":"20.00000","3":"7"},{"1":"661.85","2":"18.72951","3":"7"},{"1":"644.45","2":"18.25000","3":"7"},{"1":"644.20","2":"18.99257","3":"7"},{"1":"649.50","2":"19.88764","3":"7"},{"1":"634.40","2":"19.37895","3":"7"},{"1":"669.35","2":"20.46259","3":"7"},{"1":"641.45","2":"22.29157","3":"7"},{"1":"636.90","2":"20.70474","3":"7"},{"1":"616.30","2":"19.06005","3":"7"},{"1":"666.05","2":"20.23247","3":"7"},{"1":"664.40","2":"19.69012","3":"7"},{"1":"631.85","2":"20.36254","3":"7"},{"1":"654.90","2":"19.75422","3":"7"},{"1":"644.95","2":"19.37977","3":"7"},{"1":"663.45","2":"22.92351","3":"7"},{"1":"634.10","2":"19.37340","3":"7"},{"1":"650.90","2":"19.15516","3":"7"},{"1":"622.05","2":"21.30000","3":"7"},{"1":"674.25","2":"18.30357","3":"7"},{"1":"666.00","2":"21.07926","3":"7"},{"1":"671.30","2":"18.79121","3":"7"},{"1":"635.95","2":"19.62662","3":"7"},{"1":"695.20","2":"19.59016","3":"7"},{"1":"633.65","2":"20.87187","3":"7"},{"1":"662.45","2":"21.11500","3":"7"},{"1":"653.95","2":"20.08452","3":"7"},{"1":"639.80","2":"19.91049","3":"7"},{"1":"665.65","2":"17.81285","3":"7"},{"1":"657.55","2":"18.13333","3":"7"},{"1":"657.00","2":"19.22221","3":"7"},{"1":"681.90","2":"18.66072","3":"7"},{"1":"626.90","2":"19.60000","3":"7"},{"1":"645.75","2":"19.28384","3":"7"},{"1":"676.60","2":"22.81818","3":"7"},{"1":"633.90","2":"18.80922","3":"7"},{"1":"651.90","2":"21.37363","3":"7"},{"1":"682.15","2":"20.02041","3":"7"},{"1":"672.35","2":"21.49862","3":"7"},{"1":"663.90","2":"15.42857","3":"7"},{"1":"676.95","2":"22.40000","3":"7"},{"1":"683.40","2":"20.12709","3":"7"},{"1":"615.75","2":"19.03798","3":"7"},{"1":"694.25","2":"17.34216","3":"7"},{"1":"650.60","2":"17.01863","3":"7"},{"1":"628.40","2":"20.80000","3":"7"},{"1":"623.10","2":"21.15385","3":"7"},{"1":"618.30","2":"18.45833","3":"7"},{"1":"659.35","2":"19.14082","3":"7"},{"1":"686.70","2":"19.40766","3":"7"},{"1":"668.40","2":"19.56896","3":"7"},{"1":"616.30","2":"21.50120","3":"7"},{"1":"655.35","2":"17.52941","3":"7"},{"1":"659.40","2":"16.43017","3":"7"},{"1":"665.95","2":"19.79654","3":"7"},{"1":"656.50","2":"17.18613","3":"7"},{"1":"644.45","2":"17.61589","3":"7"},{"1":"635.75","2":"20.12537","3":"7"},{"1":"634.90","2":"22.16667","3":"7"},{"1":"658.80","2":"19.96154","3":"7"},{"1":"704.30","2":"19.03945","3":"7"},{"1":"624.95","2":"15.22436","3":"7"},{"1":"640.85","2":"21.14475","3":"7"},{"1":"646.50","2":"19.64390","3":"7"},{"1":"657.15","2":"21.04869","3":"7"},{"1":"686.05","2":"20.17544","3":"7"},{"1":"691.05","2":"21.39130","3":"7"},{"1":"655.05","2":"20.00833","3":"7"},{"1":"681.30","2":"20.29137","3":"7"},{"1":"679.80","2":"17.66667","3":"7"},{"1":"634.05","2":"18.22055","3":"7"},{"1":"659.05","2":"20.27100","3":"7"},{"1":"656.55","2":"20.19895","3":"7"},{"1":"657.50","2":"21.38424","3":"7"},{"1":"616.45","2":"20.97368","3":"7"},{"1":"625.30","2":"20.00000","3":"7"},{"1":"655.75","2":"17.15328","3":"7"},{"1":"661.95","2":"22.34977","3":"7"},{"1":"659.80","2":"22.17007","3":"7"},{"1":"635.20","2":"18.18182","3":"7"},{"1":"668.95","2":"18.95714","3":"7"},{"1":"678.05","2":"19.74533","3":"7"},{"1":"661.60","2":"16.42623","3":"7"},{"1":"645.55","2":"16.62540","3":"7"},{"1":"664.70","2":"16.38177","3":"7"},{"1":"624.55","2":"20.07416","3":"7"},{"1":"661.85","2":"17.99544","3":"7"},{"1":"661.45","2":"19.39130","3":"7"},{"1":"647.05","2":"16.42857","3":"7"},{"1":"642.40","2":"16.72949","3":"7"},{"1":"644.50","2":"24.41345","3":"7"},{"1":"626.10","2":"18.26415","3":"7"},{"1":"691.35","2":"18.95504","3":"7"},{"1":"665.35","2":"21.03896","3":"7"},{"1":"626.80","2":"20.74074","3":"7"},{"1":"684.35","2":"18.10000","3":"7"},{"1":"632.25","2":"19.84615","3":"7"},{"1":"654.20","2":"21.60000","3":"7"},{"1":"660.20","2":"22.44242","3":"7"},{"1":"639.25","2":"23.01438","3":"7"},{"1":"639.30","2":"17.74892","3":"7"},{"1":"669.35","2":"18.28664","3":"7"},{"1":"647.25","2":"19.26544","3":"7"},{"1":"630.35","2":"22.66667","3":"7"},{"1":"634.10","2":"19.29412","3":"7"},{"1":"653.55","2":"17.36364","3":"7"},{"1":"647.60","2":"19.82143","3":"7"},{"1":"638.70","2":"20.43378","3":"7"},{"1":"659.15","2":"21.03721","3":"7"},{"1":"657.75","2":"19.92462","3":"7"},{"1":"661.90","2":"19.00986","3":"7"},{"1":"636.10","2":"23.82222","3":"7"},{"1":"671.75","2":"19.36909","3":"7"},{"1":"687.55","2":"19.82857","3":"7"},{"1":"644.40","2":"15.25885","3":"7"},{"1":"637.95","2":"17.16129","3":"7"},{"1":"653.85","2":"21.81333","3":"7"},{"1":"680.05","2":"19.07471","3":"7"},{"1":"620.50","2":"25.78512","3":"7"},{"1":"671.25","2":"18.21261","3":"7"},{"1":"675.70","2":"18.16606","3":"7"},{"1":"672.05","2":"16.97297","3":"7"},{"1":"643.40","2":"21.50087","3":"7"},{"1":"643.50","2":"20.60000","3":"7"},{"1":"651.20","2":"16.99029","3":"7"},{"1":"655.40","2":"20.77954","3":"7"},{"1":"670.70","2":"15.51247","3":"7"},{"1":"673.25","2":"19.88506","3":"7"},{"1":"655.85","2":"21.39882","3":"7"},{"1":"609.00","2":"20.49751","3":"7"},{"1":"668.10","2":"19.36376","3":"7"},{"1":"637.65","2":"17.65957","3":"7"},{"1":"665.20","2":"21.01796","3":"7"},{"1":"677.25","2":"19.05565","3":"7"},{"1":"656.80","2":"22.53846","3":"7"},{"1":"655.70","2":"21.10787","3":"7"},{"1":"667.45","2":"20.05135","3":"7"},{"1":"646.90","2":"14.20176","3":"7"},{"1":"643.70","2":"18.47687","3":"7"},{"1":"651.15","2":"18.63542","3":"7"},{"1":"644.55","2":"20.94595","3":"7"},{"1":"671.90","2":"21.08548","3":"7"},{"1":"642.20","2":"18.69288","3":"7"},{"1":"662.40","2":"20.86808","3":"7"},{"1":"657.80","2":"19.82558","3":"7"},{"1":"661.35","2":"19.75000","3":"7"},{"1":"659.90","2":"19.50000","3":"7"},{"1":"645.60","2":"18.39080","3":"7"},{"1":"650.45","2":"18.78676","3":"7"},{"1":"669.10","2":"19.77018","3":"7"},{"1":"631.40","2":"19.33333","3":"7"},{"1":"666.70","2":"21.46392","3":"7"},{"1":"668.60","2":"23.08492","3":"7"},{"1":"653.40","2":"21.06299","3":"7"},{"1":"640.50","2":"18.68687","3":"7"},{"1":"651.80","2":"20.77024","3":"7"},{"1":"636.70","2":"19.30556","3":"7"},{"1":"638.20","2":"20.13280","3":"7"},{"1":"673.55","2":"20.66964","3":"7"},{"1":"619.80","2":"22.28155","3":"7"},{"1":"655.80","2":"20.60027","3":"7"},{"1":"664.15","2":"20.82734","3":"7"},{"1":"652.50","2":"19.22492","3":"7"},{"1":"667.15","2":"17.65477","3":"7"},{"1":"621.40","2":"17.00000","3":"7"},{"1":"668.65","2":"16.49773","3":"7"},{"1":"636.60","2":"19.78261","3":"7"},{"1":"651.85","2":"22.30216","3":"7"},{"1":"673.55","2":"17.73077","3":"7"},{"1":"623.45","2":"20.44836","3":"7"},{"1":"650.55","2":"20.37169","3":"7"},{"1":"662.40","2":"20.16479","3":"7"},{"1":"669.85","2":"21.61538","3":"7"},{"1":"646.70","2":"20.56143","3":"7"},{"1":"641.45","2":"19.95551","3":"7"},{"1":"671.70","2":"21.18387","3":"7"},{"1":"664.75","2":"18.81042","3":"7"},{"1":"667.45","2":"20.57838","3":"7"},{"1":"679.50","2":"18.32461","3":"7"},{"1":"658.35","2":"18.82063","3":"7"},{"1":"663.85","2":"20.81633","3":"7"},{"1":"622.60","2":"20.00000","3":"7"},{"1":"668.90","2":"19.68182","3":"7"},{"1":"649.85","2":"19.39018","3":"7"},{"1":"651.45","2":"20.92732","3":"7"},{"1":"680.45","2":"19.94437","3":"7"},{"1":"666.45","2":"20.79109","3":"7"},{"1":"654.20","2":"19.20354","3":"7"},{"1":"666.60","2":"19.02439","3":"7"},{"1":"636.50","2":"17.62058","3":"7"},{"1":"689.10","2":"20.23715","3":"7"},{"1":"640.15","2":"19.29374","3":"7"},{"1":"672.20","2":"18.82998","3":"7"},{"1":"652.10","2":"20.33949","3":"7"},{"1":"652.00","2":"19.22900","3":"7"},{"1":"648.35","2":"17.89130","3":"7"},{"1":"627.10","2":"19.51881","3":"7"},{"1":"676.55","2":"19.08451","3":"7"},{"1":"646.40","2":"19.93548","3":"7"},{"1":"665.90","2":"18.87326","3":"7"},{"1":"634.20","2":"20.14178","3":"7"},{"1":"671.60","2":"23.55637","3":"7"},{"1":"616.30","2":"21.46479","3":"7"},{"1":"698.25","2":"19.19101","3":"7"},{"1":"671.60","2":"20.13080","3":"7"},{"1":"681.30","2":"25.80000","3":"7"},{"1":"666.55","2":"18.77774","3":"7"},{"1":"643.70","2":"19.10982","3":"7"},{"1":"642.20","2":"19.70109","3":"7"},{"1":"668.00","2":"18.61594","3":"7"},{"1":"653.10","2":"20.99721","3":"7"},{"1":"655.20","2":"20.00000","3":"7"},{"1":"655.35","2":"20.98325","3":"7"},{"1":"644.70","2":"21.64262","3":"7"},{"1":"645.00","2":"20.02967","3":"7"},{"1":"638.55","2":"19.81140","3":"7"},{"1":"664.00","2":"18.00000","3":"7"},{"1":"660.30","2":"19.35811","3":"7"},{"1":"666.85","2":"20.17912","3":"7"},{"1":"633.05","2":"21.11986","3":"7"},{"1":"693.95","2":"23.38974","3":"7"},{"1":"657.40","2":"22.18182","3":"7"},{"1":"672.55","2":"19.94283","3":"7"},{"1":"652.35","2":"17.78826","3":"7"},{"1":"667.60","2":"14.70588","3":"7"},{"1":"673.30","2":"19.04077","3":"7"},{"1":"644.20","2":"20.89195","3":"7"},{"1":"653.80","2":"19.83851","3":"7"},{"1":"682.45","2":"19.52191","3":"7"},{"1":"652.30","2":"20.68622","3":"7"},{"1":"649.15","2":"18.18182","3":"7"},{"1":"666.15","2":"18.89224","3":"7"},{"1":"681.60","2":"24.88889","3":"7"},{"1":"679.65","2":"18.58064","3":"7"},{"1":"642.75","2":"18.04000","3":"7"},{"1":"634.55","2":"17.73399","3":"7"},{"1":"655.55","2":"21.45455","3":"7"},{"1":"639.50","2":"19.92343","3":"7"},{"1":"612.50","2":"20.33942","3":"7"},{"1":"627.25","2":"22.54608","3":"7"},{"1":"646.00","2":"21.10344","3":"7"},{"1":"672.05","2":"18.19743","3":"7"},{"1":"706.75","2":"20.10768","3":"7"},{"1":"645.75","2":"19.15984","3":"7"},{"1":"654.10","2":"19.54545","3":"7"},{"1":"643.05","2":"20.88889","3":"7"},{"1":"648.00","2":"18.39150","3":"7"},{"1":"691.90","2":"19.17990","3":"7"},{"1":"647.60","2":"19.39771","3":"7"},{"1":"643.20","2":"21.67827","3":"7"},{"1":"640.90","2":"19.28889","3":"7"},{"1":"673.90","2":"20.34927","3":"7"},{"1":"664.95","2":"20.96416","3":"7"},{"1":"664.95","2":"19.46039","3":"7"},{"1":"640.10","2":"19.28572","3":"7"},{"1":"641.80","2":"20.91979","3":"7"},{"1":"661.60","2":"20.90021","3":"7"},{"1":"661.20","2":"20.59575","3":"7"},{"1":"662.90","2":"19.37500","3":"7"},{"1":"666.15","2":"19.95122","3":"7"},{"1":"624.15","2":"18.84973","3":"7"},{"1":"646.35","2":"18.11787","3":"7"},{"1":"698.45","2":"19.18341","3":"7"},{"1":"672.70","2":"22.00000","3":"7"},{"1":"606.75","2":"21.58416","3":"7"},{"1":"678.80","2":"20.38889","3":"7"},{"1":"651.35","2":"16.29310","3":"7"},{"1":"643.25","2":"18.27778","3":"7"},{"1":"623.60","2":"19.37472","3":"7"},{"1":"646.55","2":"18.90909","3":"7"},{"1":"666.85","2":"16.40693","3":"7"},{"1":"666.65","2":"15.59140","3":"7"},{"1":"663.35","2":"18.70694","3":"7"},{"1":"698.20","2":"18.32985","3":"7"},{"1":"660.10","2":"17.90235","3":"7"},{"1":"690.80","2":"18.91157","3":"7"},{"1":"700.30","2":"20.32497","3":"7"},{"1":"671.90","2":"20.02457","3":"7"},{"1":"655.05","2":"24.00000","3":"7"},{"1":"645.25","2":"17.60784","3":"7"},{"1":"652.40","2":"19.34853","3":"7"},{"1":"634.00","2":"19.67846","3":"7"},{"1":"637.10","2":"18.72861","3":"7"},{"1":"639.75","2":"15.88235","3":"7"},{"1":"650.65","2":"20.05491","3":"7"},{"1":"659.40","2":"17.98825","3":"7"},{"1":"679.75","2":"16.96629","3":"7"},{"1":"662.55","2":"19.23937","3":"7"},{"1":"632.95","2":"19.19586","3":"7"},{"1":"655.05","2":"19.59906","3":"7"},{"1":"662.70","2":"20.54348","3":"7"},{"1":"657.65","2":"18.58848","3":"7"},{"1":"652.10","2":"15.60419","3":"7"},{"1":"665.10","2":"15.29304","3":"7"},{"1":"682.55","2":"17.65537","3":"7"},{"1":"695.30","2":"17.57976","3":"7"},{"1":"662.55","2":"22.33333","3":"7"},{"1":"663.50","2":"18.75000","3":"7"},{"1":"651.40","2":"18.10241","3":"7"},{"1":"635.05","2":"20.25641","3":"7"},{"1":"632.20","2":"18.80207","3":"7"},{"1":"664.15","2":"18.77230","3":"7"},{"1":"649.30","2":"20.40521","3":"7"},{"1":"669.30","2":"18.65079","3":"7"},{"1":"628.25","2":"20.70707","3":"7"},{"1":"639.35","2":"22.00000","3":"7"},{"1":"647.30","2":"17.69978","3":"7"},{"1":"640.75","2":"21.48329","3":"7"},{"1":"635.45","2":"16.70103","3":"7"},{"1":"673.05","2":"19.57567","3":"7"},{"1":"671.95","2":"17.25806","3":"7"},{"1":"638.30","2":"17.37526","3":"7"},{"1":"641.55","2":"17.34931","3":"7"},{"1":"635.60","2":"16.26229","3":"7"},{"1":"605.55","2":"17.70045","3":"7"},{"1":"643.60","2":"20.12881","3":"7"},{"1":"643.50","2":"18.26539","3":"7"},{"1":"625.85","2":"14.54214","3":"7"},{"1":"637.95","2":"19.15261","3":"7"},{"1":"645.10","2":"17.36574","3":"7"},{"1":"652.85","2":"15.13898","3":"7"},{"1":"677.95","2":"17.84266","3":"7"},{"1":"669.95","2":"15.40704","3":"7"},{"1":"634.15","2":"18.86534","3":"7"},{"1":"675.40","2":"16.47413","3":"7"},{"1":"660.05","2":"17.86263","3":"7"},{"1":"653.70","2":"21.88586","3":"7"},{"1":"628.65","2":"20.20000","3":"7"},{"1":"684.95","2":"19.03640","3":"7"},{"1":"644.45","2":"17.88991","3":"8"},{"1":"639.25","2":"21.52466","3":"8"},{"1":"616.30","2":"18.69723","3":"8"},{"1":"673.55","2":"17.35714","3":"8"},{"1":"645.75","2":"18.67133","3":"8"},{"1":"638.00","2":"21.40625","3":"8"},{"1":"636.70","2":"19.50000","3":"8"},{"1":"684.30","2":"20.89412","3":"8"},{"1":"664.70","2":"19.94737","3":"8"},{"1":"636.95","2":"20.80556","3":"8"},{"1":"671.90","2":"21.23809","3":"8"},{"1":"680.05","2":"21.00000","3":"8"},{"1":"667.45","2":"20.60000","3":"8"},{"1":"659.15","2":"20.00822","3":"8"},{"1":"644.20","2":"18.02778","3":"8"},{"1":"706.75","2":"20.25196","3":"8"},{"1":"666.15","2":"16.97787","3":"8"},{"1":"668.65","2":"16.50980","3":"8"},{"1":"654.85","2":"22.70402","3":"8"},{"1":"661.85","2":"19.91111","3":"8"},{"1":"651.35","2":"18.33333","3":"8"},{"1":"640.75","2":"22.61905","3":"8"},{"1":"634.05","2":"19.44828","3":"8"},{"1":"666.85","2":"25.05263","3":"8"},{"1":"612.65","2":"20.67544","3":"8"},{"1":"657.80","2":"18.68235","3":"8"},{"1":"647.70","2":"22.84553","3":"8"},{"1":"648.25","2":"19.26667","3":"8"},{"1":"670.70","2":"19.25000","3":"8"},{"1":"665.95","2":"20.54545","3":"8"},{"1":"638.30","2":"20.60697","3":"8"},{"1":"643.50","2":"21.07268","3":"8"},{"1":"655.75","2":"21.53581","3":"8"},{"1":"653.55","2":"19.90400","3":"8"},{"1":"640.15","2":"21.19407","3":"8"},{"1":"653.85","2":"21.86535","3":"8"},{"1":"650.90","2":"18.32965","3":"8"},{"1":"644.45","2":"16.22857","3":"8"},{"1":"641.10","2":"19.17857","3":"8"},{"1":"666.45","2":"20.27737","3":"8"},{"1":"641.55","2":"22.98614","3":"8"},{"1":"665.90","2":"20.44444","3":"8"},{"1":"637.10","2":"19.82085","3":"8"},{"1":"655.40","2":"23.20522","3":"8"},{"1":"639.75","2":"19.26697","3":"8"},{"1":"657.50","2":"23.30189","3":"8"},{"1":"646.50","2":"21.18829","3":"8"},{"1":"664.45","2":"20.87180","3":"8"},{"1":"644.20","2":"19.01749","3":"8"},{"1":"638.20","2":"21.91938","3":"8"},{"1":"656.40","2":"20.10124","3":"8"},{"1":"632.25","2":"21.47651","3":"8"},{"1":"664.95","2":"20.06579","3":"8"},{"1":"661.85","2":"20.37510","3":"8"},{"1":"645.75","2":"22.44648","3":"8"},{"1":"669.80","2":"22.89524","3":"8"},{"1":"620.50","2":"20.49797","3":"8"},{"1":"681.60","2":"20.00000","3":"8"},{"1":"694.25","2":"22.25658","3":"8"},{"1":"666.70","2":"21.56436","3":"8"},{"1":"641.80","2":"19.47737","3":"8"},{"1":"654.85","2":"17.67002","3":"8"},{"1":"668.95","2":"21.94756","3":"8"},{"1":"628.40","2":"21.78339","3":"8"},{"1":"635.45","2":"19.14000","3":"8"},{"1":"644.55","2":"18.11050","3":"8"},{"1":"662.90","2":"20.68242","3":"8"},{"1":"642.20","2":"22.62361","3":"8"},{"1":"652.85","2":"21.78650","3":"8"},{"1":"657.40","2":"18.58293","3":"8"},{"1":"659.80","2":"21.54545","3":"8"},{"1":"646.90","2":"21.15289","3":"8"},{"1":"617.35","2":"16.63333","3":"8"},{"1":"646.55","2":"21.14438","3":"8"},{"1":"653.70","2":"19.78182","3":"8"},{"1":"654.20","2":"18.98373","3":"8"},{"1":"651.80","2":"17.66767","3":"8"},{"1":"672.05","2":"17.75499","3":"8"},{"1":"661.35","2":"15.27273","3":"8"},{"1":"635.60","2":"14.00000","3":"8"},{"1":"696.55","2":"20.59613","3":"8"},{"1":"671.70","2":"16.31169","3":"8"},{"1":"641.45","2":"21.12796","3":"8"},{"1":"666.55","2":"17.48801","3":"8"},{"1":"636.50","2":"17.88679","3":"8"},{"1":"683.40","2":"19.30676","3":"8"},{"1":"699.10","2":"20.89231","3":"8"},{"1":"660.20","2":"21.28684","3":"8"},{"1":"633.05","2":"20.19560","3":"8"},{"1":"637.65","2":"24.95000","3":"8"},{"1":"666.85","2":"18.13043","3":"8"},{"1":"663.90","2":"20.00000","3":"8"},{"1":"673.05","2":"18.72951","3":"8"},{"1":"698.45","2":"18.25000","3":"8"},{"1":"676.60","2":"18.99257","3":"8"},{"1":"642.40","2":"19.88764","3":"8"},{"1":"626.90","2":"19.37895","3":"8"},{"1":"672.35","2":"20.46259","3":"8"},{"1":"620.30","2":"22.29157","3":"8"},{"1":"634.55","2":"20.70474","3":"8"},{"1":"639.85","2":"19.06005","3":"8"},{"1":"641.45","2":"20.23247","3":"8"},{"1":"637.00","2":"19.69012","3":"8"},{"1":"660.05","2":"20.36254","3":"8"},{"1":"675.70","2":"19.75422","3":"8"},{"1":"653.50","2":"19.37977","3":"8"},{"1":"664.30","2":"22.92351","3":"8"},{"1":"686.70","2":"19.37340","3":"8"},{"1":"656.80","2":"19.15516","3":"8"},{"1":"634.40","2":"21.30000","3":"8"},{"1":"648.20","2":"18.30357","3":"8"},{"1":"669.30","2":"21.07926","3":"8"},{"1":"698.20","2":"18.79121","3":"8"},{"1":"662.50","2":"19.62662","3":"8"},{"1":"661.60","2":"19.59016","3":"8"},{"1":"678.05","2":"20.87187","3":"8"},{"1":"643.70","2":"21.11500","3":"8"},{"1":"653.40","2":"20.08452","3":"8"},{"1":"658.35","2":"19.91049","3":"8"},{"1":"642.75","2":"17.81285","3":"8"},{"1":"643.40","2":"18.13333","3":"8"},{"1":"632.95","2":"19.22221","3":"8"},{"1":"651.85","2":"18.66072","3":"8"},{"1":"627.30","2":"19.60000","3":"8"},{"1":"673.30","2":"19.28384","3":"8"},{"1":"642.20","2":"22.81818","3":"8"},{"1":"648.00","2":"18.80922","3":"8"},{"1":"669.10","2":"21.37363","3":"8"},{"1":"691.90","2":"20.02041","3":"8"},{"1":"652.10","2":"21.49862","3":"8"},{"1":"671.75","2":"15.42857","3":"8"},{"1":"651.90","2":"22.40000","3":"8"},{"1":"637.95","2":"20.12709","3":"8"},{"1":"616.30","2":"19.03798","3":"8"},{"1":"647.05","2":"17.34216","3":"8"},{"1":"656.65","2":"17.01863","3":"8"},{"1":"680.45","2":"20.80000","3":"8"},{"1":"655.35","2":"21.15385","3":"8"},{"1":"660.10","2":"18.45833","3":"8"},{"1":"632.85","2":"19.14082","3":"8"},{"1":"665.20","2":"19.40766","3":"8"},{"1":"695.30","2":"19.56896","3":"8"},{"1":"663.35","2":"21.50120","3":"8"},{"1":"662.55","2":"17.52941","3":"8"},{"1":"678.80","2":"16.43017","3":"8"},{"1":"631.95","2":"19.79654","3":"8"},{"1":"635.05","2":"17.18613","3":"8"},{"1":"639.80","2":"17.61589","3":"8"},{"1":"618.05","2":"20.12537","3":"8"},{"1":"640.50","2":"22.16667","3":"8"},{"1":"612.50","2":"19.96154","3":"8"},{"1":"660.75","2":"19.03945","3":"8"},{"1":"665.10","2":"15.22436","3":"8"},{"1":"646.70","2":"21.14475","3":"8"},{"1":"650.65","2":"19.64390","3":"8"},{"1":"630.55","2":"21.04869","3":"8"},{"1":"631.40","2":"20.17544","3":"8"},{"1":"636.10","2":"21.39130","3":"8"},{"1":"655.20","2":"20.00833","3":"8"},{"1":"693.95","2":"20.29137","3":"8"},{"1":"673.90","2":"17.66667","3":"8"},{"1":"634.20","2":"18.22055","3":"8"},{"1":"646.40","2":"20.27100","3":"8"},{"1":"657.15","2":"20.19895","3":"8"},{"1":"645.25","2":"21.38424","3":"8"},{"1":"627.25","2":"20.97368","3":"8"},{"1":"669.30","2":"20.00000","3":"8"},{"1":"672.45","2":"17.15328","3":"8"},{"1":"654.60","2":"22.34977","3":"8"},{"1":"645.55","2":"22.17007","3":"8"},{"1":"616.45","2":"18.18182","3":"8"},{"1":"682.65","2":"18.95714","3":"8"},{"1":"638.55","2":"19.74533","3":"8"},{"1":"634.90","2":"16.42623","3":"8"},{"1":"666.00","2":"16.62540","3":"8"},{"1":"676.55","2":"16.38177","3":"8"},{"1":"666.65","2":"20.07416","3":"8"},{"1":"629.80","2":"17.99544","3":"8"},{"1":"679.40","2":"19.39130","3":"8"},{"1":"694.80","2":"16.42857","3":"8"},{"1":"661.60","2":"16.72949","3":"8"},{"1":"684.95","2":"24.41345","3":"8"},{"1":"615.75","2":"18.26415","3":"8"},{"1":"665.35","2":"18.95504","3":"8"},{"1":"643.05","2":"21.03896","3":"8"},{"1":"630.55","2":"20.74074","3":"8"},{"1":"635.60","2":"18.10000","3":"8"},{"1":"652.50","2":"19.84615","3":"8"},{"1":"677.95","2":"21.60000","3":"8"},{"1":"662.40","2":"22.44242","3":"8"},{"1":"634.10","2":"23.01438","3":"8"},{"1":"655.05","2":"17.74892","3":"8"},{"1":"635.75","2":"18.28664","3":"8"},{"1":"657.75","2":"19.26544","3":"8"},{"1":"657.65","2":"22.66667","3":"8"},{"1":"656.80","2":"19.29412","3":"8"},{"1":"653.80","2":"17.36364","3":"8"},{"1":"623.60","2":"19.82143","3":"8"},{"1":"683.35","2":"20.43378","3":"8"},{"1":"645.00","2":"21.03721","3":"8"},{"1":"645.10","2":"19.92462","3":"8"},{"1":"686.05","2":"19.00986","3":"8"},{"1":"660.95","2":"23.82222","3":"8"},{"1":"655.70","2":"19.36909","3":"8"},{"1":"663.45","2":"19.82857","3":"8"},{"1":"671.60","2":"15.25885","3":"8"},{"1":"628.55","2":"17.16129","3":"8"},{"1":"663.85","2":"21.81333","3":"8"},{"1":"700.30","2":"19.07471","3":"8"},{"1":"684.35","2":"25.78512","3":"8"},{"1":"661.20","2":"18.21261","3":"8"},{"1":"652.10","2":"18.16606","3":"8"},{"1":"635.20","2":"16.97297","3":"8"},{"1":"695.20","2":"21.50087","3":"8"},{"1":"619.80","2":"20.60000","3":"8"},{"1":"655.85","2":"16.99029","3":"8"},{"1":"679.65","2":"20.77954","3":"8"},{"1":"621.75","2":"15.51247","3":"8"},{"1":"668.80","2":"19.88506","3":"8"},{"1":"666.05","2":"21.39882","3":"8"},{"1":"649.70","2":"20.49751","3":"8"},{"1":"655.05","2":"19.36376","3":"8"},{"1":"636.90","2":"17.65957","3":"8"},{"1":"637.35","2":"21.01796","3":"8"},{"1":"655.35","2":"19.05565","3":"8"},{"1":"676.95","2":"22.53846","3":"8"},{"1":"666.15","2":"21.10787","3":"8"},{"1":"672.05","2":"20.05135","3":"8"},{"1":"632.00","2":"14.20176","3":"8"},{"1":"626.10","2":"18.47687","3":"8"},{"1":"664.15","2":"18.63542","3":"8"},{"1":"652.30","2":"20.94595","3":"8"},{"1":"640.10","2":"21.08548","3":"8"},{"1":"640.85","2":"18.69288","3":"8"},{"1":"638.30","2":"20.86808","3":"8"},{"1":"659.90","2":"19.82558","3":"8"},{"1":"643.50","2":"19.75000","3":"8"},{"1":"698.25","2":"19.50000","3":"8"},{"1":"624.15","2":"18.39080","3":"8"},{"1":"672.55","2":"18.78676","3":"8"},{"1":"657.00","2":"19.77018","3":"8"},{"1":"616.30","2":"19.33333","3":"8"},{"1":"648.95","2":"21.46392","3":"8"},{"1":"653.55","2":"23.08492","3":"8"},{"1":"644.95","2":"21.06299","3":"8"},{"1":"689.10","2":"18.68687","3":"8"},{"1":"658.00","2":"20.77024","3":"8"},{"1":"649.50","2":"19.30556","3":"8"},{"1":"671.65","2":"20.13280","3":"8"},{"1":"669.95","2":"20.66964","3":"8"},{"1":"659.35","2":"22.28155","3":"8"},{"1":"679.75","2":"20.60027","3":"8"},{"1":"651.45","2":"20.82734","3":"8"},{"1":"628.75","2":"19.22492","3":"8"},{"1":"645.55","2":"17.65477","3":"8"},{"1":"633.15","2":"17.00000","3":"8"},{"1":"636.60","2":"16.49773","3":"8"},{"1":"652.40","2":"19.78261","3":"8"},{"1":"630.35","2":"22.30216","3":"8"},{"1":"672.70","2":"17.73077","3":"8"},{"1":"661.90","2":"20.44836","3":"8"},{"1":"674.25","2":"20.37169","3":"8"},{"1":"643.70","2":"20.16479","3":"8"},{"1":"650.45","2":"21.61538","3":"8"},{"1":"623.10","2":"20.56143","3":"8"},{"1":"655.30","2":"19.95551","3":"8"},{"1":"656.70","2":"21.18387","3":"8"},{"1":"663.85","2":"18.81042","3":"8"},{"1":"643.60","2":"20.57838","3":"8"},{"1":"647.60","2":"18.32461","3":"8"},{"1":"643.20","2":"18.82063","3":"8"},{"1":"655.55","2":"20.81633","3":"8"},{"1":"646.95","2":"20.00000","3":"8"},{"1":"664.15","2":"19.68182","3":"8"},{"1":"677.25","2":"19.39018","3":"8"},{"1":"672.20","2":"20.92732","3":"8"},{"1":"666.65","2":"19.94437","3":"8"},{"1":"691.05","2":"20.79109","3":"8"},{"1":"681.30","2":"19.20354","3":"8"},{"1":"668.00","2":"19.02439","3":"8"},{"1":"667.15","2":"17.62058","3":"8"},{"1":"626.80","2":"20.23715","3":"8"},{"1":"656.50","2":"19.29374","3":"8"},{"1":"647.60","2":"18.82998","3":"8"},{"1":"638.35","2":"20.33949","3":"8"},{"1":"671.30","2":"19.22900","3":"8"},{"1":"658.80","2":"17.89130","3":"8"},{"1":"652.00","2":"19.51881","3":"8"},{"1":"635.95","2":"19.08451","3":"8"},{"1":"668.10","2":"19.93548","3":"8"},{"1":"679.50","2":"18.87326","3":"8"},{"1":"671.60","2":"20.14178","3":"8"},{"1":"647.30","2":"23.55637","3":"8"},{"1":"650.90","2":"21.46479","3":"8"},{"1":"634.10","2":"19.19101","3":"8"},{"1":"673.25","2":"20.13080","3":"8"},{"1":"659.05","2":"25.80000","3":"8"},{"1":"606.75","2":"18.77774","3":"8"},{"1":"657.00","2":"19.10982","3":"8"},{"1":"631.90","2":"19.70109","3":"8"},{"1":"653.95","2":"18.61594","3":"8"},{"1":"625.85","2":"20.99721","3":"8"},{"1":"661.85","2":"20.00000","3":"8"},{"1":"659.40","2":"20.98325","3":"8"},{"1":"691.35","2":"21.64262","3":"8"},{"1":"687.55","2":"20.02967","3":"8"},{"1":"646.35","2":"19.81140","3":"8"},{"1":"651.40","2":"18.00000","3":"8"},{"1":"671.25","2":"19.35811","3":"8"},{"1":"669.35","2":"20.17912","3":"8"},{"1":"639.30","2":"21.11986","3":"8"},{"1":"639.50","2":"23.38974","3":"8"},{"1":"625.30","2":"22.18182","3":"8"},{"1":"671.90","2":"19.94283","3":"8"},{"1":"634.95","2":"17.78826","3":"8"},{"1":"623.45","2":"14.70588","3":"8"},{"1":"682.55","2":"19.04077","3":"8"},{"1":"660.30","2":"20.89195","3":"8"},{"1":"667.45","2":"19.83851","3":"8"},{"1":"649.15","2":"19.52191","3":"8"},{"1":"643.25","2":"20.68622","3":"8"},{"1":"653.10","2":"18.18182","3":"8"},{"1":"639.90","2":"18.89224","3":"8"},{"1":"666.60","2":"24.88889","3":"8"},{"1":"657.90","2":"18.58064","3":"8"},{"1":"647.25","2":"18.04000","3":"8"},{"1":"632.20","2":"17.73399","3":"8"},{"1":"628.25","2":"21.45455","3":"8"},{"1":"644.40","2":"19.92343","3":"8"},{"1":"682.45","2":"20.33942","3":"8"},{"1":"652.35","2":"22.54608","3":"8"},{"1":"627.10","2":"21.10344","3":"8"},{"1":"621.40","2":"18.19743","3":"8"},{"1":"659.40","2":"20.10768","3":"8"},{"1":"646.00","2":"19.15984","3":"8"},{"1":"658.60","2":"19.54545","3":"8"},{"1":"664.00","2":"20.88889","3":"8"},{"1":"667.60","2":"18.39150","3":"8"},{"1":"667.20","2":"19.17990","3":"8"},{"1":"679.80","2":"19.39771","3":"8"},{"1":"661.45","2":"21.67827","3":"8"},{"1":"637.95","2":"19.28889","3":"8"},{"1":"684.80","2":"20.34927","3":"8"},{"1":"648.70","2":"20.96416","3":"8"},{"1":"664.40","2":"19.46039","3":"8"},{"1":"655.05","2":"19.28572","3":"8"},{"1":"632.45","2":"20.91979","3":"8"},{"1":"609.00","2":"20.90021","3":"8"},{"1":"668.90","2":"20.59575","3":"8"},{"1":"633.90","2":"19.37500","3":"8"},{"1":"649.30","2":"19.95122","3":"8"},{"1":"634.70","2":"18.84973","3":"8"},{"1":"704.30","2":"18.11787","3":"8"},{"1":"657.55","2":"19.18341","3":"8"},{"1":"654.30","2":"22.00000","3":"8"},{"1":"623.20","2":"21.58416","3":"8"},{"1":"668.40","2":"20.38889","3":"8"},{"1":"654.10","2":"16.29310","3":"8"},{"1":"661.95","2":"18.27778","3":"8"},{"1":"652.40","2":"19.37472","3":"8"},{"1":"670.00","2":"18.90909","3":"8"},{"1":"624.55","2":"16.40693","3":"8"},{"1":"666.10","2":"15.59140","3":"8"},{"1":"639.35","2":"18.70694","3":"8"},{"1":"650.60","2":"18.32985","3":"8"},{"1":"605.55","2":"17.90235","3":"8"},{"1":"661.90","2":"18.91157","3":"8"},{"1":"664.00","2":"20.32497","3":"8"},{"1":"662.55","2":"20.02457","3":"8"},{"1":"628.65","2":"24.00000","3":"8"},{"1":"649.85","2":"17.60784","3":"8"},{"1":"664.95","2":"19.34853","3":"8"},{"1":"646.20","2":"19.67846","3":"8"},{"1":"650.55","2":"18.72861","3":"8"},{"1":"652.30","2":"15.88235","3":"8"},{"1":"669.35","2":"20.05491","3":"8"},{"1":"640.90","2":"17.98825","3":"8"},{"1":"624.95","2":"16.96629","3":"8"},{"1":"678.40","2":"19.23937","3":"8"},{"1":"644.50","2":"19.19586","3":"8"},{"1":"681.90","2":"19.59906","3":"8"},{"1":"648.35","2":"20.54348","3":"8"},{"1":"651.20","2":"18.58848","3":"8"},{"1":"630.40","2":"15.60419","3":"8"},{"1":"644.70","2":"15.29304","3":"8"},{"1":"631.05","2":"17.65537","3":"8"},{"1":"645.60","2":"17.57976","3":"8"},{"1":"631.85","2":"22.33333","3":"8"},{"1":"671.95","2":"18.75000","3":"8"},{"1":"622.05","2":"18.10241","3":"8"},{"1":"662.75","2":"20.25641","3":"8"},{"1":"633.65","2":"18.80207","3":"8"},{"1":"662.70","2":"18.77230","3":"8"},{"1":"651.15","2":"20.40521","3":"8"},{"1":"672.30","2":"18.65079","3":"8"},{"1":"662.40","2":"20.70707","3":"8"},{"1":"656.55","2":"22.00000","3":"8"},{"1":"664.75","2":"17.69978","3":"8"},{"1":"662.45","2":"21.48329","3":"8"},{"1":"690.80","2":"16.70103","3":"8"},{"1":"668.60","2":"19.57567","3":"8"},{"1":"655.80","2":"17.25806","3":"8"},{"1":"654.20","2":"17.37526","3":"8"},{"1":"622.60","2":"17.34931","3":"8"},{"1":"638.70","2":"16.26229","3":"8"},{"1":"634.00","2":"17.70045","3":"8"},{"1":"634.15","2":"20.12881","3":"8"},{"1":"665.65","2":"18.26539","3":"8"},{"1":"662.65","2":"14.54214","3":"8"},{"1":"675.40","2":"19.15261","3":"8"},{"1":"673.55","2":"17.36574","3":"8"},{"1":"643.40","2":"15.13898","3":"8"},{"1":"654.90","2":"17.84266","3":"8"},{"1":"618.30","2":"15.40704","3":"8"},{"1":"663.50","2":"18.86534","3":"8"},{"1":"669.85","2":"16.47413","3":"8"},{"1":"676.85","2":"17.86263","3":"8"},{"1":"682.15","2":"21.88586","3":"8"},{"1":"676.15","2":"20.20000","3":"8"},{"1":"681.30","2":"19.03640","3":"8"},{"1":"667.15","2":"17.88991","3":"9"},{"1":"661.45","2":"21.52466","3":"9"},{"1":"645.75","2":"18.69723","3":"9"},{"1":"643.40","2":"17.35714","3":"9"},{"1":"654.60","2":"18.67133","3":"9"},{"1":"660.05","2":"21.40625","3":"9"},{"1":"624.95","2":"19.50000","3":"9"},{"1":"671.60","2":"20.89412","3":"9"},{"1":"634.40","2":"19.94737","3":"9"},{"1":"698.45","2":"20.80556","3":"9"},{"1":"675.40","2":"21.23809","3":"9"},{"1":"668.65","2":"21.00000","3":"9"},{"1":"638.30","2":"20.60000","3":"9"},{"1":"641.55","2":"20.00822","3":"9"},{"1":"635.20","2":"18.02778","3":"9"},{"1":"648.00","2":"20.25196","3":"9"},{"1":"669.35","2":"16.97787","3":"9"},{"1":"657.00","2":"16.50980","3":"9"},{"1":"654.20","2":"22.70402","3":"9"},{"1":"641.80","2":"19.91111","3":"9"},{"1":"639.30","2":"18.33333","3":"9"},{"1":"654.10","2":"22.61905","3":"9"},{"1":"634.95","2":"19.44828","3":"9"},{"1":"648.35","2":"25.05263","3":"9"},{"1":"668.80","2":"20.67544","3":"9"},{"1":"676.55","2":"18.68235","3":"9"},{"1":"668.10","2":"22.84553","3":"9"},{"1":"679.50","2":"19.26667","3":"9"},{"1":"668.40","2":"19.25000","3":"9"},{"1":"656.70","2":"20.54545","3":"9"},{"1":"635.60","2":"20.60697","3":"9"},{"1":"654.20","2":"21.07268","3":"9"},{"1":"653.40","2":"21.53581","3":"9"},{"1":"674.25","2":"19.90400","3":"9"},{"1":"642.75","2":"21.19407","3":"9"},{"1":"652.10","2":"21.86535","3":"9"},{"1":"627.25","2":"18.32965","3":"9"},{"1":"655.85","2":"16.22857","3":"9"},{"1":"652.00","2":"19.17857","3":"9"},{"1":"617.35","2":"20.27737","3":"9"},{"1":"675.70","2":"22.98614","3":"9"},{"1":"646.70","2":"20.44444","3":"9"},{"1":"658.80","2":"19.82085","3":"9"},{"1":"634.10","2":"23.20522","3":"9"},{"1":"651.80","2":"19.26697","3":"9"},{"1":"649.85","2":"23.30189","3":"9"},{"1":"676.85","2":"21.18829","3":"9"},{"1":"642.20","2":"20.87180","3":"9"},{"1":"659.15","2":"19.01749","3":"9"},{"1":"662.65","2":"21.91938","3":"9"},{"1":"694.25","2":"20.10124","3":"9"},{"1":"633.65","2":"21.47651","3":"9"},{"1":"676.15","2":"20.06579","3":"9"},{"1":"646.00","2":"20.37510","3":"9"},{"1":"637.95","2":"22.44648","3":"9"},{"1":"652.40","2":"22.89524","3":"9"},{"1":"624.55","2":"20.49797","3":"9"},{"1":"651.45","2":"20.00000","3":"9"},{"1":"644.20","2":"22.25658","3":"9"},{"1":"662.55","2":"21.56436","3":"9"},{"1":"643.60","2":"19.47737","3":"9"},{"1":"673.55","2":"17.67002","3":"9"},{"1":"623.45","2":"21.94756","3":"9"},{"1":"634.90","2":"21.78339","3":"9"},{"1":"672.70","2":"19.14000","3":"9"},{"1":"624.15","2":"18.11050","3":"9"},{"1":"619.80","2":"20.68242","3":"9"},{"1":"651.90","2":"22.62361","3":"9"},{"1":"647.05","2":"21.78650","3":"9"},{"1":"655.40","2":"18.58293","3":"9"},{"1":"691.05","2":"21.54545","3":"9"},{"1":"635.60","2":"21.15289","3":"9"},{"1":"673.30","2":"16.63333","3":"9"},{"1":"678.80","2":"21.14438","3":"9"},{"1":"648.95","2":"19.78182","3":"9"},{"1":"666.70","2":"18.98373","3":"9"},{"1":"655.35","2":"17.66767","3":"9"},{"1":"666.60","2":"17.75499","3":"9"},{"1":"639.50","2":"15.27273","3":"9"},{"1":"686.70","2":"14.00000","3":"9"},{"1":"690.80","2":"20.59613","3":"9"},{"1":"637.95","2":"16.31169","3":"9"},{"1":"623.10","2":"21.12796","3":"9"},{"1":"665.35","2":"17.48801","3":"9"},{"1":"648.25","2":"17.88679","3":"9"},{"1":"670.00","2":"19.30676","3":"9"},{"1":"655.70","2":"20.89231","3":"9"},{"1":"657.00","2":"21.28684","3":"9"},{"1":"642.40","2":"20.19560","3":"9"},{"1":"657.50","2":"24.95000","3":"9"},{"1":"664.00","2":"18.13043","3":"9"},{"1":"632.20","2":"20.00000","3":"9"},{"1":"659.40","2":"18.72951","3":"9"},{"1":"673.90","2":"18.25000","3":"9"},{"1":"665.65","2":"18.99257","3":"9"},{"1":"661.20","2":"19.88764","3":"9"},{"1":"618.30","2":"19.37895","3":"9"},{"1":"639.75","2":"20.46259","3":"9"},{"1":"699.10","2":"22.29157","3":"9"},{"1":"638.00","2":"20.70474","3":"9"},{"1":"636.10","2":"19.06005","3":"9"},{"1":"627.10","2":"20.23247","3":"9"},{"1":"664.75","2":"19.69012","3":"9"},{"1":"634.20","2":"20.36254","3":"9"},{"1":"634.05","2":"19.75422","3":"9"},{"1":"667.45","2":"19.37977","3":"9"},{"1":"657.80","2":"22.92351","3":"9"},{"1":"660.20","2":"19.37340","3":"9"},{"1":"647.60","2":"19.15516","3":"9"},{"1":"645.10","2":"21.30000","3":"9"},{"1":"620.30","2":"18.30357","3":"9"},{"1":"664.15","2":"21.07926","3":"9"},{"1":"639.25","2":"18.79121","3":"9"},{"1":"644.55","2":"19.62662","3":"9"},{"1":"662.40","2":"19.59016","3":"9"},{"1":"665.10","2":"20.87187","3":"9"},{"1":"673.25","2":"21.11500","3":"9"},{"1":"644.50","2":"20.08452","3":"9"},{"1":"686.05","2":"19.91049","3":"9"},{"1":"653.55","2":"17.81285","3":"9"},{"1":"645.60","2":"18.13333","3":"9"},{"1":"696.55","2":"19.22221","3":"9"},{"1":"654.85","2":"18.66072","3":"9"},{"1":"606.75","2":"19.60000","3":"9"},{"1":"632.00","2":"19.28384","3":"9"},{"1":"664.95","2":"22.81818","3":"9"},{"1":"616.30","2":"18.80922","3":"9"},{"1":"646.55","2":"21.37363","3":"9"},{"1":"681.90","2":"20.02041","3":"9"},{"1":"653.50","2":"21.49862","3":"9"},{"1":"643.05","2":"15.42857","3":"9"},{"1":"660.75","2":"22.40000","3":"9"},{"1":"656.65","2":"20.12709","3":"9"},{"1":"695.20","2":"19.03798","3":"9"},{"1":"647.60","2":"17.34216","3":"9"},{"1":"679.75","2":"17.01863","3":"9"},{"1":"657.90","2":"20.80000","3":"9"},{"1":"640.90","2":"21.15385","3":"9"},{"1":"680.05","2":"18.45833","3":"9"},{"1":"676.95","2":"19.14082","3":"9"},{"1":"671.75","2":"19.40766","3":"9"},{"1":"667.60","2":"19.56896","3":"9"},{"1":"633.05","2":"21.50120","3":"9"},{"1":"664.40","2":"17.52941","3":"9"},{"1":"704.30","2":"16.43017","3":"9"},{"1":"644.40","2":"19.79654","3":"9"},{"1":"662.90","2":"17.18613","3":"9"},{"1":"628.40","2":"17.61589","3":"9"},{"1":"640.50","2":"20.12537","3":"9"},{"1":"635.45","2":"22.16667","3":"9"},{"1":"656.55","2":"19.96154","3":"9"},{"1":"679.65","2":"19.03945","3":"9"},{"1":"695.30","2":"15.22436","3":"9"},{"1":"634.70","2":"21.14475","3":"9"},{"1":"612.65","2":"19.64390","3":"9"},{"1":"653.70","2":"21.04869","3":"9"},{"1":"630.40","2":"20.17544","3":"9"},{"1":"647.25","2":"21.39130","3":"9"},{"1":"645.55","2":"20.00833","3":"9"},{"1":"663.50","2":"20.29137","3":"9"},{"1":"615.75","2":"17.66667","3":"9"},{"1":"668.00","2":"18.22055","3":"9"},{"1":"616.30","2":"20.27100","3":"9"},{"1":"649.50","2":"20.19895","3":"9"},{"1":"669.30","2":"21.38424","3":"9"},{"1":"678.40","2":"20.97368","3":"9"},{"1":"643.50","2":"20.00000","3":"9"},{"1":"631.85","2":"17.15328","3":"9"},{"1":"672.30","2":"22.34977","3":"9"},{"1":"642.20","2":"22.17007","3":"9"},{"1":"634.10","2":"18.18182","3":"9"},{"1":"672.35","2":"18.95714","3":"9"},{"1":"655.35","2":"19.74533","3":"9"},{"1":"665.95","2":"16.42623","3":"9"},{"1":"684.95","2":"16.62540","3":"9"},{"1":"650.65","2":"16.38177","3":"9"},{"1":"663.85","2":"20.07416","3":"9"},{"1":"664.45","2":"17.99544","3":"9"},{"1":"659.05","2":"19.39130","3":"9"},{"1":"666.65","2":"16.42857","3":"9"},{"1":"683.35","2":"16.72949","3":"9"},{"1":"645.55","2":"24.41345","3":"9"},{"1":"632.25","2":"18.26415","3":"9"},{"1":"687.55","2":"18.95504","3":"9"},{"1":"638.30","2":"21.03896","3":"9"},{"1":"643.70","2":"20.74074","3":"9"},{"1":"660.95","2":"18.10000","3":"9"},{"1":"644.70","2":"19.84615","3":"9"},{"1":"646.95","2":"21.60000","3":"9"},{"1":"668.95","2":"22.44242","3":"9"},{"1":"681.30","2":"23.01438","3":"9"},{"1":"649.70","2":"17.74892","3":"9"},{"1":"646.20","2":"18.28664","3":"9"},{"1":"643.25","2":"19.26544","3":"9"},{"1":"671.95","2":"22.66667","3":"9"},{"1":"643.40","2":"19.29412","3":"9"},{"1":"673.05","2":"17.36364","3":"9"},{"1":"700.30","2":"19.82143","3":"9"},{"1":"652.35","2":"20.43378","3":"9"},{"1":"609.00","2":"21.03721","3":"9"},{"1":"636.95","2":"19.92462","3":"9"},{"1":"662.70","2":"19.00986","3":"9"},{"1":"671.90","2":"23.82222","3":"9"},{"1":"629.80","2":"19.36909","3":"9"},{"1":"667.45","2":"19.82857","3":"9"},{"1":"644.20","2":"15.25885","3":"9"},{"1":"645.00","2":"17.16129","3":"9"},{"1":"673.55","2":"21.81333","3":"9"},{"1":"656.50","2":"19.07471","3":"9"},{"1":"646.35","2":"25.78512","3":"9"},{"1":"633.90","2":"18.21261","3":"9"},{"1":"627.30","2":"18.16606","3":"9"},{"1":"645.75","2":"16.97297","3":"9"},{"1":"634.15","2":"21.50087","3":"9"},{"1":"631.95","2":"20.60000","3":"9"},{"1":"639.80","2":"16.99029","3":"9"},{"1":"628.65","2":"20.77954","3":"9"},{"1":"651.35","2":"15.51247","3":"9"},{"1":"663.45","2":"19.88506","3":"9"},{"1":"631.05","2":"21.39882","3":"9"},{"1":"628.25","2":"20.49751","3":"9"},{"1":"655.30","2":"19.36376","3":"9"},{"1":"664.15","2":"17.65957","3":"9"},{"1":"626.90","2":"21.01796","3":"9"},{"1":"641.10","2":"19.05565","3":"9"},{"1":"645.25","2":"22.53846","3":"9"},{"1":"654.90","2":"21.10787","3":"9"},{"1":"666.00","2":"20.05135","3":"9"},{"1":"632.45","2":"14.20176","3":"9"},{"1":"640.15","2":"18.47687","3":"9"},{"1":"635.95","2":"18.63542","3":"9"},{"1":"665.90","2":"20.94595","3":"9"},{"1":"646.40","2":"21.08548","3":"9"},{"1":"653.55","2":"18.69288","3":"9"},{"1":"668.60","2":"20.86808","3":"9"},{"1":"643.50","2":"19.82558","3":"9"},{"1":"666.15","2":"19.75000","3":"9"},{"1":"653.80","2":"19.50000","3":"9"},{"1":"658.60","2":"18.39080","3":"9"},{"1":"662.50","2":"18.78676","3":"9"},{"1":"605.55","2":"19.77018","3":"9"},{"1":"661.35","2":"19.33333","3":"9"},{"1":"681.60","2":"21.46392","3":"9"},{"1":"656.80","2":"23.08492","3":"9"},{"1":"636.50","2":"21.06299","3":"9"},{"1":"637.65","2":"18.68687","3":"9"},{"1":"666.15","2":"20.77024","3":"9"},{"1":"641.45","2":"19.30556","3":"9"},{"1":"657.15","2":"20.13280","3":"9"},{"1":"641.45","2":"20.66964","3":"9"},{"1":"623.20","2":"22.28155","3":"9"},{"1":"661.85","2":"20.60027","3":"9"},{"1":"637.35","2":"20.82734","3":"9"},{"1":"637.00","2":"19.22492","3":"9"},{"1":"662.45","2":"17.65477","3":"9"},{"1":"653.95","2":"17.00000","3":"9"},{"1":"649.15","2":"16.49773","3":"9"},{"1":"661.60","2":"19.78261","3":"9"},{"1":"669.85","2":"22.30216","3":"9"},{"1":"651.20","2":"17.73077","3":"9"},{"1":"650.45","2":"20.44836","3":"9"},{"1":"669.30","2":"20.37169","3":"9"},{"1":"652.10","2":"20.16479","3":"9"},{"1":"666.10","2":"21.61538","3":"9"},{"1":"663.35","2":"20.56143","3":"9"},{"1":"654.30","2":"19.95551","3":"9"},{"1":"691.35","2":"21.18387","3":"9"},{"1":"630.35","2":"18.81042","3":"9"},{"1":"655.20","2":"20.57838","3":"9"},{"1":"643.70","2":"18.32461","3":"9"},{"1":"657.55","2":"18.82063","3":"9"},{"1":"672.05","2":"20.81633","3":"9"},{"1":"650.60","2":"20.00000","3":"9"},{"1":"662.75","2":"19.68182","3":"9"},{"1":"658.00","2":"19.39018","3":"9"},{"1":"651.85","2":"20.92732","3":"9"},{"1":"625.30","2":"19.94437","3":"9"},{"1":"612.50","2":"20.79109","3":"9"},{"1":"657.40","2":"19.20354","3":"9"},{"1":"640.85","2":"19.02439","3":"9"},{"1":"628.55","2":"17.62058","3":"9"},{"1":"706.75","2":"20.23715","3":"9"},{"1":"647.70","2":"19.29374","3":"9"},{"1":"644.45","2":"18.82998","3":"9"},{"1":"663.90","2":"20.33949","3":"9"},{"1":"644.95","2":"19.22900","3":"9"},{"1":"684.35","2":"17.89130","3":"9"},{"1":"677.95","2":"19.51881","3":"9"},{"1":"643.20","2":"19.08451","3":"9"},{"1":"640.10","2":"19.93548","3":"9"},{"1":"650.55","2":"18.87326","3":"9"},{"1":"623.60","2":"20.14178","3":"9"},{"1":"680.45","2":"23.55637","3":"9"},{"1":"652.40","2":"21.46479","3":"9"},{"1":"650.90","2":"19.19101","3":"9"},{"1":"653.10","2":"20.13080","3":"9"},{"1":"655.05","2":"25.80000","3":"9"},{"1":"652.30","2":"18.77774","3":"9"},{"1":"635.05","2":"19.10982","3":"9"},{"1":"659.90","2":"19.70109","3":"9"},{"1":"651.40","2":"18.61594","3":"9"},{"1":"621.40","2":"20.99721","3":"9"},{"1":"636.90","2":"20.00000","3":"9"},{"1":"647.30","2":"20.98325","3":"9"},{"1":"638.20","2":"21.64262","3":"9"},{"1":"662.55","2":"20.02967","3":"9"},{"1":"682.15","2":"19.81140","3":"9"},{"1":"666.05","2":"18.00000","3":"9"},{"1":"646.50","2":"19.35811","3":"9"},{"1":"664.95","2":"20.17912","3":"9"},{"1":"684.80","2":"21.11986","3":"9"},{"1":"632.85","2":"23.38974","3":"9"},{"1":"666.85","2":"22.18182","3":"9"},{"1":"671.70","2":"19.94283","3":"9"},{"1":"661.90","2":"17.78826","3":"9"},{"1":"672.05","2":"14.70588","3":"9"},{"1":"652.30","2":"19.04077","3":"9"},{"1":"646.90","2":"20.89195","3":"9"},{"1":"659.40","2":"19.83851","3":"9"},{"1":"655.80","2":"19.52191","3":"9"},{"1":"671.25","2":"20.68622","3":"9"},{"1":"672.20","2":"18.18182","3":"9"},{"1":"669.95","2":"18.89224","3":"9"},{"1":"670.70","2":"24.88889","3":"9"},{"1":"684.30","2":"18.58064","3":"9"},{"1":"691.90","2":"18.04000","3":"9"},{"1":"622.05","2":"17.73399","3":"9"},{"1":"689.10","2":"21.45455","3":"9"},{"1":"636.60","2":"19.92343","3":"9"},{"1":"640.75","2":"20.33942","3":"9"},{"1":"621.75","2":"22.54608","3":"9"},{"1":"655.55","2":"21.10344","3":"9"},{"1":"634.55","2":"18.19743","3":"9"},{"1":"639.85","2":"20.10768","3":"9"},{"1":"632.95","2":"19.15984","3":"9"},{"1":"622.60","2":"19.54545","3":"9"},{"1":"672.55","2":"20.88889","3":"9"},{"1":"681.30","2":"18.39150","3":"9"},{"1":"616.45","2":"19.17990","3":"9"},{"1":"664.70","2":"19.39771","3":"9"},{"1":"671.60","2":"21.67827","3":"9"},{"1":"649.30","2":"19.28889","3":"9"},{"1":"668.90","2":"20.34927","3":"9"},{"1":"656.40","2":"20.96416","3":"9"},{"1":"698.25","2":"19.46039","3":"9"},{"1":"676.60","2":"19.28572","3":"9"},{"1":"631.90","2":"20.91979","3":"9"},{"1":"671.30","2":"20.90021","3":"9"},{"1":"659.80","2":"20.59575","3":"9"},{"1":"694.80","2":"19.37500","3":"9"},{"1":"626.80","2":"19.95122","3":"9"},{"1":"671.90","2":"18.84973","3":"9"},{"1":"634.00","2":"18.11787","3":"9"},{"1":"639.90","2":"19.18341","3":"9"},{"1":"630.55","2":"22.00000","3":"9"},{"1":"669.10","2":"21.58416","3":"9"},{"1":"651.15","2":"20.38889","3":"9"},{"1":"661.90","2":"16.29310","3":"9"},{"1":"661.85","2":"18.27778","3":"9"},{"1":"682.45","2":"19.37472","3":"9"},{"1":"666.85","2":"18.90909","3":"9"},{"1":"679.40","2":"16.40693","3":"9"},{"1":"666.65","2":"15.59140","3":"9"},{"1":"638.70","2":"18.70694","3":"9"},{"1":"671.65","2":"18.32985","3":"9"},{"1":"636.70","2":"17.90235","3":"9"},{"1":"657.75","2":"18.91157","3":"9"},{"1":"625.85","2":"20.32497","3":"9"},{"1":"665.20","2":"20.02457","3":"9"},{"1":"667.20","2":"24.00000","3":"9"},{"1":"660.10","2":"17.60784","3":"9"},{"1":"672.45","2":"19.34853","3":"9"},{"1":"654.85","2":"19.67846","3":"9"},{"1":"662.40","2":"18.72861","3":"9"},{"1":"669.80","2":"15.88235","3":"9"},{"1":"655.75","2":"20.05491","3":"9"},{"1":"693.95","2":"17.98825","3":"9"},{"1":"661.85","2":"16.96629","3":"9"},{"1":"638.55","2":"19.23937","3":"9"},{"1":"648.20","2":"19.19586","3":"9"},{"1":"631.40","2":"19.59906","3":"9"},{"1":"663.85","2":"20.54348","3":"9"},{"1":"655.05","2":"18.58848","3":"9"},{"1":"677.25","2":"15.60419","3":"9"},{"1":"616.30","2":"15.29304","3":"9"},{"1":"660.30","2":"17.65537","3":"9"},{"1":"650.90","2":"17.57976","3":"9"},{"1":"659.35","2":"22.33333","3":"9"},{"1":"620.50","2":"18.75000","3":"9"},{"1":"664.00","2":"18.10241","3":"9"},{"1":"666.55","2":"20.25641","3":"9"},{"1":"655.05","2":"18.80207","3":"9"},{"1":"679.80","2":"18.77230","3":"9"},{"1":"653.85","2":"20.40521","3":"9"},{"1":"683.40","2":"18.65079","3":"9"},{"1":"644.45","2":"20.70707","3":"9"},{"1":"618.05","2":"22.00000","3":"9"},{"1":"669.35","2":"17.69978","3":"9"},{"1":"666.45","2":"21.48329","3":"9"},{"1":"658.35","2":"16.70103","3":"9"},{"1":"628.75","2":"19.57567","3":"9"},{"1":"637.10","2":"17.25806","3":"9"},{"1":"678.05","2":"17.37526","3":"9"},{"1":"664.30","2":"17.34931","3":"9"},{"1":"661.60","2":"16.26229","3":"9"},{"1":"656.80","2":"17.70045","3":"9"},{"1":"657.65","2":"20.12881","3":"9"},{"1":"661.95","2":"18.26539","3":"9"},{"1":"638.35","2":"14.54214","3":"9"},{"1":"698.20","2":"19.15261","3":"9"},{"1":"652.50","2":"17.36574","3":"9"},{"1":"682.65","2":"15.13898","3":"9"},{"1":"630.55","2":"17.84266","3":"9"},{"1":"648.70","2":"15.40704","3":"9"},{"1":"633.15","2":"18.86534","3":"9"},{"1":"652.85","2":"16.47413","3":"9"},{"1":"682.55","2":"17.86263","3":"9"},{"1":"626.10","2":"21.88586","3":"9"},{"1":"639.35","2":"20.20000","3":"9"},{"1":"635.75","2":"19.03640","3":"9"},{"1":"666.70","2":"17.88991","3":"10"},{"1":"649.70","2":"21.52466","3":"10"},{"1":"684.80","2":"18.69723","3":"10"},{"1":"631.95","2":"17.35714","3":"10"},{"1":"662.45","2":"18.67133","3":"10"},{"1":"700.30","2":"21.40625","3":"10"},{"1":"643.05","2":"19.50000","3":"10"},{"1":"660.95","2":"20.89412","3":"10"},{"1":"632.25","2":"19.94737","3":"10"},{"1":"683.40","2":"20.80556","3":"10"},{"1":"665.35","2":"21.23809","3":"10"},{"1":"691.35","2":"21.00000","3":"10"},{"1":"636.60","2":"20.60000","3":"10"},{"1":"637.95","2":"20.00822","3":"10"},{"1":"664.40","2":"18.02778","3":"10"},{"1":"651.40","2":"20.25196","3":"10"},{"1":"672.45","2":"16.97787","3":"10"},{"1":"652.85","2":"16.50980","3":"10"},{"1":"679.75","2":"22.70402","3":"10"},{"1":"658.00","2":"19.91111","3":"10"},{"1":"662.55","2":"18.33333","3":"10"},{"1":"640.85","2":"22.61905","3":"10"},{"1":"628.40","2":"19.44828","3":"10"},{"1":"618.30","2":"25.05263","3":"10"},{"1":"643.50","2":"20.67544","3":"10"},{"1":"679.40","2":"18.68235","3":"10"},{"1":"662.50","2":"22.84553","3":"10"},{"1":"654.30","2":"19.26667","3":"10"},{"1":"636.50","2":"19.25000","3":"10"},{"1":"652.35","2":"20.54545","3":"10"},{"1":"668.65","2":"20.60697","3":"10"},{"1":"698.45","2":"21.07268","3":"10"},{"1":"666.45","2":"21.53581","3":"10"},{"1":"654.85","2":"19.90400","3":"10"},{"1":"650.90","2":"21.19407","3":"10"},{"1":"670.00","2":"21.86535","3":"10"},{"1":"656.70","2":"18.32965","3":"10"},{"1":"647.25","2":"16.22857","3":"10"},{"1":"654.20","2":"19.17857","3":"10"},{"1":"624.15","2":"20.27737","3":"10"},{"1":"656.55","2":"22.98614","3":"10"},{"1":"655.20","2":"20.44444","3":"10"},{"1":"616.30","2":"19.82085","3":"10"},{"1":"646.50","2":"23.20522","3":"10"},{"1":"665.65","2":"19.26697","3":"10"},{"1":"654.60","2":"23.30189","3":"10"},{"1":"636.95","2":"21.18829","3":"10"},{"1":"653.80","2":"20.87180","3":"10"},{"1":"651.90","2":"19.01749","3":"10"},{"1":"661.35","2":"21.91938","3":"10"},{"1":"647.05","2":"20.10124","3":"10"},{"1":"641.80","2":"21.47651","3":"10"},{"1":"643.20","2":"20.06579","3":"10"},{"1":"671.95","2":"20.37510","3":"10"},{"1":"626.80","2":"22.44648","3":"10"},{"1":"653.95","2":"22.89524","3":"10"},{"1":"622.60","2":"20.49797","3":"10"},{"1":"655.55","2":"20.00000","3":"10"},{"1":"652.40","2":"22.25658","3":"10"},{"1":"643.50","2":"21.56436","3":"10"},{"1":"645.75","2":"19.47737","3":"10"},{"1":"662.70","2":"17.67002","3":"10"},{"1":"678.40","2":"21.94756","3":"10"},{"1":"682.65","2":"21.78339","3":"10"},{"1":"674.25","2":"19.14000","3":"10"},{"1":"634.00","2":"18.11050","3":"10"},{"1":"638.35","2":"20.68242","3":"10"},{"1":"632.20","2":"22.62361","3":"10"},{"1":"668.90","2":"21.78650","3":"10"},{"1":"649.15","2":"18.58293","3":"10"},{"1":"616.45","2":"21.54545","3":"10"},{"1":"640.10","2":"21.15289","3":"10"},{"1":"640.75","2":"16.63333","3":"10"},{"1":"654.10","2":"21.14438","3":"10"},{"1":"637.00","2":"19.78182","3":"10"},{"1":"646.00","2":"18.98373","3":"10"},{"1":"676.60","2":"17.66767","3":"10"},{"1":"634.70","2":"17.75499","3":"10"},{"1":"637.35","2":"15.27273","3":"10"},{"1":"657.65","2":"14.00000","3":"10"},{"1":"666.15","2":"20.59613","3":"10"},{"1":"644.45","2":"16.31169","3":"10"},{"1":"696.55","2":"21.12796","3":"10"},{"1":"653.55","2":"17.48801","3":"10"},{"1":"669.35","2":"17.88679","3":"10"},{"1":"625.30","2":"19.30676","3":"10"},{"1":"656.50","2":"20.89231","3":"10"},{"1":"668.80","2":"21.28684","3":"10"},{"1":"625.85","2":"20.19560","3":"10"},{"1":"659.15","2":"24.95000","3":"10"},{"1":"655.05","2":"18.13043","3":"10"},{"1":"651.85","2":"20.00000","3":"10"},{"1":"639.25","2":"18.72951","3":"10"},{"1":"668.95","2":"18.25000","3":"10"},{"1":"612.50","2":"18.99257","3":"10"},{"1":"644.70","2":"19.88764","3":"10"},{"1":"664.15","2":"19.37895","3":"10"},{"1":"681.30","2":"20.46259","3":"10"},{"1":"634.90","2":"22.29157","3":"10"},{"1":"633.15","2":"20.70474","3":"10"},{"1":"652.10","2":"19.06005","3":"10"},{"1":"645.25","2":"20.23247","3":"10"},{"1":"687.55","2":"19.69012","3":"10"},{"1":"649.85","2":"20.36254","3":"10"},{"1":"671.25","2":"19.75422","3":"10"},{"1":"666.65","2":"19.37977","3":"10"},{"1":"638.55","2":"22.92351","3":"10"},{"1":"640.90","2":"19.37340","3":"10"},{"1":"676.55","2":"19.15516","3":"10"},{"1":"658.80","2":"21.30000","3":"10"},{"1":"662.75","2":"18.30357","3":"10"},{"1":"694.25","2":"21.07926","3":"10"},{"1":"663.35","2":"18.79121","3":"10"},{"1":"667.45","2":"19.62662","3":"10"},{"1":"645.00","2":"19.59016","3":"10"},{"1":"639.30","2":"20.87187","3":"10"},{"1":"673.05","2":"21.11500","3":"10"},{"1":"623.20","2":"20.08452","3":"10"},{"1":"652.40","2":"19.91049","3":"10"},{"1":"631.85","2":"17.81285","3":"10"},{"1":"634.10","2":"18.13333","3":"10"},{"1":"672.20","2":"19.22221","3":"10"},{"1":"655.75","2":"18.66072","3":"10"},{"1":"668.10","2":"19.60000","3":"10"},{"1":"641.55","2":"19.28384","3":"10"},{"1":"645.55","2":"22.81818","3":"10"},{"1":"643.40","2":"18.80922","3":"10"},{"1":"679.50","2":"21.37363","3":"10"},{"1":"665.20","2":"20.02041","3":"10"},{"1":"633.90","2":"21.49862","3":"10"},{"1":"632.00","2":"15.42857","3":"10"},{"1":"652.30","2":"22.40000","3":"10"},{"1":"635.60","2":"20.12709","3":"10"},{"1":"644.40","2":"19.03798","3":"10"},{"1":"647.60","2":"17.34216","3":"10"},{"1":"655.35","2":"17.01863","3":"10"},{"1":"676.15","2":"20.80000","3":"10"},{"1":"671.60","2":"21.15385","3":"10"},{"1":"669.80","2":"18.45833","3":"10"},{"1":"649.30","2":"19.14082","3":"10"},{"1":"655.05","2":"19.40766","3":"10"},{"1":"661.20","2":"19.56896","3":"10"},{"1":"659.40","2":"21.50120","3":"10"},{"1":"659.40","2":"17.52941","3":"10"},{"1":"656.80","2":"16.43017","3":"10"},{"1":"664.45","2":"19.79654","3":"10"},{"1":"655.80","2":"17.18613","3":"10"},{"1":"661.60","2":"17.61589","3":"10"},{"1":"651.80","2":"20.12537","3":"10"},{"1":"680.45","2":"22.16667","3":"10"},{"1":"635.05","2":"19.96154","3":"10"},{"1":"669.10","2":"19.03945","3":"10"},{"1":"623.60","2":"15.22436","3":"10"},{"1":"635.45","2":"21.14475","3":"10"},{"1":"650.45","2":"19.64390","3":"10"},{"1":"671.30","2":"21.04869","3":"10"},{"1":"683.35","2":"20.17544","3":"10"},{"1":"661.45","2":"21.39130","3":"10"},{"1":"637.65","2":"20.00833","3":"10"},{"1":"667.60","2":"20.29137","3":"10"},{"1":"630.55","2":"17.66667","3":"10"},{"1":"639.35","2":"18.22055","3":"10"},{"1":"628.25","2":"20.27100","3":"10"},{"1":"617.35","2":"20.19895","3":"10"},{"1":"672.05","2":"21.38424","3":"10"},{"1":"677.95","2":"20.97368","3":"10"},{"1":"673.25","2":"20.00000","3":"10"},{"1":"678.80","2":"17.15328","3":"10"},{"1":"639.50","2":"22.34977","3":"10"},{"1":"646.35","2":"22.17007","3":"10"},{"1":"686.70","2":"18.18182","3":"10"},{"1":"655.30","2":"18.95714","3":"10"},{"1":"656.80","2":"19.74533","3":"10"},{"1":"677.25","2":"16.42623","3":"10"},{"1":"653.70","2":"16.62540","3":"10"},{"1":"676.95","2":"16.38177","3":"10"},{"1":"640.50","2":"20.07416","3":"10"},{"1":"664.15","2":"17.99544","3":"10"},{"1":"682.15","2":"19.39130","3":"10"},{"1":"661.60","2":"16.42857","3":"10"},{"1":"669.30","2":"16.72949","3":"10"},{"1":"657.00","2":"24.41345","3":"10"},{"1":"663.90","2":"18.26415","3":"10"},{"1":"636.90","2":"18.95504","3":"10"},{"1":"616.30","2":"21.03896","3":"10"},{"1":"628.65","2":"20.74074","3":"10"},{"1":"666.05","2":"18.10000","3":"10"},{"1":"666.85","2":"19.84615","3":"10"},{"1":"664.95","2":"21.60000","3":"10"},{"1":"638.30","2":"22.44242","3":"10"},{"1":"634.05","2":"23.01438","3":"10"},{"1":"633.65","2":"17.74892","3":"10"},{"1":"648.20","2":"18.28664","3":"10"},{"1":"671.90","2":"19.26544","3":"10"},{"1":"675.70","2":"22.66667","3":"10"},{"1":"641.45","2":"19.29412","3":"10"},{"1":"616.30","2":"17.36364","3":"10"},{"1":"655.70","2":"19.82143","3":"10"},{"1":"642.20","2":"20.43378","3":"10"},{"1":"657.40","2":"21.03721","3":"10"},{"1":"661.95","2":"19.92462","3":"10"},{"1":"648.70","2":"19.00986","3":"10"},{"1":"646.40","2":"23.82222","3":"10"},{"1":"668.40","2":"19.36909","3":"10"},{"1":"671.90","2":"19.82857","3":"10"},{"1":"673.55","2":"15.25885","3":"10"},{"1":"682.55","2":"17.16129","3":"10"},{"1":"681.30","2":"21.81333","3":"10"},{"1":"660.20","2":"19.07471","3":"10"},{"1":"637.95","2":"25.78512","3":"10"},{"1":"654.85","2":"18.21261","3":"10"},{"1":"623.10","2":"18.16606","3":"10"},{"1":"669.35","2":"16.97297","3":"10"},{"1":"651.35","2":"21.50087","3":"10"},{"1":"642.75","2":"20.60000","3":"10"},{"1":"671.70","2":"16.99029","3":"10"},{"1":"660.75","2":"20.77954","3":"10"},{"1":"699.10","2":"15.51247","3":"10"},{"1":"698.25","2":"19.88506","3":"10"},{"1":"659.35","2":"21.39882","3":"10"},{"1":"672.30","2":"20.49751","3":"10"},{"1":"620.30","2":"19.36376","3":"10"},{"1":"662.55","2":"17.65957","3":"10"},{"1":"668.00","2":"21.01796","3":"10"},{"1":"623.45","2":"19.05565","3":"10"},{"1":"650.65","2":"22.53846","3":"10"},{"1":"628.55","2":"21.10787","3":"10"},{"1":"664.70","2":"20.05135","3":"10"},{"1":"671.75","2":"14.20176","3":"10"},{"1":"663.50","2":"18.47687","3":"10"},{"1":"671.65","2":"18.63542","3":"10"},{"1":"691.90","2":"20.94595","3":"10"},{"1":"682.45","2":"21.08548","3":"10"},{"1":"661.90","2":"18.69288","3":"10"},{"1":"666.65","2":"20.86808","3":"10"},{"1":"650.90","2":"19.82558","3":"10"},{"1":"671.60","2":"19.75000","3":"10"},{"1":"657.80","2":"19.50000","3":"10"},{"1":"641.45","2":"18.39080","3":"10"},{"1":"647.70","2":"18.78676","3":"10"},{"1":"693.95","2":"19.77018","3":"10"},{"1":"657.00","2":"19.33333","3":"10"},{"1":"644.95","2":"21.46392","3":"10"},{"1":"630.55","2":"23.08492","3":"10"},{"1":"684.35","2":"21.06299","3":"10"},{"1":"662.65","2":"18.68687","3":"10"},{"1":"684.95","2":"20.77024","3":"10"},{"1":"627.30","2":"19.30556","3":"10"},{"1":"621.75","2":"20.13280","3":"10"},{"1":"643.25","2":"20.66964","3":"10"},{"1":"644.50","2":"22.28155","3":"10"},{"1":"665.90","2":"20.60027","3":"10"},{"1":"624.95","2":"20.82734","3":"10"},{"1":"664.95","2":"19.22492","3":"10"},{"1":"653.55","2":"17.65477","3":"10"},{"1":"642.20","2":"17.00000","3":"10"},{"1":"653.85","2":"16.49773","3":"10"},{"1":"672.35","2":"19.78261","3":"10"},{"1":"666.00","2":"22.30216","3":"10"},{"1":"675.40","2":"17.73077","3":"10"},{"1":"630.35","2":"20.44836","3":"10"},{"1":"652.10","2":"20.37169","3":"10"},{"1":"695.20","2":"20.16479","3":"10"},{"1":"638.00","2":"21.61538","3":"10"},{"1":"645.55","2":"20.56143","3":"10"},{"1":"605.55","2":"19.95551","3":"10"},{"1":"645.75","2":"21.18387","3":"10"},{"1":"634.15","2":"18.81042","3":"10"},{"1":"678.05","2":"20.57838","3":"10"},{"1":"672.70","2":"18.32461","3":"10"},{"1":"639.90","2":"18.82063","3":"10"},{"1":"679.65","2":"20.81633","3":"10"},{"1":"657.75","2":"20.00000","3":"10"},{"1":"644.20","2":"19.68182","3":"10"},{"1":"651.15","2":"19.39018","3":"10"},{"1":"647.30","2":"20.92732","3":"10"},{"1":"652.00","2":"19.94437","3":"10"},{"1":"655.85","2":"20.79109","3":"10"},{"1":"643.60","2":"19.20354","3":"10"},{"1":"628.75","2":"19.02439","3":"10"},{"1":"642.40","2":"17.62058","3":"10"},{"1":"684.30","2":"20.23715","3":"10"},{"1":"665.95","2":"19.29374","3":"10"},{"1":"630.40","2":"18.82998","3":"10"},{"1":"639.85","2":"20.33949","3":"10"},{"1":"680.05","2":"19.22900","3":"10"},{"1":"634.40","2":"17.89130","3":"10"},{"1":"652.30","2":"19.51881","3":"10"},{"1":"640.15","2":"19.08451","3":"10"},{"1":"631.05","2":"19.93548","3":"10"},{"1":"651.20","2":"18.87326","3":"10"},{"1":"621.40","2":"20.14178","3":"10"},{"1":"655.05","2":"23.55637","3":"10"},{"1":"655.35","2":"21.46479","3":"10"},{"1":"650.55","2":"19.19101","3":"10"},{"1":"657.15","2":"20.13080","3":"10"},{"1":"663.85","2":"25.80000","3":"10"},{"1":"635.75","2":"18.77774","3":"10"},{"1":"652.50","2":"19.10982","3":"10"},{"1":"689.10","2":"19.70109","3":"10"},{"1":"649.50","2":"18.61594","3":"10"},{"1":"660.05","2":"20.99721","3":"10"},{"1":"636.10","2":"20.00000","3":"10"},{"1":"634.95","2":"20.98325","3":"10"},{"1":"665.10","2":"21.64262","3":"10"},{"1":"648.25","2":"20.02967","3":"10"},{"1":"670.70","2":"19.81140","3":"10"},{"1":"698.20","2":"18.00000","3":"10"},{"1":"650.60","2":"19.35811","3":"10"},{"1":"676.85","2":"20.17912","3":"10"},{"1":"662.40","2":"21.11986","3":"10"},{"1":"706.75","2":"23.38974","3":"10"},{"1":"626.90","2":"22.18182","3":"10"},{"1":"639.80","2":"19.94283","3":"10"},{"1":"661.90","2":"17.78826","3":"10"},{"1":"666.55","2":"14.70588","3":"10"},{"1":"633.05","2":"19.04077","3":"10"},{"1":"646.95","2":"20.89195","3":"10"},{"1":"668.60","2":"19.83851","3":"10"},{"1":"694.80","2":"19.52191","3":"10"},{"1":"631.90","2":"20.68622","3":"10"},{"1":"669.85","2":"18.18182","3":"10"},{"1":"632.45","2":"18.89224","3":"10"},{"1":"673.90","2":"24.88889","3":"10"},{"1":"644.45","2":"18.58064","3":"10"},{"1":"690.80","2":"18.04000","3":"10"},{"1":"661.85","2":"17.73399","3":"10"},{"1":"653.50","2":"21.45455","3":"10"},{"1":"635.20","2":"19.92343","3":"10"},{"1":"664.30","2":"20.33942","3":"10"},{"1":"662.90","2":"22.54608","3":"10"},{"1":"654.90","2":"21.10344","3":"10"},{"1":"669.30","2":"18.19743","3":"10"},{"1":"643.40","2":"20.10768","3":"10"},{"1":"673.30","2":"19.15984","3":"10"},{"1":"643.70","2":"19.54545","3":"10"},{"1":"673.55","2":"20.88889","3":"10"},{"1":"648.00","2":"18.39150","3":"10"},{"1":"648.35","2":"19.17990","3":"10"},{"1":"619.80","2":"19.39771","3":"10"},{"1":"661.85","2":"21.67827","3":"10"},{"1":"646.20","2":"19.28889","3":"10"},{"1":"664.00","2":"20.34927","3":"10"},{"1":"658.60","2":"20.96416","3":"10"},{"1":"704.30","2":"19.46039","3":"10"},{"1":"663.85","2":"19.28572","3":"10"},{"1":"639.75","2":"20.91979","3":"10"},{"1":"669.95","2":"20.90021","3":"10"},{"1":"657.55","2":"20.59575","3":"10"},{"1":"657.90","2":"19.37500","3":"10"},{"1":"626.10","2":"19.95122","3":"10"},{"1":"659.80","2":"18.84973","3":"10"},{"1":"612.65","2":"18.11787","3":"10"},{"1":"609.00","2":"19.18341","3":"10"},{"1":"663.45","2":"22.00000","3":"10"},{"1":"667.20","2":"21.58416","3":"10"},{"1":"618.05","2":"20.38889","3":"10"},{"1":"659.05","2":"16.29310","3":"10"},{"1":"661.85","2":"18.27778","3":"10"},{"1":"660.10","2":"19.37472","3":"10"},{"1":"615.75","2":"18.90909","3":"10"},{"1":"658.35","2":"16.40693","3":"10"},{"1":"681.60","2":"15.59140","3":"10"},{"1":"634.55","2":"18.70694","3":"10"},{"1":"634.20","2":"18.32985","3":"10"},{"1":"635.60","2":"17.90235","3":"10"},{"1":"672.55","2":"18.91157","3":"10"},{"1":"644.55","2":"20.32497","3":"10"},{"1":"657.50","2":"20.02457","3":"10"},{"1":"638.20","2":"24.00000","3":"10"},{"1":"645.60","2":"17.60784","3":"10"},{"1":"624.55","2":"19.34853","3":"10"},{"1":"659.90","2":"19.67846","3":"10"},{"1":"632.95","2":"18.72861","3":"10"},{"1":"637.10","2":"15.88235","3":"10"},{"1":"691.05","2":"20.05491","3":"10"},{"1":"655.40","2":"17.98825","3":"10"},{"1":"672.05","2":"16.96629","3":"10"},{"1":"634.10","2":"19.23937","3":"10"},{"1":"606.75","2":"19.19586","3":"10"},{"1":"666.10","2":"19.59906","3":"10"},{"1":"646.90","2":"20.54348","3":"10"},{"1":"645.10","2":"18.58848","3":"10"},{"1":"629.80","2":"15.60419","3":"10"},{"1":"635.95","2":"15.29304","3":"10"},{"1":"648.95","2":"17.65537","3":"10"},{"1":"664.00","2":"17.57976","3":"10"},{"1":"656.40","2":"22.33333","3":"10"},{"1":"620.50","2":"18.75000","3":"10"},{"1":"656.65","2":"18.10241","3":"10"},{"1":"631.40","2":"20.25641","3":"10"},{"1":"667.45","2":"18.80207","3":"10"},{"1":"653.10","2":"18.77230","3":"10"},{"1":"681.90","2":"20.40521","3":"10"},{"1":"686.05","2":"18.65079","3":"10"},{"1":"641.10","2":"20.70707","3":"10"},{"1":"654.20","2":"22.00000","3":"10"},{"1":"643.70","2":"17.69978","3":"10"},{"1":"632.85","2":"21.48329","3":"10"},{"1":"660.30","2":"16.70103","3":"10"},{"1":"646.70","2":"19.57567","3":"10"},{"1":"622.05","2":"17.25806","3":"10"},{"1":"666.60","2":"17.37526","3":"10"},{"1":"647.60","2":"17.34931","3":"10"},{"1":"662.40","2":"16.26229","3":"10"},{"1":"653.40","2":"17.70045","3":"10"},{"1":"664.75","2":"20.12881","3":"10"},{"1":"666.15","2":"18.26539","3":"10"},{"1":"679.80","2":"14.54214","3":"10"},{"1":"627.25","2":"19.15261","3":"10"},{"1":"638.70","2":"17.36574","3":"10"},{"1":"646.55","2":"15.13898","3":"10"},{"1":"636.70","2":"17.84266","3":"10"},{"1":"667.15","2":"15.40704","3":"10"},{"1":"644.20","2":"18.86534","3":"10"},{"1":"638.30","2":"16.47413","3":"10"},{"1":"651.45","2":"17.86263","3":"10"},{"1":"695.30","2":"21.88586","3":"10"},{"1":"666.85","2":"20.20000","3":"10"},{"1":"627.10","2":"19.03640","3":"10"},{"1":"659.90","2":"17.88991","3":"11"},{"1":"661.85","2":"21.52466","3":"11"},{"1":"640.50","2":"18.69723","3":"11"},{"1":"672.70","2":"17.35714","3":"11"},{"1":"639.35","2":"18.67133","3":"11"},{"1":"683.40","2":"21.40625","3":"11"},{"1":"657.15","2":"19.50000","3":"11"},{"1":"652.35","2":"20.89412","3":"11"},{"1":"635.60","2":"19.94737","3":"11"},{"1":"631.95","2":"20.80556","3":"11"},{"1":"673.55","2":"21.23809","3":"11"},{"1":"661.85","2":"21.00000","3":"11"},{"1":"666.55","2":"20.60000","3":"11"},{"1":"669.30","2":"20.00822","3":"11"},{"1":"649.15","2":"18.02778","3":"11"},{"1":"654.20","2":"20.25196","3":"11"},{"1":"651.20","2":"16.97787","3":"11"},{"1":"625.85","2":"16.50980","3":"11"},{"1":"650.45","2":"22.70402","3":"11"},{"1":"665.10","2":"19.91111","3":"11"},{"1":"660.30","2":"18.33333","3":"11"},{"1":"642.40","2":"22.61905","3":"11"},{"1":"637.65","2":"19.44828","3":"11"},{"1":"642.75","2":"25.05263","3":"11"},{"1":"661.35","2":"20.67544","3":"11"},{"1":"658.60","2":"18.68235","3":"11"},{"1":"653.40","2":"22.84553","3":"11"},{"1":"634.90","2":"19.26667","3":"11"},{"1":"665.35","2":"19.25000","3":"11"},{"1":"678.80","2":"20.54545","3":"11"},{"1":"629.80","2":"20.60697","3":"11"},{"1":"634.10","2":"21.07268","3":"11"},{"1":"640.75","2":"21.53581","3":"11"},{"1":"654.60","2":"19.90400","3":"11"},{"1":"657.80","2":"21.19407","3":"11"},{"1":"653.70","2":"21.86535","3":"11"},{"1":"624.55","2":"18.32965","3":"11"},{"1":"654.20","2":"16.22857","3":"11"},{"1":"636.95","2":"19.17857","3":"11"},{"1":"666.60","2":"20.27737","3":"11"},{"1":"667.60","2":"22.98614","3":"11"},{"1":"645.00","2":"20.44444","3":"11"},{"1":"640.15","2":"19.82085","3":"11"},{"1":"676.55","2":"23.20522","3":"11"},{"1":"635.95","2":"19.26697","3":"11"},{"1":"645.75","2":"23.30189","3":"11"},{"1":"671.90","2":"21.18829","3":"11"},{"1":"646.90","2":"20.87180","3":"11"},{"1":"647.60","2":"19.01749","3":"11"},{"1":"620.30","2":"21.91938","3":"11"},{"1":"637.35","2":"20.10124","3":"11"},{"1":"654.85","2":"21.47651","3":"11"},{"1":"641.55","2":"20.06579","3":"11"},{"1":"644.20","2":"20.37510","3":"11"},{"1":"648.20","2":"22.44648","3":"11"},{"1":"671.60","2":"22.89524","3":"11"},{"1":"621.40","2":"20.49797","3":"11"},{"1":"666.15","2":"20.00000","3":"11"},{"1":"628.75","2":"22.25658","3":"11"},{"1":"624.15","2":"21.56436","3":"11"},{"1":"658.80","2":"19.47737","3":"11"},{"1":"676.95","2":"17.67002","3":"11"},{"1":"647.05","2":"21.94756","3":"11"},{"1":"661.20","2":"21.78339","3":"11"},{"1":"672.45","2":"19.14000","3":"11"},{"1":"665.90","2":"18.11050","3":"11"},{"1":"668.95","2":"20.68242","3":"11"},{"1":"623.45","2":"22.62361","3":"11"},{"1":"669.95","2":"21.78650","3":"11"},{"1":"631.90","2":"18.58293","3":"11"},{"1":"661.60","2":"21.54545","3":"11"},{"1":"687.55","2":"21.15289","3":"11"},{"1":"645.60","2":"16.63333","3":"11"},{"1":"655.85","2":"21.14438","3":"11"},{"1":"648.25","2":"19.78182","3":"11"},{"1":"638.35","2":"18.98373","3":"11"},{"1":"652.85","2":"17.66767","3":"11"},{"1":"626.90","2":"17.75499","3":"11"},{"1":"631.05","2":"15.27273","3":"11"},{"1":"651.35","2":"14.00000","3":"11"},{"1":"653.55","2":"20.59613","3":"11"},{"1":"645.75","2":"16.31169","3":"11"},{"1":"653.80","2":"21.12796","3":"11"},{"1":"667.45","2":"17.48801","3":"11"},{"1":"644.95","2":"17.88679","3":"11"},{"1":"662.40","2":"19.30676","3":"11"},{"1":"663.45","2":"20.89231","3":"11"},{"1":"633.65","2":"21.28684","3":"11"},{"1":"695.30","2":"20.19560","3":"11"},{"1":"632.20","2":"24.95000","3":"11"},{"1":"630.35","2":"18.13043","3":"11"},{"1":"662.65","2":"20.00000","3":"11"},{"1":"638.30","2":"18.72951","3":"11"},{"1":"646.00","2":"18.25000","3":"11"},{"1":"679.50","2":"18.99257","3":"11"},{"1":"646.95","2":"19.88764","3":"11"},{"1":"638.30","2":"19.37895","3":"11"},{"1":"643.40","2":"20.46259","3":"11"},{"1":"682.15","2":"22.29157","3":"11"},{"1":"669.35","2":"20.70474","3":"11"},{"1":"661.85","2":"19.06005","3":"11"},{"1":"618.30","2":"20.23247","3":"11"},{"1":"632.85","2":"19.69012","3":"11"},{"1":"666.15","2":"20.36254","3":"11"},{"1":"694.80","2":"19.75422","3":"11"},{"1":"616.45","2":"19.37977","3":"11"},{"1":"664.15","2":"22.92351","3":"11"},{"1":"672.30","2":"19.37340","3":"11"},{"1":"694.25","2":"19.15516","3":"11"},{"1":"669.10","2":"21.30000","3":"11"},{"1":"664.40","2":"18.30357","3":"11"},{"1":"679.75","2":"21.07926","3":"11"},{"1":"652.00","2":"18.79121","3":"11"},{"1":"666.45","2":"19.62662","3":"11"},{"1":"616.30","2":"19.59016","3":"11"},{"1":"643.70","2":"20.87187","3":"11"},{"1":"657.90","2":"21.11500","3":"11"},{"1":"620.50","2":"20.08452","3":"11"},{"1":"666.65","2":"19.91049","3":"11"},{"1":"650.55","2":"17.81285","3":"11"},{"1":"627.30","2":"18.13333","3":"11"},{"1":"622.05","2":"19.22221","3":"11"},{"1":"634.70","2":"18.66072","3":"11"},{"1":"665.65","2":"19.60000","3":"11"},{"1":"642.20","2":"19.28384","3":"11"},{"1":"628.65","2":"22.81818","3":"11"},{"1":"659.40","2":"18.80922","3":"11"},{"1":"684.80","2":"21.37363","3":"11"},{"1":"666.85","2":"20.02041","3":"11"},{"1":"682.55","2":"21.49862","3":"11"},{"1":"653.85","2":"15.42857","3":"11"},{"1":"666.65","2":"22.40000","3":"11"},{"1":"659.05","2":"20.12709","3":"11"},{"1":"660.10","2":"19.03798","3":"11"},{"1":"661.95","2":"17.34216","3":"11"},{"1":"650.90","2":"17.01863","3":"11"},{"1":"652.30","2":"20.80000","3":"11"},{"1":"678.40","2":"21.15385","3":"11"},{"1":"630.55","2":"18.45833","3":"11"},{"1":"664.95","2":"19.14082","3":"11"},{"1":"648.35","2":"19.40766","3":"11"},{"1":"671.95","2":"19.56896","3":"11"},{"1":"644.70","2":"21.50120","3":"11"},{"1":"654.90","2":"17.52941","3":"11"},{"1":"636.50","2":"16.43017","3":"11"},{"1":"698.45","2":"19.79654","3":"11"},{"1":"655.40","2":"17.18613","3":"11"},{"1":"654.10","2":"17.61589","3":"11"},{"1":"649.30","2":"20.12537","3":"11"},{"1":"634.15","2":"22.16667","3":"11"},{"1":"677.25","2":"19.96154","3":"11"},{"1":"664.30","2":"19.03945","3":"11"},{"1":"630.40","2":"15.22436","3":"11"},{"1":"699.10","2":"21.14475","3":"11"},{"1":"657.00","2":"19.64390","3":"11"},{"1":"655.05","2":"21.04869","3":"11"},{"1":"668.65","2":"20.17544","3":"11"},{"1":"645.25","2":"21.39130","3":"11"},{"1":"666.00","2":"20.00833","3":"11"},{"1":"623.60","2":"20.29137","3":"11"},{"1":"654.30","2":"17.66667","3":"11"},{"1":"668.60","2":"18.22055","3":"11"},{"1":"637.10","2":"20.27100","3":"11"},{"1":"662.40","2":"20.19895","3":"11"},{"1":"638.00","2":"21.38424","3":"11"},{"1":"673.25","2":"20.97368","3":"11"},{"1":"664.00","2":"20.00000","3":"11"},{"1":"673.55","2":"17.15328","3":"11"},{"1":"686.70","2":"22.34977","3":"11"},{"1":"639.90","2":"22.17007","3":"11"},{"1":"637.00","2":"18.18182","3":"11"},{"1":"663.85","2":"18.95714","3":"11"},{"1":"675.40","2":"19.74533","3":"11"},{"1":"670.70","2":"16.42623","3":"11"},{"1":"648.95","2":"16.62540","3":"11"},{"1":"664.45","2":"16.38177","3":"11"},{"1":"674.25","2":"20.07416","3":"11"},{"1":"664.15","2":"17.99544","3":"11"},{"1":"663.90","2":"19.39130","3":"11"},{"1":"638.70","2":"16.42857","3":"11"},{"1":"680.45","2":"16.72949","3":"11"},{"1":"628.25","2":"24.41345","3":"11"},{"1":"630.55","2":"18.26415","3":"11"},{"1":"623.20","2":"18.95504","3":"11"},{"1":"649.50","2":"21.03896","3":"11"},{"1":"644.20","2":"20.74074","3":"11"},{"1":"643.40","2":"18.10000","3":"11"},{"1":"671.65","2":"19.84615","3":"11"},{"1":"637.95","2":"21.60000","3":"11"},{"1":"704.30","2":"22.44242","3":"11"},{"1":"695.20","2":"23.01438","3":"11"},{"1":"662.55","2":"17.74892","3":"11"},{"1":"657.65","2":"18.28664","3":"11"},{"1":"623.10","2":"19.26544","3":"11"},{"1":"639.75","2":"22.66667","3":"11"},{"1":"636.60","2":"19.29412","3":"11"},{"1":"618.05","2":"17.36364","3":"11"},{"1":"655.55","2":"19.82143","3":"11"},{"1":"653.10","2":"20.43378","3":"11"},{"1":"645.10","2":"21.03721","3":"11"},{"1":"669.80","2":"19.92462","3":"11"},{"1":"671.60","2":"19.00986","3":"11"},{"1":"696.55","2":"23.82222","3":"11"},{"1":"650.90","2":"19.36909","3":"11"},{"1":"672.05","2":"19.82857","3":"11"},{"1":"664.70","2":"15.25885","3":"11"},{"1":"654.85","2":"17.16129","3":"11"},{"1":"634.40","2":"21.81333","3":"11"},{"1":"641.45","2":"19.07471","3":"11"},{"1":"632.95","2":"25.78512","3":"11"},{"1":"643.60","2":"18.21261","3":"11"},{"1":"636.10","2":"18.16606","3":"11"},{"1":"678.05","2":"16.97297","3":"11"},{"1":"670.00","2":"21.50087","3":"11"},{"1":"672.05","2":"20.60000","3":"11"},{"1":"665.20","2":"16.99029","3":"11"},{"1":"632.00","2":"20.77954","3":"11"},{"1":"634.05","2":"15.51247","3":"11"},{"1":"634.55","2":"19.88506","3":"11"},{"1":"638.20","2":"21.39882","3":"11"},{"1":"680.05","2":"20.49751","3":"11"},{"1":"605.55","2":"19.36376","3":"11"},{"1":"635.20","2":"17.65957","3":"11"},{"1":"655.80","2":"21.01796","3":"11"},{"1":"691.90","2":"19.05565","3":"11"},{"1":"649.85","2":"22.53846","3":"11"},{"1":"644.45","2":"21.10787","3":"11"},{"1":"659.35","2":"20.05135","3":"11"},{"1":"641.10","2":"14.20176","3":"11"},{"1":"609.00","2":"18.47687","3":"11"},{"1":"652.10","2":"18.63542","3":"11"},{"1":"616.30","2":"20.94595","3":"11"},{"1":"646.20","2":"21.08548","3":"11"},{"1":"655.35","2":"18.69288","3":"11"},{"1":"655.05","2":"20.86808","3":"11"},{"1":"649.70","2":"19.82558","3":"11"},{"1":"615.75","2":"19.75000","3":"11"},{"1":"657.50","2":"19.50000","3":"11"},{"1":"625.30","2":"18.39080","3":"11"},{"1":"668.90","2":"18.78676","3":"11"},{"1":"676.15","2":"19.77018","3":"11"},{"1":"693.95","2":"19.33333","3":"11"},{"1":"617.35","2":"21.46392","3":"11"},{"1":"639.30","2":"23.08492","3":"11"},{"1":"646.35","2":"21.06299","3":"11"},{"1":"690.80","2":"18.68687","3":"11"},{"1":"641.80","2":"20.77024","3":"11"},{"1":"656.80","2":"19.30556","3":"11"},{"1":"656.50","2":"20.13280","3":"11"},{"1":"653.95","2":"20.66964","3":"11"},{"1":"683.35","2":"22.28155","3":"11"},{"1":"653.50","2":"20.60027","3":"11"},{"1":"656.80","2":"20.82734","3":"11"},{"1":"622.60","2":"19.22492","3":"11"},{"1":"641.45","2":"17.65477","3":"11"},{"1":"638.55","2":"17.00000","3":"11"},{"1":"668.10","2":"16.49773","3":"11"},{"1":"643.70","2":"19.78261","3":"11"},{"1":"672.35","2":"22.30216","3":"11"},{"1":"663.85","2":"17.73077","3":"11"},{"1":"684.95","2":"20.44836","3":"11"},{"1":"684.35","2":"20.37169","3":"11"},{"1":"635.60","2":"20.16479","3":"11"},{"1":"637.95","2":"21.61538","3":"11"},{"1":"666.05","2":"20.56143","3":"11"},{"1":"634.95","2":"19.95551","3":"11"},{"1":"648.70","2":"21.18387","3":"11"},{"1":"660.20","2":"18.81042","3":"11"},{"1":"647.70","2":"20.57838","3":"11"},{"1":"634.20","2":"18.32461","3":"11"},{"1":"668.40","2":"18.82063","3":"11"},{"1":"669.30","2":"20.81633","3":"11"},{"1":"676.60","2":"20.00000","3":"11"},{"1":"662.50","2":"19.68182","3":"11"},{"1":"656.55","2":"19.39018","3":"11"},{"1":"643.50","2":"20.92732","3":"11"},{"1":"667.20","2":"19.94437","3":"11"},{"1":"644.50","2":"20.79109","3":"11"},{"1":"681.30","2":"19.20354","3":"11"},{"1":"635.05","2":"19.02439","3":"11"},{"1":"669.85","2":"17.62058","3":"11"},{"1":"661.90","2":"20.23715","3":"11"},{"1":"689.10","2":"19.29374","3":"11"},{"1":"652.10","2":"18.82998","3":"11"},{"1":"626.10","2":"20.33949","3":"11"},{"1":"646.40","2":"19.22900","3":"11"},{"1":"671.25","2":"17.89130","3":"11"},{"1":"679.80","2":"19.51881","3":"11"},{"1":"663.35","2":"19.08451","3":"11"},{"1":"675.70","2":"19.93548","3":"11"},{"1":"655.05","2":"18.87326","3":"11"},{"1":"671.90","2":"20.14178","3":"11"},{"1":"681.30","2":"23.55637","3":"11"},{"1":"643.25","2":"21.46479","3":"11"},{"1":"671.70","2":"19.19101","3":"11"},{"1":"681.60","2":"20.13080","3":"11"},{"1":"661.90","2":"25.80000","3":"11"},{"1":"639.50","2":"18.77774","3":"11"},{"1":"651.80","2":"19.10982","3":"11"},{"1":"666.70","2":"19.70109","3":"11"},{"1":"643.50","2":"18.61594","3":"11"},{"1":"676.85","2":"20.99721","3":"11"},{"1":"664.00","2":"20.00000","3":"11"},{"1":"658.00","2":"20.98325","3":"11"},{"1":"651.90","2":"21.64262","3":"11"},{"1":"653.55","2":"20.02967","3":"11"},{"1":"661.60","2":"19.81140","3":"11"},{"1":"646.55","2":"18.00000","3":"11"},{"1":"632.25","2":"19.35811","3":"11"},{"1":"660.75","2":"20.17912","3":"11"},{"1":"652.40","2":"21.11986","3":"11"},{"1":"646.70","2":"23.38974","3":"11"},{"1":"679.65","2":"22.18182","3":"11"},{"1":"668.00","2":"19.94283","3":"11"},{"1":"660.95","2":"17.78826","3":"11"},{"1":"616.30","2":"14.70588","3":"11"},{"1":"698.25","2":"19.04077","3":"11"},{"1":"639.85","2":"20.89195","3":"11"},{"1":"662.70","2":"19.83851","3":"11"},{"1":"645.55","2":"19.52191","3":"11"},{"1":"651.85","2":"20.68622","3":"11"},{"1":"633.90","2":"18.18182","3":"11"},{"1":"659.40","2":"18.89224","3":"11"},{"1":"682.45","2":"24.88889","3":"11"},{"1":"628.40","2":"18.58064","3":"11"},{"1":"606.75","2":"18.04000","3":"11"},{"1":"657.40","2":"17.73399","3":"11"},{"1":"642.20","2":"21.45455","3":"11"},{"1":"644.40","2":"19.92343","3":"11"},{"1":"652.30","2":"20.33942","3":"11"},{"1":"643.20","2":"22.54608","3":"11"},{"1":"633.15","2":"21.10344","3":"11"},{"1":"655.35","2":"18.19743","3":"11"},{"1":"669.35","2":"20.10768","3":"11"},{"1":"612.65","2":"19.15984","3":"11"},{"1":"659.80","2":"19.54545","3":"11"},{"1":"657.75","2":"20.88889","3":"11"},{"1":"643.05","2":"18.39150","3":"11"},{"1":"636.70","2":"19.17990","3":"11"},{"1":"655.70","2":"19.39771","3":"11"},{"1":"665.95","2":"21.67827","3":"11"},{"1":"647.30","2":"19.28889","3":"11"},{"1":"635.75","2":"20.34927","3":"11"},{"1":"666.85","2":"20.96416","3":"11"},{"1":"627.10","2":"19.46039","3":"11"},{"1":"650.65","2":"19.28572","3":"11"},{"1":"671.75","2":"20.91979","3":"11"},{"1":"646.50","2":"20.90021","3":"11"},{"1":"656.70","2":"20.59575","3":"11"},{"1":"679.40","2":"19.37500","3":"11"},{"1":"640.10","2":"19.95122","3":"11"},{"1":"664.75","2":"18.84973","3":"11"},{"1":"656.40","2":"18.11787","3":"11"},{"1":"657.00","2":"19.18341","3":"11"},{"1":"660.05","2":"22.00000","3":"11"},{"1":"656.65","2":"21.58416","3":"11"},{"1":"644.45","2":"20.38889","3":"11"},{"1":"700.30","2":"16.29310","3":"11"},{"1":"661.45","2":"18.27778","3":"11"},{"1":"682.65","2":"19.37472","3":"11"},{"1":"662.75","2":"18.90909","3":"11"},{"1":"673.90","2":"16.40693","3":"11"},{"1":"671.30","2":"15.59140","3":"11"},{"1":"621.75","2":"18.70694","3":"11"},{"1":"652.40","2":"18.32985","3":"11"},{"1":"631.40","2":"17.90235","3":"11"},{"1":"672.20","2":"18.91157","3":"11"},{"1":"650.60","2":"20.32497","3":"11"},{"1":"647.25","2":"20.02457","3":"11"},{"1":"667.45","2":"24.00000","3":"11"},{"1":"691.05","2":"17.60784","3":"11"},{"1":"684.30","2":"19.34853","3":"11"},{"1":"627.25","2":"19.67846","3":"11"},{"1":"639.25","2":"18.72861","3":"11"},{"1":"681.90","2":"15.88235","3":"11"},{"1":"655.75","2":"20.05491","3":"11"},{"1":"658.35","2":"17.98825","3":"11"},{"1":"644.55","2":"16.96629","3":"11"},{"1":"640.85","2":"19.23937","3":"11"},{"1":"691.35","2":"19.19586","3":"11"},{"1":"648.00","2":"19.59906","3":"11"},{"1":"645.55","2":"20.54348","3":"11"},{"1":"667.15","2":"18.58848","3":"11"},{"1":"628.55","2":"15.60419","3":"11"},{"1":"647.60","2":"15.29304","3":"11"},{"1":"677.95","2":"17.65537","3":"11"},{"1":"640.90","2":"17.57976","3":"11"},{"1":"632.45","2":"22.33333","3":"11"},{"1":"634.00","2":"18.75000","3":"11"},{"1":"664.95","2":"18.10241","3":"11"},{"1":"698.20","2":"20.25641","3":"11"},{"1":"639.80","2":"18.80207","3":"11"},{"1":"651.45","2":"18.77230","3":"11"},{"1":"626.80","2":"20.40521","3":"11"},{"1":"666.10","2":"18.65079","3":"11"},{"1":"655.20","2":"20.70707","3":"11"},{"1":"686.05","2":"22.00000","3":"11"},{"1":"657.55","2":"17.69978","3":"11"},{"1":"635.45","2":"21.48329","3":"11"},{"1":"655.30","2":"16.70103","3":"11"},{"1":"634.10","2":"19.57567","3":"11"},{"1":"612.50","2":"17.25806","3":"11"},{"1":"651.40","2":"17.37526","3":"11"},{"1":"659.15","2":"17.34931","3":"11"},{"1":"668.80","2":"16.26229","3":"11"},{"1":"673.05","2":"17.70045","3":"11"},{"1":"633.05","2":"20.12881","3":"11"},{"1":"624.95","2":"18.26539","3":"11"},{"1":"652.50","2":"14.54214","3":"11"},{"1":"662.55","2":"19.15261","3":"11"},{"1":"662.45","2":"17.36574","3":"11"},{"1":"619.80","2":"15.13898","3":"11"},{"1":"662.90","2":"17.84266","3":"11"},{"1":"636.90","2":"15.40704","3":"11"},{"1":"631.85","2":"18.86534","3":"11"},{"1":"673.30","2":"16.47413","3":"11"},{"1":"663.50","2":"17.86263","3":"11"},{"1":"706.75","2":"21.88586","3":"11"},{"1":"651.15","2":"20.20000","3":"11"},{"1":"672.55","2":"19.03640","3":"11"},{"1":"646.95","2":"17.88991","3":"12"},{"1":"616.45","2":"21.52466","3":"12"},{"1":"651.90","2":"18.69723","3":"12"},{"1":"690.80","2":"17.35714","3":"12"},{"1":"668.65","2":"18.67133","3":"12"},{"1":"662.45","2":"21.40625","3":"12"},{"1":"632.95","2":"19.50000","3":"12"},{"1":"643.50","2":"20.89412","3":"12"},{"1":"674.25","2":"19.94737","3":"12"},{"1":"657.50","2":"20.80556","3":"12"},{"1":"634.10","2":"21.23809","3":"12"},{"1":"657.00","2":"21.00000","3":"12"},{"1":"646.40","2":"20.60000","3":"12"},{"1":"645.10","2":"20.00822","3":"12"},{"1":"637.65","2":"18.02778","3":"12"},{"1":"666.60","2":"20.25196","3":"12"},{"1":"662.75","2":"16.97787","3":"12"},{"1":"657.00","2":"16.50980","3":"12"},{"1":"634.90","2":"22.70402","3":"12"},{"1":"676.60","2":"19.91111","3":"12"},{"1":"686.05","2":"18.33333","3":"12"},{"1":"669.30","2":"22.61905","3":"12"},{"1":"655.20","2":"19.44828","3":"12"},{"1":"620.30","2":"25.05263","3":"12"},{"1":"653.10","2":"20.67544","3":"12"},{"1":"652.85","2":"18.68235","3":"12"},{"1":"635.95","2":"22.84553","3":"12"},{"1":"650.90","2":"19.26667","3":"12"},{"1":"649.85","2":"19.25000","3":"12"},{"1":"628.25","2":"20.54545","3":"12"},{"1":"653.95","2":"20.60697","3":"12"},{"1":"671.60","2":"21.07268","3":"12"},{"1":"631.85","2":"21.53581","3":"12"},{"1":"661.90","2":"19.90400","3":"12"},{"1":"678.80","2":"21.19407","3":"12"},{"1":"681.90","2":"21.86535","3":"12"},{"1":"632.00","2":"18.32965","3":"12"},{"1":"672.05","2":"16.22857","3":"12"},{"1":"641.55","2":"19.17857","3":"12"},{"1":"659.40","2":"20.27737","3":"12"},{"1":"647.30","2":"22.98614","3":"12"},{"1":"644.55","2":"20.44444","3":"12"},{"1":"635.20","2":"19.82085","3":"12"},{"1":"649.70","2":"23.20522","3":"12"},{"1":"668.95","2":"19.26697","3":"12"},{"1":"620.50","2":"23.30189","3":"12"},{"1":"680.05","2":"21.18829","3":"12"},{"1":"643.70","2":"20.87180","3":"12"},{"1":"616.30","2":"19.01749","3":"12"},{"1":"653.40","2":"21.91938","3":"12"},{"1":"642.75","2":"20.10124","3":"12"},{"1":"627.25","2":"21.47651","3":"12"},{"1":"657.55","2":"20.06579","3":"12"},{"1":"664.15","2":"20.37510","3":"12"},{"1":"684.95","2":"22.44648","3":"12"},{"1":"645.00","2":"22.89524","3":"12"},{"1":"634.70","2":"20.49797","3":"12"},{"1":"669.10","2":"20.00000","3":"12"},{"1":"645.25","2":"22.25658","3":"12"},{"1":"650.45","2":"21.56436","3":"12"},{"1":"682.65","2":"19.47737","3":"12"},{"1":"683.40","2":"17.67002","3":"12"},{"1":"638.00","2":"21.94756","3":"12"},{"1":"627.10","2":"21.78339","3":"12"},{"1":"653.55","2":"19.14000","3":"12"},{"1":"612.65","2":"18.11050","3":"12"},{"1":"634.10","2":"20.68242","3":"12"},{"1":"647.60","2":"22.62361","3":"12"},{"1":"643.40","2":"21.78650","3":"12"},{"1":"666.15","2":"18.58293","3":"12"},{"1":"655.85","2":"21.54545","3":"12"},{"1":"631.05","2":"21.15289","3":"12"},{"1":"668.00","2":"16.63333","3":"12"},{"1":"664.95","2":"21.14438","3":"12"},{"1":"689.10","2":"19.78182","3":"12"},{"1":"657.75","2":"18.98373","3":"12"},{"1":"671.60","2":"17.66767","3":"12"},{"1":"655.30","2":"17.75499","3":"12"},{"1":"639.35","2":"15.27273","3":"12"},{"1":"623.45","2":"14.00000","3":"12"},{"1":"641.10","2":"20.59613","3":"12"},{"1":"644.95","2":"16.31169","3":"12"},{"1":"645.60","2":"21.12796","3":"12"},{"1":"636.95","2":"17.48801","3":"12"},{"1":"643.25","2":"17.88679","3":"12"},{"1":"706.75","2":"19.30676","3":"12"},{"1":"652.30","2":"20.89231","3":"12"},{"1":"635.75","2":"21.28684","3":"12"},{"1":"679.65","2":"20.19560","3":"12"},{"1":"609.00","2":"24.95000","3":"12"},{"1":"666.85","2":"18.13043","3":"12"},{"1":"655.35","2":"20.00000","3":"12"},{"1":"655.70","2":"18.72951","3":"12"},{"1":"669.95","2":"18.25000","3":"12"},{"1":"682.45","2":"18.99257","3":"12"},{"1":"646.50","2":"19.88764","3":"12"},{"1":"667.20","2":"19.37895","3":"12"},{"1":"652.50","2":"20.46259","3":"12"},{"1":"651.35","2":"22.29157","3":"12"},{"1":"672.35","2":"20.70474","3":"12"},{"1":"619.80","2":"19.06005","3":"12"},{"1":"653.80","2":"20.23247","3":"12"},{"1":"656.70","2":"19.69012","3":"12"},{"1":"648.20","2":"20.36254","3":"12"},{"1":"646.55","2":"19.75422","3":"12"},{"1":"642.20","2":"19.37977","3":"12"},{"1":"681.60","2":"22.92351","3":"12"},{"1":"645.75","2":"19.37340","3":"12"},{"1":"673.55","2":"19.15516","3":"12"},{"1":"652.30","2":"21.30000","3":"12"},{"1":"637.00","2":"18.30357","3":"12"},{"1":"649.50","2":"21.07926","3":"12"},{"1":"661.90","2":"18.79121","3":"12"},{"1":"660.30","2":"19.62662","3":"12"},{"1":"656.80","2":"19.59016","3":"12"},{"1":"643.05","2":"20.87187","3":"12"},{"1":"671.90","2":"21.11500","3":"12"},{"1":"663.35","2":"20.08452","3":"12"},{"1":"649.30","2":"19.91049","3":"12"},{"1":"668.60","2":"17.81285","3":"12"},{"1":"644.20","2":"18.13333","3":"12"},{"1":"673.05","2":"19.22221","3":"12"},{"1":"651.80","2":"18.66072","3":"12"},{"1":"666.65","2":"19.60000","3":"12"},{"1":"652.40","2":"19.28384","3":"12"},{"1":"651.40","2":"22.81818","3":"12"},{"1":"643.70","2":"18.80922","3":"12"},{"1":"623.10","2":"21.37363","3":"12"},{"1":"629.80","2":"20.02041","3":"12"},{"1":"628.55","2":"21.49862","3":"12"},{"1":"654.85","2":"15.42857","3":"12"},{"1":"643.20","2":"22.40000","3":"12"},{"1":"632.85","2":"20.12709","3":"12"},{"1":"626.80","2":"19.03798","3":"12"},{"1":"639.75","2":"17.34216","3":"12"},{"1":"679.75","2":"17.01863","3":"12"},{"1":"654.85","2":"20.80000","3":"12"},{"1":"672.20","2":"21.15385","3":"12"},{"1":"667.45","2":"18.45833","3":"12"},{"1":"684.30","2":"19.14082","3":"12"},{"1":"653.85","2":"19.40766","3":"12"},{"1":"647.60","2":"19.56896","3":"12"},{"1":"662.70","2":"21.50120","3":"12"},{"1":"693.95","2":"17.52941","3":"12"},{"1":"654.20","2":"16.43017","3":"12"},{"1":"664.00","2":"19.79654","3":"12"},{"1":"648.00","2":"17.18613","3":"12"},{"1":"673.55","2":"17.61589","3":"12"},{"1":"644.70","2":"20.12537","3":"12"},{"1":"670.00","2":"22.16667","3":"12"},{"1":"659.40","2":"19.96154","3":"12"},{"1":"641.45","2":"19.03945","3":"12"},{"1":"630.55","2":"15.22436","3":"12"},{"1":"672.45","2":"21.14475","3":"12"},{"1":"651.20","2":"19.64390","3":"12"},{"1":"662.65","2":"21.04869","3":"12"},{"1":"633.05","2":"20.17544","3":"12"},{"1":"665.35","2":"21.39130","3":"12"},{"1":"605.55","2":"20.00833","3":"12"},{"1":"649.15","2":"20.29137","3":"12"},{"1":"650.60","2":"17.66667","3":"12"},{"1":"644.40","2":"18.22055","3":"12"},{"1":"665.10","2":"20.27100","3":"12"},{"1":"640.85","2":"20.19895","3":"12"},{"1":"643.60","2":"21.38424","3":"12"},{"1":"639.90","2":"20.97368","3":"12"},{"1":"646.90","2":"20.00000","3":"12"},{"1":"634.15","2":"17.15328","3":"12"},{"1":"677.25","2":"22.34977","3":"12"},{"1":"654.30","2":"22.17007","3":"12"},{"1":"615.75","2":"18.18182","3":"12"},{"1":"635.60","2":"18.95714","3":"12"},{"1":"681.30","2":"19.74533","3":"12"},{"1":"662.40","2":"16.42623","3":"12"},{"1":"635.60","2":"16.62540","3":"12"},{"1":"655.05","2":"16.38177","3":"12"},{"1":"630.40","2":"20.07416","3":"12"},{"1":"661.95","2":"17.99544","3":"12"},{"1":"691.90","2":"19.39130","3":"12"},{"1":"671.90","2":"16.42857","3":"12"},{"1":"663.85","2":"16.72949","3":"12"},{"1":"652.10","2":"24.41345","3":"12"},{"1":"642.40","2":"18.26415","3":"12"},{"1":"627.30","2":"18.95504","3":"12"},{"1":"658.00","2":"21.03896","3":"12"},{"1":"659.05","2":"20.74074","3":"12"},{"1":"698.25","2":"18.10000","3":"12"},{"1":"666.15","2":"19.84615","3":"12"},{"1":"664.15","2":"21.60000","3":"12"},{"1":"655.05","2":"22.44242","3":"12"},{"1":"661.45","2":"23.01438","3":"12"},{"1":"644.45","2":"17.74892","3":"12"},{"1":"676.55","2":"18.28664","3":"12"},{"1":"676.95","2":"19.26544","3":"12"},{"1":"662.40","2":"22.66667","3":"12"},{"1":"666.85","2":"19.29412","3":"12"},{"1":"651.15","2":"17.36364","3":"12"},{"1":"624.95","2":"19.82143","3":"12"},{"1":"700.30","2":"20.43378","3":"12"},{"1":"635.45","2":"21.03721","3":"12"},{"1":"676.85","2":"19.92462","3":"12"},{"1":"622.60","2":"19.00986","3":"12"},{"1":"637.35","2":"23.82222","3":"12"},{"1":"659.80","2":"19.36909","3":"12"},{"1":"664.95","2":"19.82857","3":"12"},{"1":"616.30","2":"15.25885","3":"12"},{"1":"678.05","2":"17.16129","3":"12"},{"1":"663.50","2":"21.81333","3":"12"},{"1":"675.70","2":"19.07471","3":"12"},{"1":"665.95","2":"25.78512","3":"12"},{"1":"668.10","2":"18.21261","3":"12"},{"1":"656.65","2":"18.16606","3":"12"},{"1":"671.65","2":"16.97297","3":"12"},{"1":"677.95","2":"21.50087","3":"12"},{"1":"666.05","2":"20.60000","3":"12"},{"1":"660.95","2":"16.99029","3":"12"},{"1":"666.10","2":"20.77954","3":"12"},{"1":"638.70","2":"15.51247","3":"12"},{"1":"661.85","2":"19.88506","3":"12"},{"1":"659.15","2":"21.39882","3":"12"},{"1":"676.15","2":"20.49751","3":"12"},{"1":"618.30","2":"19.36376","3":"12"},{"1":"653.55","2":"17.65957","3":"12"},{"1":"637.10","2":"21.01796","3":"12"},{"1":"665.65","2":"19.05565","3":"12"},{"1":"640.50","2":"22.53846","3":"12"},{"1":"639.85","2":"21.10787","3":"12"},{"1":"691.05","2":"20.05135","3":"12"},{"1":"652.00","2":"14.20176","3":"12"},{"1":"648.25","2":"18.47687","3":"12"},{"1":"680.45","2":"18.63542","3":"12"},{"1":"670.70","2":"20.94595","3":"12"},{"1":"626.90","2":"21.08548","3":"12"},{"1":"656.40","2":"18.69288","3":"12"},{"1":"663.45","2":"20.86808","3":"12"},{"1":"698.20","2":"19.82558","3":"12"},{"1":"638.30","2":"19.75000","3":"12"},{"1":"640.75","2":"19.50000","3":"12"},{"1":"665.20","2":"18.39080","3":"12"},{"1":"684.80","2":"18.78676","3":"12"},{"1":"644.45","2":"19.77018","3":"12"},{"1":"666.55","2":"19.33333","3":"12"},{"1":"664.70","2":"21.46392","3":"12"},{"1":"651.45","2":"23.08492","3":"12"},{"1":"646.20","2":"21.06299","3":"12"},{"1":"673.30","2":"18.68687","3":"12"},{"1":"675.40","2":"20.77024","3":"12"},{"1":"687.55","2":"19.30556","3":"12"},{"1":"672.55","2":"20.13280","3":"12"},{"1":"664.45","2":"20.66964","3":"12"},{"1":"660.75","2":"22.28155","3":"12"},{"1":"667.15","2":"20.60027","3":"12"},{"1":"666.70","2":"20.82734","3":"12"},{"1":"632.45","2":"19.22492","3":"12"},{"1":"671.25","2":"17.65477","3":"12"},{"1":"638.30","2":"17.00000","3":"12"},{"1":"655.75","2":"16.49773","3":"12"},{"1":"658.60","2":"19.78261","3":"12"},{"1":"646.35","2":"22.30216","3":"12"},{"1":"634.40","2":"17.73077","3":"12"},{"1":"657.15","2":"20.44836","3":"12"},{"1":"622.05","2":"20.37169","3":"12"},{"1":"655.80","2":"20.16479","3":"12"},{"1":"669.80","2":"21.61538","3":"12"},{"1":"661.85","2":"20.56143","3":"12"},{"1":"684.35","2":"19.95551","3":"12"},{"1":"679.50","2":"21.18387","3":"12"},{"1":"652.40","2":"18.81042","3":"12"},{"1":"679.40","2":"20.57838","3":"12"},{"1":"654.90","2":"18.32461","3":"12"},{"1":"634.55","2":"18.82063","3":"12"},{"1":"671.95","2":"20.81633","3":"12"},{"1":"654.60","2":"20.00000","3":"12"},{"1":"652.10","2":"19.68182","3":"12"},{"1":"646.00","2":"19.39018","3":"12"},{"1":"628.65","2":"20.92732","3":"12"},{"1":"657.65","2":"19.94437","3":"12"},{"1":"654.10","2":"20.79109","3":"12"},{"1":"650.65","2":"19.20354","3":"12"},{"1":"656.80","2":"19.02439","3":"12"},{"1":"630.35","2":"17.62058","3":"12"},{"1":"621.40","2":"20.23715","3":"12"},{"1":"662.55","2":"19.29374","3":"12"},{"1":"666.65","2":"18.82998","3":"12"},{"1":"641.80","2":"20.33949","3":"12"},{"1":"623.20","2":"19.22900","3":"12"},{"1":"669.35","2":"17.89130","3":"12"},{"1":"668.40","2":"19.51881","3":"12"},{"1":"647.70","2":"19.08451","3":"12"},{"1":"686.70","2":"19.93548","3":"12"},{"1":"645.75","2":"18.87326","3":"12"},{"1":"662.90","2":"20.14178","3":"12"},{"1":"633.15","2":"23.55637","3":"12"},{"1":"639.30","2":"21.46479","3":"12"},{"1":"668.90","2":"19.19101","3":"12"},{"1":"647.25","2":"20.13080","3":"12"},{"1":"652.35","2":"25.80000","3":"12"},{"1":"621.75","2":"18.77774","3":"12"},{"1":"653.70","2":"19.10982","3":"12"},{"1":"624.55","2":"19.70109","3":"12"},{"1":"663.90","2":"18.61594","3":"12"},{"1":"671.70","2":"20.99721","3":"12"},{"1":"636.90","2":"20.00000","3":"12"},{"1":"618.05","2":"20.98325","3":"12"},{"1":"655.05","2":"21.64262","3":"12"},{"1":"625.30","2":"20.02967","3":"12"},{"1":"660.05","2":"19.81140","3":"12"},{"1":"664.40","2":"18.00000","3":"12"},{"1":"639.50","2":"19.35811","3":"12"},{"1":"625.85","2":"20.17912","3":"12"},{"1":"667.45","2":"21.11986","3":"12"},{"1":"660.20","2":"23.38974","3":"12"},{"1":"634.00","2":"22.18182","3":"12"},{"1":"654.20","2":"19.94283","3":"12"},{"1":"636.50","2":"17.78826","3":"12"},{"1":"646.70","2":"14.70588","3":"12"},{"1":"661.35","2":"19.04077","3":"12"},{"1":"696.55","2":"20.89195","3":"12"},{"1":"655.40","2":"19.83851","3":"12"},{"1":"648.35","2":"19.52191","3":"12"},{"1":"653.50","2":"20.68622","3":"12"},{"1":"669.30","2":"18.18182","3":"12"},{"1":"636.70","2":"18.89224","3":"12"},{"1":"655.55","2":"24.88889","3":"12"},{"1":"632.20","2":"18.58064","3":"12"},{"1":"656.55","2":"18.04000","3":"12"},{"1":"665.90","2":"17.73399","3":"12"},{"1":"628.75","2":"21.45455","3":"12"},{"1":"634.20","2":"19.92343","3":"12"},{"1":"632.25","2":"20.33942","3":"12"},{"1":"634.05","2":"22.54608","3":"12"},{"1":"694.80","2":"21.10344","3":"12"},{"1":"659.90","2":"18.19743","3":"12"},{"1":"638.20","2":"20.10768","3":"12"},{"1":"659.35","2":"19.15984","3":"12"},{"1":"635.05","2":"19.54545","3":"12"},{"1":"633.65","2":"20.88889","3":"12"},{"1":"691.35","2":"18.39150","3":"12"},{"1":"682.15","2":"19.17990","3":"12"},{"1":"633.90","2":"19.39771","3":"12"},{"1":"624.15","2":"21.67827","3":"12"},{"1":"658.35","2":"19.28889","3":"12"},{"1":"694.25","2":"20.34927","3":"12"},{"1":"664.00","2":"20.96416","3":"12"},{"1":"623.60","2":"19.46039","3":"12"},{"1":"616.30","2":"19.28572","3":"12"},{"1":"666.45","2":"20.91979","3":"12"},{"1":"644.20","2":"20.90021","3":"12"},{"1":"634.95","2":"20.59575","3":"12"},{"1":"671.30","2":"19.37500","3":"12"},{"1":"647.05","2":"19.95122","3":"12"},{"1":"681.30","2":"18.84973","3":"12"},{"1":"678.40","2":"18.11787","3":"12"},{"1":"662.50","2":"19.18341","3":"12"},{"1":"657.80","2":"22.00000","3":"12"},{"1":"640.90","2":"21.58416","3":"12"},{"1":"679.80","2":"20.38889","3":"12"},{"1":"669.35","2":"16.29310","3":"12"},{"1":"639.25","2":"18.27778","3":"12"},{"1":"661.60","2":"19.37472","3":"12"},{"1":"661.20","2":"18.90909","3":"12"},{"1":"650.90","2":"16.40693","3":"12"},{"1":"642.20","2":"15.59140","3":"12"},{"1":"657.90","2":"18.70694","3":"12"},{"1":"631.40","2":"18.32985","3":"12"},{"1":"699.10","2":"17.90235","3":"12"},{"1":"672.05","2":"18.91157","3":"12"},{"1":"664.75","2":"20.32497","3":"12"},{"1":"631.95","2":"20.02457","3":"12"},{"1":"606.75","2":"24.00000","3":"12"},{"1":"648.70","2":"17.60784","3":"12"},{"1":"648.95","2":"19.34853","3":"12"},{"1":"612.50","2":"19.67846","3":"12"},{"1":"638.55","2":"18.72861","3":"12"},{"1":"637.95","2":"15.88235","3":"12"},{"1":"662.55","2":"20.05491","3":"12"},{"1":"640.15","2":"17.98825","3":"12"},{"1":"645.55","2":"16.96629","3":"12"},{"1":"660.10","2":"19.23937","3":"12"},{"1":"651.85","2":"19.19586","3":"12"},{"1":"626.10","2":"19.59906","3":"12"},{"1":"644.50","2":"20.54348","3":"12"},{"1":"664.30","2":"18.58848","3":"12"},{"1":"663.85","2":"15.60419","3":"12"},{"1":"658.80","2":"15.29304","3":"12"},{"1":"657.40","2":"17.65537","3":"12"},{"1":"640.10","2":"17.57976","3":"12"},{"1":"631.90","2":"22.33333","3":"12"},{"1":"637.95","2":"18.75000","3":"12"},{"1":"668.80","2":"18.10241","3":"12"},{"1":"617.35","2":"20.25641","3":"12"},{"1":"671.75","2":"18.80207","3":"12"},{"1":"643.50","2":"18.77230","3":"12"},{"1":"673.25","2":"20.40521","3":"12"},{"1":"636.60","2":"18.65079","3":"12"},{"1":"639.80","2":"20.70707","3":"12"},{"1":"666.00","2":"22.00000","3":"12"},{"1":"698.45","2":"17.69978","3":"12"},{"1":"673.90","2":"21.48329","3":"12"},{"1":"704.30","2":"16.70103","3":"12"},{"1":"682.55","2":"19.57567","3":"12"},{"1":"672.30","2":"17.25806","3":"12"},{"1":"655.35","2":"17.37526","3":"12"},{"1":"683.35","2":"17.34931","3":"12"},{"1":"645.55","2":"16.26229","3":"12"},{"1":"661.60","2":"17.70045","3":"12"},{"1":"695.30","2":"20.12881","3":"12"},{"1":"656.50","2":"18.26539","3":"12"},{"1":"641.45","2":"14.54214","3":"12"},{"1":"628.40","2":"19.15261","3":"12"},{"1":"667.60","2":"17.36574","3":"12"},{"1":"672.70","2":"15.13898","3":"12"},{"1":"669.85","2":"17.84266","3":"12"},{"1":"661.85","2":"15.40704","3":"12"},{"1":"630.55","2":"18.86534","3":"12"},{"1":"636.10","2":"16.47413","3":"12"},{"1":"643.40","2":"17.86263","3":"12"},{"1":"695.20","2":"21.88586","3":"12"},{"1":"650.55","2":"20.20000","3":"12"},{"1":"638.35","2":"19.03640","3":"12"},{"1":"649.50","2":"17.88991","3":"13"},{"1":"637.35","2":"21.52466","3":"13"},{"1":"656.65","2":"18.69723","3":"13"},{"1":"653.40","2":"17.35714","3":"13"},{"1":"639.50","2":"18.67133","3":"13"},{"1":"669.35","2":"21.40625","3":"13"},{"1":"628.65","2":"19.50000","3":"13"},{"1":"650.55","2":"20.89412","3":"13"},{"1":"683.35","2":"19.94737","3":"13"},{"1":"684.80","2":"20.80556","3":"13"},{"1":"652.40","2":"21.23809","3":"13"},{"1":"654.85","2":"21.00000","3":"13"},{"1":"641.45","2":"20.60000","3":"13"},{"1":"635.95","2":"20.00822","3":"13"},{"1":"700.30","2":"18.02778","3":"13"},{"1":"624.95","2":"20.25196","3":"13"},{"1":"676.95","2":"16.97787","3":"13"},{"1":"666.05","2":"16.50980","3":"13"},{"1":"664.40","2":"22.70402","3":"13"},{"1":"638.35","2":"19.91111","3":"13"},{"1":"606.75","2":"18.33333","3":"13"},{"1":"645.55","2":"22.61905","3":"13"},{"1":"638.30","2":"19.44828","3":"13"},{"1":"615.75","2":"25.05263","3":"13"},{"1":"663.90","2":"20.67544","3":"13"},{"1":"650.45","2":"18.68235","3":"13"},{"1":"652.00","2":"22.84553","3":"13"},{"1":"627.30","2":"19.26667","3":"13"},{"1":"682.45","2":"19.25000","3":"13"},{"1":"651.80","2":"20.54545","3":"13"},{"1":"691.90","2":"20.60697","3":"13"},{"1":"660.30","2":"21.07268","3":"13"},{"1":"643.40","2":"21.53581","3":"13"},{"1":"666.85","2":"19.90400","3":"13"},{"1":"644.40","2":"21.19407","3":"13"},{"1":"663.45","2":"21.86535","3":"13"},{"1":"616.45","2":"18.32965","3":"13"},{"1":"647.60","2":"16.22857","3":"13"},{"1":"673.30","2":"19.17857","3":"13"},{"1":"655.30","2":"20.27737","3":"13"},{"1":"616.30","2":"22.98614","3":"13"},{"1":"649.30","2":"20.44444","3":"13"},{"1":"630.35","2":"19.82085","3":"13"},{"1":"665.95","2":"23.20522","3":"13"},{"1":"662.50","2":"19.26697","3":"13"},{"1":"683.40","2":"23.30189","3":"13"},{"1":"662.40","2":"21.18829","3":"13"},{"1":"632.20","2":"20.87180","3":"13"},{"1":"650.65","2":"19.01749","3":"13"},{"1":"659.80","2":"21.91938","3":"13"},{"1":"661.60","2":"20.10124","3":"13"},{"1":"624.55","2":"21.47651","3":"13"},{"1":"669.10","2":"20.06579","3":"13"},{"1":"668.90","2":"20.37510","3":"13"},{"1":"657.55","2":"22.44648","3":"13"},{"1":"668.00","2":"22.89524","3":"13"},{"1":"646.50","2":"20.49797","3":"13"},{"1":"646.90","2":"20.00000","3":"13"},{"1":"654.30","2":"22.25658","3":"13"},{"1":"675.40","2":"21.56436","3":"13"},{"1":"621.40","2":"19.47737","3":"13"},{"1":"638.20","2":"17.67002","3":"13"},{"1":"637.00","2":"21.94756","3":"13"},{"1":"639.90","2":"21.78339","3":"13"},{"1":"669.30","2":"19.14000","3":"13"},{"1":"657.00","2":"18.11050","3":"13"},{"1":"630.55","2":"20.68242","3":"13"},{"1":"650.90","2":"22.62361","3":"13"},{"1":"666.15","2":"21.78650","3":"13"},{"1":"646.70","2":"18.58293","3":"13"},{"1":"666.65","2":"21.54545","3":"13"},{"1":"654.10","2":"21.15289","3":"13"},{"1":"655.75","2":"16.63333","3":"13"},{"1":"677.25","2":"21.14438","3":"13"},{"1":"633.90","2":"19.78182","3":"13"},{"1":"643.25","2":"18.98373","3":"13"},{"1":"653.50","2":"17.66767","3":"13"},{"1":"661.35","2":"17.75499","3":"13"},{"1":"662.75","2":"15.27273","3":"13"},{"1":"661.20","2":"14.00000","3":"13"},{"1":"673.90","2":"20.59613","3":"13"},{"1":"671.90","2":"16.31169","3":"13"},{"1":"666.70","2":"21.12796","3":"13"},{"1":"660.20","2":"17.48801","3":"13"},{"1":"655.35","2":"17.88679","3":"13"},{"1":"654.85","2":"19.30676","3":"13"},{"1":"647.70","2":"20.89231","3":"13"},{"1":"628.40","2":"21.28684","3":"13"},{"1":"681.30","2":"20.19560","3":"13"},{"1":"623.20","2":"24.95000","3":"13"},{"1":"628.55","2":"18.13043","3":"13"},{"1":"653.85","2":"20.00000","3":"13"},{"1":"661.45","2":"18.72951","3":"13"},{"1":"686.70","2":"18.25000","3":"13"},{"1":"647.30","2":"18.99257","3":"13"},{"1":"644.55","2":"19.88764","3":"13"},{"1":"640.75","2":"19.37895","3":"13"},{"1":"668.40","2":"20.46259","3":"13"},{"1":"672.30","2":"22.29157","3":"13"},{"1":"624.15","2":"20.70474","3":"13"},{"1":"634.15","2":"19.06005","3":"13"},{"1":"672.45","2":"20.23247","3":"13"},{"1":"666.85","2":"19.69012","3":"13"},{"1":"687.55","2":"20.36254","3":"13"},{"1":"645.25","2":"19.75422","3":"13"},{"1":"677.95","2":"19.37977","3":"13"},{"1":"664.95","2":"22.92351","3":"13"},{"1":"646.95","2":"19.37340","3":"13"},{"1":"655.05","2":"19.15516","3":"13"},{"1":"656.50","2":"21.30000","3":"13"},{"1":"645.75","2":"18.30357","3":"13"},{"1":"659.40","2":"21.07926","3":"13"},{"1":"637.65","2":"18.79121","3":"13"},{"1":"658.00","2":"19.62662","3":"13"},{"1":"638.55","2":"19.59016","3":"13"},{"1":"634.05","2":"20.87187","3":"13"},{"1":"635.05","2":"21.11500","3":"13"},{"1":"665.10","2":"20.08452","3":"13"},{"1":"646.55","2":"19.91049","3":"13"},{"1":"651.15","2":"17.81285","3":"13"},{"1":"616.30","2":"18.13333","3":"13"},{"1":"632.95","2":"19.22221","3":"13"},{"1":"669.80","2":"18.66072","3":"13"},{"1":"641.55","2":"19.60000","3":"13"},{"1":"651.90","2":"19.28384","3":"13"},{"1":"657.65","2":"22.81818","3":"13"},{"1":"676.55","2":"18.80922","3":"13"},{"1":"679.65","2":"21.37363","3":"13"},{"1":"693.95","2":"20.02041","3":"13"},{"1":"637.95","2":"21.49862","3":"13"},{"1":"634.20","2":"15.42857","3":"13"},{"1":"658.35","2":"22.40000","3":"13"},{"1":"662.55","2":"20.12709","3":"13"},{"1":"638.70","2":"19.03798","3":"13"},{"1":"665.90","2":"17.34216","3":"13"},{"1":"661.85","2":"17.01863","3":"13"},{"1":"679.75","2":"20.80000","3":"13"},{"1":"630.55","2":"21.15385","3":"13"},{"1":"664.30","2":"18.45833","3":"13"},{"1":"643.40","2":"19.14082","3":"13"},{"1":"650.90","2":"19.40766","3":"13"},{"1":"648.70","2":"19.56896","3":"13"},{"1":"628.75","2":"21.50120","3":"13"},{"1":"623.10","2":"17.52941","3":"13"},{"1":"691.05","2":"16.43017","3":"13"},{"1":"657.15","2":"19.79654","3":"13"},{"1":"651.40","2":"17.18613","3":"13"},{"1":"646.00","2":"17.61589","3":"13"},{"1":"648.20","2":"20.12537","3":"13"},{"1":"667.45","2":"22.16667","3":"13"},{"1":"671.65","2":"19.96154","3":"13"},{"1":"631.05","2":"19.03945","3":"13"},{"1":"657.50","2":"15.22436","3":"13"},{"1":"628.25","2":"21.14475","3":"13"},{"1":"662.55","2":"19.64390","3":"13"},{"1":"629.80","2":"21.04869","3":"13"},{"1":"655.05","2":"20.17544","3":"13"},{"1":"671.95","2":"21.39130","3":"13"},{"1":"652.30","2":"20.00833","3":"13"},{"1":"664.15","2":"20.29137","3":"13"},{"1":"626.10","2":"17.66667","3":"13"},{"1":"673.05","2":"18.22055","3":"13"},{"1":"631.85","2":"20.27100","3":"13"},{"1":"621.75","2":"20.19895","3":"13"},{"1":"616.30","2":"21.38424","3":"13"},{"1":"634.55","2":"20.97368","3":"13"},{"1":"678.40","2":"20.00000","3":"13"},{"1":"609.00","2":"17.15328","3":"13"},{"1":"680.05","2":"22.34977","3":"13"},{"1":"671.60","2":"22.17007","3":"13"},{"1":"673.55","2":"18.18182","3":"13"},{"1":"653.95","2":"18.95714","3":"13"},{"1":"663.50","2":"19.74533","3":"13"},{"1":"637.10","2":"16.42623","3":"13"},{"1":"643.50","2":"16.62540","3":"13"},{"1":"646.20","2":"16.38177","3":"13"},{"1":"668.65","2":"20.07416","3":"13"},{"1":"690.80","2":"17.99544","3":"13"},{"1":"666.65","2":"19.39130","3":"13"},{"1":"653.80","2":"16.42857","3":"13"},{"1":"649.15","2":"16.72949","3":"13"},{"1":"655.55","2":"24.41345","3":"13"},{"1":"670.00","2":"18.26415","3":"13"},{"1":"640.90","2":"18.95504","3":"13"},{"1":"696.55","2":"21.03896","3":"13"},{"1":"671.75","2":"20.74074","3":"13"},{"1":"670.70","2":"18.10000","3":"13"},{"1":"681.30","2":"19.84615","3":"13"},{"1":"684.35","2":"21.60000","3":"13"},{"1":"635.60","2":"22.44242","3":"13"},{"1":"644.45","2":"23.01438","3":"13"},{"1":"620.50","2":"17.74892","3":"13"},{"1":"694.80","2":"18.28664","3":"13"},{"1":"632.25","2":"19.26544","3":"13"},{"1":"655.80","2":"22.66667","3":"13"},{"1":"660.10","2":"19.29412","3":"13"},{"1":"651.35","2":"17.36364","3":"13"},{"1":"678.05","2":"19.82143","3":"13"},{"1":"682.55","2":"20.43378","3":"13"},{"1":"649.70","2":"21.03721","3":"13"},{"1":"639.80","2":"19.92462","3":"13"},{"1":"659.05","2":"19.00986","3":"13"},{"1":"651.20","2":"23.82222","3":"13"},{"1":"638.00","2":"19.36909","3":"13"},{"1":"634.40","2":"19.82857","3":"13"},{"1":"660.75","2":"15.25885","3":"13"},{"1":"667.15","2":"17.16129","3":"13"},{"1":"655.20","2":"21.81333","3":"13"},{"1":"668.95","2":"19.07471","3":"13"},{"1":"643.20","2":"25.78512","3":"13"},{"1":"689.10","2":"18.21261","3":"13"},{"1":"653.10","2":"18.16606","3":"13"},{"1":"658.60","2":"16.97297","3":"13"},{"1":"638.30","2":"21.50087","3":"13"},{"1":"625.85","2":"20.60000","3":"13"},{"1":"664.70","2":"16.99029","3":"13"},{"1":"626.80","2":"20.77954","3":"13"},{"1":"640.85","2":"15.51247","3":"13"},{"1":"636.10","2":"19.88506","3":"13"},{"1":"674.25","2":"21.39882","3":"13"},{"1":"671.70","2":"20.49751","3":"13"},{"1":"675.70","2":"19.36376","3":"13"},{"1":"651.85","2":"17.65957","3":"13"},{"1":"647.60","2":"21.01796","3":"13"},{"1":"664.75","2":"19.05565","3":"13"},{"1":"644.20","2":"22.53846","3":"13"},{"1":"666.55","2":"21.10787","3":"13"},{"1":"634.10","2":"20.05135","3":"13"},{"1":"673.25","2":"14.20176","3":"13"},{"1":"652.10","2":"18.47687","3":"13"},{"1":"666.10","2":"18.63542","3":"13"},{"1":"631.40","2":"20.94595","3":"13"},{"1":"657.80","2":"21.08548","3":"13"},{"1":"681.60","2":"18.69288","3":"13"},{"1":"653.70","2":"20.86808","3":"13"},{"1":"648.25","2":"19.82558","3":"13"},{"1":"662.70","2":"19.75000","3":"13"},{"1":"605.55","2":"19.50000","3":"13"},{"1":"672.55","2":"18.39080","3":"13"},{"1":"676.15","2":"18.78676","3":"13"},{"1":"652.85","2":"19.77018","3":"13"},{"1":"656.80","2":"19.33333","3":"13"},{"1":"655.40","2":"21.46392","3":"13"},{"1":"639.25","2":"23.08492","3":"13"},{"1":"672.05","2":"21.06299","3":"13"},{"1":"691.35","2":"18.68687","3":"13"},{"1":"655.70","2":"20.77024","3":"13"},{"1":"684.95","2":"19.30556","3":"13"},{"1":"639.30","2":"20.13280","3":"13"},{"1":"648.35","2":"20.66964","3":"13"},{"1":"634.90","2":"22.28155","3":"13"},{"1":"634.70","2":"20.60027","3":"13"},{"1":"694.25","2":"20.82734","3":"13"},{"1":"661.85","2":"19.22492","3":"13"},{"1":"653.55","2":"17.65477","3":"13"},{"1":"618.05","2":"17.00000","3":"13"},{"1":"640.15","2":"16.49773","3":"13"},{"1":"619.80","2":"19.78261","3":"13"},{"1":"664.45","2":"22.30216","3":"13"},{"1":"667.60","2":"17.73077","3":"13"},{"1":"643.50","2":"20.44836","3":"13"},{"1":"657.75","2":"20.37169","3":"13"},{"1":"643.70","2":"20.16479","3":"13"},{"1":"640.10","2":"21.61538","3":"13"},{"1":"639.85","2":"20.56143","3":"13"},{"1":"655.85","2":"19.95551","3":"13"},{"1":"669.30","2":"21.18387","3":"13"},{"1":"654.20","2":"18.81042","3":"13"},{"1":"671.30","2":"20.57838","3":"13"},{"1":"646.40","2":"18.32461","3":"13"},{"1":"658.80","2":"18.82063","3":"13"},{"1":"654.20","2":"20.81633","3":"13"},{"1":"664.95","2":"20.00000","3":"13"},{"1":"644.95","2":"19.68182","3":"13"},{"1":"656.80","2":"19.39018","3":"13"},{"1":"662.45","2":"20.92732","3":"13"},{"1":"620.30","2":"19.94437","3":"13"},{"1":"666.45","2":"20.79109","3":"13"},{"1":"632.45","2":"19.20354","3":"13"},{"1":"648.95","2":"19.02439","3":"13"},{"1":"706.75","2":"17.62058","3":"13"},{"1":"643.60","2":"20.23715","3":"13"},{"1":"642.40","2":"19.29374","3":"13"},{"1":"642.20","2":"18.82998","3":"13"},{"1":"623.60","2":"20.33949","3":"13"},{"1":"641.80","2":"19.22900","3":"13"},{"1":"669.95","2":"17.89130","3":"13"},{"1":"672.70","2":"19.51881","3":"13"},{"1":"642.75","2":"19.08451","3":"13"},{"1":"654.90","2":"19.93548","3":"13"},{"1":"644.70","2":"18.87326","3":"13"},{"1":"646.35","2":"20.14178","3":"13"},{"1":"656.55","2":"23.55637","3":"13"},{"1":"664.00","2":"21.46479","3":"13"},{"1":"672.35","2":"19.19101","3":"13"},{"1":"632.00","2":"20.13080","3":"13"},{"1":"633.15","2":"25.80000","3":"13"},{"1":"649.85","2":"18.77774","3":"13"},{"1":"636.95","2":"19.10982","3":"13"},{"1":"633.05","2":"19.70109","3":"13"},{"1":"644.45","2":"18.61594","3":"13"},{"1":"643.70","2":"20.99721","3":"13"},{"1":"676.85","2":"20.00000","3":"13"},{"1":"682.15","2":"20.98325","3":"13"},{"1":"652.40","2":"21.64262","3":"13"},{"1":"671.60","2":"20.02967","3":"13"},{"1":"679.50","2":"19.81140","3":"13"},{"1":"635.75","2":"18.00000","3":"13"},{"1":"633.65","2":"19.35811","3":"13"},{"1":"635.45","2":"20.17912","3":"13"},{"1":"645.10","2":"21.11986","3":"13"},{"1":"659.90","2":"23.38974","3":"13"},{"1":"647.25","2":"22.18182","3":"13"},{"1":"669.85","2":"19.94283","3":"13"},{"1":"634.95","2":"17.78826","3":"13"},{"1":"661.90","2":"14.70588","3":"13"},{"1":"657.40","2":"19.04077","3":"13"},{"1":"663.85","2":"20.89195","3":"13"},{"1":"662.65","2":"19.83851","3":"13"},{"1":"645.60","2":"19.52191","3":"13"},{"1":"648.00","2":"20.68622","3":"13"},{"1":"612.50","2":"18.18182","3":"13"},{"1":"640.50","2":"18.89224","3":"13"},{"1":"679.80","2":"24.88889","3":"13"},{"1":"661.85","2":"18.58064","3":"13"},{"1":"655.35","2":"18.04000","3":"13"},{"1":"655.05","2":"17.73399","3":"13"},{"1":"651.45","2":"21.45455","3":"13"},{"1":"660.95","2":"19.92343","3":"13"},{"1":"659.15","2":"20.33942","3":"13"},{"1":"668.80","2":"22.54608","3":"13"},{"1":"623.45","2":"21.10344","3":"13"},{"1":"625.30","2":"18.19743","3":"13"},{"1":"626.90","2":"20.10768","3":"13"},{"1":"676.60","2":"19.15984","3":"13"},{"1":"645.00","2":"19.54545","3":"13"},{"1":"663.85","2":"20.88889","3":"13"},{"1":"656.40","2":"18.39150","3":"13"},{"1":"631.90","2":"19.17990","3":"13"},{"1":"644.20","2":"19.39771","3":"13"},{"1":"666.00","2":"21.67827","3":"13"},{"1":"663.35","2":"19.28889","3":"13"},{"1":"636.50","2":"20.34927","3":"13"},{"1":"656.70","2":"20.96416","3":"13"},{"1":"661.60","2":"19.46039","3":"13"},{"1":"698.20","2":"19.28572","3":"13"},{"1":"672.20","2":"20.91979","3":"13"},{"1":"641.10","2":"20.90021","3":"13"},{"1":"612.65","2":"20.59575","3":"13"},{"1":"664.00","2":"19.37500","3":"13"},{"1":"665.20","2":"19.95122","3":"13"},{"1":"636.90","2":"18.84973","3":"13"},{"1":"686.05","2":"18.11787","3":"13"},{"1":"652.35","2":"19.18341","3":"13"},{"1":"671.25","2":"22.00000","3":"13"},{"1":"637.95","2":"21.58416","3":"13"},{"1":"695.20","2":"20.38889","3":"13"},{"1":"659.35","2":"16.29310","3":"13"},{"1":"704.30","2":"18.27778","3":"13"},{"1":"639.35","2":"19.37472","3":"13"},{"1":"631.95","2":"18.90909","3":"13"},{"1":"622.60","2":"16.40693","3":"13"},{"1":"647.05","2":"15.59140","3":"13"},{"1":"639.75","2":"18.70694","3":"13"},{"1":"662.40","2":"18.32985","3":"13"},{"1":"698.45","2":"17.90235","3":"13"},{"1":"634.00","2":"18.91157","3":"13"},{"1":"662.90","2":"20.32497","3":"13"},{"1":"695.30","2":"20.02457","3":"13"},{"1":"657.00","2":"24.00000","3":"13"},{"1":"627.25","2":"17.60784","3":"13"},{"1":"672.05","2":"19.34853","3":"13"},{"1":"657.90","2":"19.67846","3":"13"},{"1":"660.05","2":"18.72861","3":"13"},{"1":"671.90","2":"15.88235","3":"13"},{"1":"667.45","2":"20.05491","3":"13"},{"1":"642.20","2":"17.98825","3":"13"},{"1":"661.95","2":"16.96629","3":"13"},{"1":"668.60","2":"19.23937","3":"13"},{"1":"666.60","2":"19.19586","3":"13"},{"1":"667.20","2":"19.59906","3":"13"},{"1":"664.15","2":"20.54348","3":"13"},{"1":"654.60","2":"18.58848","3":"13"},{"1":"652.50","2":"15.60419","3":"13"},{"1":"622.05","2":"15.29304","3":"13"},{"1":"641.45","2":"17.65537","3":"13"},{"1":"618.30","2":"17.57976","3":"13"},{"1":"665.35","2":"22.33333","3":"13"},{"1":"680.45","2":"18.75000","3":"13"},{"1":"679.40","2":"18.10241","3":"13"},{"1":"659.40","2":"20.25641","3":"13"},{"1":"682.65","2":"18.80207","3":"13"},{"1":"665.65","2":"18.77230","3":"13"},{"1":"645.75","2":"20.40521","3":"13"},{"1":"635.20","2":"18.65079","3":"13"},{"1":"666.15","2":"20.70707","3":"13"},{"1":"678.80","2":"22.00000","3":"13"},{"1":"698.25","2":"17.69978","3":"13"},{"1":"634.10","2":"21.48329","3":"13"},{"1":"652.10","2":"16.70103","3":"13"},{"1":"653.55","2":"19.57567","3":"13"},{"1":"668.10","2":"17.25806","3":"13"},{"1":"632.85","2":"17.37526","3":"13"},{"1":"650.60","2":"17.34931","3":"13"},{"1":"635.60","2":"16.26229","3":"13"},{"1":"636.70","2":"17.70045","3":"13"},{"1":"669.35","2":"20.12881","3":"13"},{"1":"627.10","2":"18.26539","3":"13"},{"1":"661.90","2":"14.54214","3":"13"},{"1":"673.55","2":"19.15261","3":"13"},{"1":"630.40","2":"17.36574","3":"13"},{"1":"684.30","2":"15.13898","3":"13"},{"1":"652.30","2":"17.84266","3":"13"},{"1":"643.05","2":"15.40704","3":"13"},{"1":"636.60","2":"18.86534","3":"13"},{"1":"645.55","2":"16.47413","3":"13"},{"1":"699.10","2":"17.86263","3":"13"},{"1":"617.35","2":"21.88586","3":"13"},{"1":"681.90","2":"20.20000","3":"13"},{"1":"644.50","2":"19.03640","3":"13"},{"1":"636.95","2":"17.88991","3":"14"},{"1":"641.55","2":"21.52466","3":"14"},{"1":"660.10","2":"18.69723","3":"14"},{"1":"681.90","2":"17.35714","3":"14"},{"1":"654.90","2":"18.67133","3":"14"},{"1":"643.40","2":"21.40625","3":"14"},{"1":"667.45","2":"19.50000","3":"14"},{"1":"664.00","2":"20.89412","3":"14"},{"1":"675.40","2":"19.94737","3":"14"},{"1":"657.75","2":"20.80556","3":"14"},{"1":"695.20","2":"21.23809","3":"14"},{"1":"637.95","2":"21.00000","3":"14"},{"1":"656.50","2":"20.60000","3":"14"},{"1":"655.35","2":"20.00822","3":"14"},{"1":"648.20","2":"18.02778","3":"14"},{"1":"672.35","2":"20.25196","3":"14"},{"1":"656.40","2":"16.97787","3":"14"},{"1":"655.40","2":"16.50980","3":"14"},{"1":"631.95","2":"22.70402","3":"14"},{"1":"642.20","2":"19.91111","3":"14"},{"1":"666.70","2":"18.33333","3":"14"},{"1":"683.35","2":"22.61905","3":"14"},{"1":"642.40","2":"19.44828","3":"14"},{"1":"667.15","2":"25.05263","3":"14"},{"1":"624.55","2":"20.67544","3":"14"},{"1":"651.90","2":"18.68235","3":"14"},{"1":"648.25","2":"22.84553","3":"14"},{"1":"638.55","2":"19.26667","3":"14"},{"1":"662.40","2":"19.25000","3":"14"},{"1":"668.95","2":"20.54545","3":"14"},{"1":"653.55","2":"20.60697","3":"14"},{"1":"652.00","2":"21.07268","3":"14"},{"1":"671.90","2":"21.53581","3":"14"},{"1":"668.10","2":"19.90400","3":"14"},{"1":"662.70","2":"21.19407","3":"14"},{"1":"623.45","2":"21.86535","3":"14"},{"1":"635.95","2":"18.32965","3":"14"},{"1":"638.30","2":"16.22857","3":"14"},{"1":"643.50","2":"19.17857","3":"14"},{"1":"682.55","2":"20.27737","3":"14"},{"1":"643.70","2":"22.98614","3":"14"},{"1":"616.30","2":"20.44444","3":"14"},{"1":"672.55","2":"19.82085","3":"14"},{"1":"679.65","2":"23.20522","3":"14"},{"1":"673.55","2":"19.26697","3":"14"},{"1":"668.40","2":"23.30189","3":"14"},{"1":"649.50","2":"21.18829","3":"14"},{"1":"698.20","2":"20.87180","3":"14"},{"1":"645.00","2":"19.01749","3":"14"},{"1":"686.70","2":"21.91938","3":"14"},{"1":"691.05","2":"20.10124","3":"14"},{"1":"640.50","2":"21.47651","3":"14"},{"1":"663.85","2":"20.06579","3":"14"},{"1":"651.35","2":"20.37510","3":"14"},{"1":"661.60","2":"22.44648","3":"14"},{"1":"648.00","2":"22.89524","3":"14"},{"1":"656.80","2":"20.49797","3":"14"},{"1":"638.70","2":"20.00000","3":"14"},{"1":"643.40","2":"22.25658","3":"14"},{"1":"639.90","2":"21.56436","3":"14"},{"1":"632.95","2":"19.47737","3":"14"},{"1":"647.70","2":"17.67002","3":"14"},{"1":"647.05","2":"21.94756","3":"14"},{"1":"678.40","2":"21.78339","3":"14"},{"1":"681.30","2":"19.14000","3":"14"},{"1":"646.50","2":"18.11050","3":"14"},{"1":"659.35","2":"20.68242","3":"14"},{"1":"676.55","2":"22.62361","3":"14"},{"1":"632.45","2":"21.78650","3":"14"},{"1":"666.15","2":"18.58293","3":"14"},{"1":"664.70","2":"21.54545","3":"14"},{"1":"663.85","2":"21.15289","3":"14"},{"1":"704.30","2":"16.63333","3":"14"},{"1":"630.40","2":"21.14438","3":"14"},{"1":"661.95","2":"19.78182","3":"14"},{"1":"653.40","2":"18.98373","3":"14"},{"1":"636.50","2":"17.66767","3":"14"},{"1":"639.85","2":"17.75499","3":"14"},{"1":"666.05","2":"15.27273","3":"14"},{"1":"657.15","2":"14.00000","3":"14"},{"1":"659.15","2":"20.59613","3":"14"},{"1":"634.15","2":"16.31169","3":"14"},{"1":"656.55","2":"21.12796","3":"14"},{"1":"659.80","2":"17.48801","3":"14"},{"1":"673.90","2":"17.88679","3":"14"},{"1":"645.55","2":"19.30676","3":"14"},{"1":"653.10","2":"20.89231","3":"14"},{"1":"706.75","2":"21.28684","3":"14"},{"1":"606.75","2":"20.19560","3":"14"},{"1":"671.65","2":"24.95000","3":"14"},{"1":"630.35","2":"18.13043","3":"14"},{"1":"653.85","2":"20.00000","3":"14"},{"1":"672.05","2":"18.72951","3":"14"},{"1":"663.90","2":"18.25000","3":"14"},{"1":"660.30","2":"18.99257","3":"14"},{"1":"666.45","2":"19.88764","3":"14"},{"1":"655.05","2":"19.37895","3":"14"},{"1":"655.70","2":"20.46259","3":"14"},{"1":"651.40","2":"22.29157","3":"14"},{"1":"655.55","2":"20.70474","3":"14"},{"1":"659.05","2":"19.06005","3":"14"},{"1":"651.80","2":"20.23247","3":"14"},{"1":"682.45","2":"19.69012","3":"14"},{"1":"638.00","2":"20.36254","3":"14"},{"1":"635.75","2":"19.75422","3":"14"},{"1":"646.20","2":"19.37977","3":"14"},{"1":"623.10","2":"22.92351","3":"14"},{"1":"659.90","2":"19.37340","3":"14"},{"1":"643.50","2":"19.15516","3":"14"},{"1":"632.20","2":"21.30000","3":"14"},{"1":"663.35","2":"18.30357","3":"14"},{"1":"645.75","2":"21.07926","3":"14"},{"1":"650.90","2":"18.79121","3":"14"},{"1":"698.45","2":"19.62662","3":"14"},{"1":"677.25","2":"19.59016","3":"14"},{"1":"655.85","2":"20.87187","3":"14"},{"1":"646.90","2":"21.11500","3":"14"},{"1":"665.20","2":"20.08452","3":"14"},{"1":"617.35","2":"19.91049","3":"14"},{"1":"636.60","2":"17.81285","3":"14"},{"1":"666.10","2":"18.13333","3":"14"},{"1":"620.30","2":"19.22221","3":"14"},{"1":"669.30","2":"18.66072","3":"14"},{"1":"605.55","2":"19.60000","3":"14"},{"1":"628.55","2":"19.28384","3":"14"},{"1":"636.10","2":"22.81818","3":"14"},{"1":"682.65","2":"18.80922","3":"14"},{"1":"647.30","2":"21.37363","3":"14"},{"1":"672.05","2":"20.02041","3":"14"},{"1":"676.15","2":"21.49862","3":"14"},{"1":"646.95","2":"15.42857","3":"14"},{"1":"684.30","2":"22.40000","3":"14"},{"1":"637.00","2":"20.12709","3":"14"},{"1":"684.35","2":"19.03798","3":"14"},{"1":"658.60","2":"17.34216","3":"14"},{"1":"673.55","2":"17.01863","3":"14"},{"1":"626.90","2":"20.80000","3":"14"},{"1":"618.05","2":"21.15385","3":"14"},{"1":"655.05","2":"18.45833","3":"14"},{"1":"622.05","2":"19.14082","3":"14"},{"1":"645.60","2":"19.40766","3":"14"},{"1":"657.40","2":"19.56896","3":"14"},{"1":"675.70","2":"21.50120","3":"14"},{"1":"657.90","2":"17.52941","3":"14"},{"1":"686.05","2":"16.43017","3":"14"},{"1":"616.30","2":"19.79654","3":"14"},{"1":"664.75","2":"17.18613","3":"14"},{"1":"699.10","2":"17.61589","3":"14"},{"1":"629.80","2":"20.12537","3":"14"},{"1":"640.85","2":"22.16667","3":"14"},{"1":"642.20","2":"19.96154","3":"14"},{"1":"680.05","2":"19.03945","3":"14"},{"1":"670.70","2":"15.22436","3":"14"},{"1":"633.15","2":"21.14475","3":"14"},{"1":"652.40","2":"19.64390","3":"14"},{"1":"661.90","2":"21.04869","3":"14"},{"1":"657.80","2":"20.17544","3":"14"},{"1":"669.10","2":"21.39130","3":"14"},{"1":"641.45","2":"20.00833","3":"14"},{"1":"648.95","2":"20.29137","3":"14"},{"1":"630.55","2":"17.66667","3":"14"},{"1":"681.60","2":"18.22055","3":"14"},{"1":"656.70","2":"20.27100","3":"14"},{"1":"657.65","2":"20.19895","3":"14"},{"1":"635.20","2":"21.38424","3":"14"},{"1":"684.80","2":"20.97368","3":"14"},{"1":"664.95","2":"20.00000","3":"14"},{"1":"648.35","2":"17.15328","3":"14"},{"1":"637.10","2":"22.34977","3":"14"},{"1":"666.65","2":"22.17007","3":"14"},{"1":"700.30","2":"18.18182","3":"14"},{"1":"661.85","2":"18.95714","3":"14"},{"1":"653.55","2":"19.74533","3":"14"},{"1":"665.90","2":"16.42623","3":"14"},{"1":"646.70","2":"16.62540","3":"14"},{"1":"609.00","2":"16.38177","3":"14"},{"1":"674.25","2":"20.07416","3":"14"},{"1":"691.35","2":"17.99544","3":"14"},{"1":"653.95","2":"19.39130","3":"14"},{"1":"616.45","2":"16.42857","3":"14"},{"1":"644.50","2":"16.72949","3":"14"},{"1":"652.40","2":"24.41345","3":"14"},{"1":"646.40","2":"18.26415","3":"14"},{"1":"647.60","2":"18.95504","3":"14"},{"1":"673.05","2":"21.03896","3":"14"},{"1":"659.40","2":"20.74074","3":"14"},{"1":"669.30","2":"18.10000","3":"14"},{"1":"663.50","2":"19.84615","3":"14"},{"1":"649.85","2":"21.60000","3":"14"},{"1":"655.80","2":"22.44242","3":"14"},{"1":"638.30","2":"23.01438","3":"14"},{"1":"671.60","2":"17.74892","3":"14"},{"1":"646.00","2":"18.28664","3":"14"},{"1":"672.30","2":"19.26544","3":"14"},{"1":"643.20","2":"22.66667","3":"14"},{"1":"621.40","2":"19.29412","3":"14"},{"1":"664.45","2":"17.36364","3":"14"},{"1":"667.60","2":"19.82143","3":"14"},{"1":"657.00","2":"20.43378","3":"14"},{"1":"633.05","2":"21.03721","3":"14"},{"1":"666.00","2":"19.92462","3":"14"},{"1":"687.55","2":"19.00986","3":"14"},{"1":"649.30","2":"23.82222","3":"14"},{"1":"639.30","2":"19.36909","3":"14"},{"1":"695.30","2":"19.82857","3":"14"},{"1":"669.35","2":"15.25885","3":"14"},{"1":"644.20","2":"17.16129","3":"14"},{"1":"624.95","2":"21.81333","3":"14"},{"1":"628.25","2":"19.07471","3":"14"},{"1":"696.55","2":"25.78512","3":"14"},{"1":"620.50","2":"18.21261","3":"14"},{"1":"640.90","2":"18.16606","3":"14"},{"1":"650.65","2":"16.97297","3":"14"},{"1":"639.25","2":"21.50087","3":"14"},{"1":"635.60","2":"20.60000","3":"14"},{"1":"666.55","2":"16.99029","3":"14"},{"1":"653.50","2":"20.77954","3":"14"},{"1":"647.25","2":"15.51247","3":"14"},{"1":"642.75","2":"19.88506","3":"14"},{"1":"625.85","2":"21.39882","3":"14"},{"1":"664.30","2":"20.49751","3":"14"},{"1":"657.55","2":"19.36376","3":"14"},{"1":"634.40","2":"17.65957","3":"14"},{"1":"633.90","2":"21.01796","3":"14"},{"1":"679.75","2":"19.05565","3":"14"},{"1":"672.70","2":"22.53846","3":"14"},{"1":"630.55","2":"21.10787","3":"14"},{"1":"644.45","2":"20.05135","3":"14"},{"1":"665.65","2":"14.20176","3":"14"},{"1":"627.10","2":"18.47687","3":"14"},{"1":"644.55","2":"18.63542","3":"14"},{"1":"639.75","2":"20.94595","3":"14"},{"1":"662.75","2":"21.08548","3":"14"},{"1":"619.80","2":"18.69288","3":"14"},{"1":"622.60","2":"20.86808","3":"14"},{"1":"666.85","2":"19.82558","3":"14"},{"1":"664.00","2":"19.75000","3":"14"},{"1":"628.75","2":"19.50000","3":"14"},{"1":"671.60","2":"18.39080","3":"14"},{"1":"672.45","2":"18.78676","3":"14"},{"1":"638.35","2":"19.77018","3":"14"},{"1":"661.60","2":"19.33333","3":"14"},{"1":"673.30","2":"21.46392","3":"14"},{"1":"623.60","2":"23.08492","3":"14"},{"1":"671.70","2":"21.06299","3":"14"},{"1":"652.30","2":"18.68687","3":"14"},{"1":"624.15","2":"20.77024","3":"14"},{"1":"670.00","2":"19.30556","3":"14"},{"1":"643.05","2":"20.13280","3":"14"},{"1":"654.20","2":"20.66964","3":"14"},{"1":"641.45","2":"22.28155","3":"14"},{"1":"679.50","2":"20.60027","3":"14"},{"1":"634.00","2":"20.82734","3":"14"},{"1":"654.30","2":"19.22492","3":"14"},{"1":"649.70","2":"17.65477","3":"14"},{"1":"649.15","2":"17.00000","3":"14"},{"1":"652.85","2":"16.49773","3":"14"},{"1":"666.15","2":"19.78261","3":"14"},{"1":"662.65","2":"22.30216","3":"14"},{"1":"683.40","2":"17.73077","3":"14"},{"1":"657.00","2":"20.44836","3":"14"},{"1":"665.10","2":"20.37169","3":"14"},{"1":"669.80","2":"20.16479","3":"14"},{"1":"651.85","2":"21.61538","3":"14"},{"1":"653.70","2":"20.56143","3":"14"},{"1":"652.35","2":"19.95551","3":"14"},{"1":"636.70","2":"21.18387","3":"14"},{"1":"638.20","2":"18.81042","3":"14"},{"1":"615.75","2":"20.57838","3":"14"},{"1":"634.95","2":"18.32461","3":"14"},{"1":"694.80","2":"18.82063","3":"14"},{"1":"661.35","2":"20.81633","3":"14"},{"1":"668.80","2":"20.00000","3":"14"},{"1":"644.95","2":"19.68182","3":"14"},{"1":"639.35","2":"19.39018","3":"14"},{"1":"676.95","2":"20.92732","3":"14"},{"1":"659.40","2":"19.94437","3":"14"},{"1":"660.95","2":"20.79109","3":"14"},{"1":"645.55","2":"19.20354","3":"14"},{"1":"626.10","2":"19.02439","3":"14"},{"1":"654.85","2":"17.62058","3":"14"},{"1":"633.65","2":"20.23715","3":"14"},{"1":"645.25","2":"19.29374","3":"14"},{"1":"637.35","2":"18.82998","3":"14"},{"1":"652.30","2":"20.33949","3":"14"},{"1":"641.80","2":"19.22900","3":"14"},{"1":"668.60","2":"17.89130","3":"14"},{"1":"661.90","2":"19.51881","3":"14"},{"1":"631.85","2":"19.08451","3":"14"},{"1":"616.30","2":"19.93548","3":"14"},{"1":"660.75","2":"18.87326","3":"14"},{"1":"668.00","2":"20.14178","3":"14"},{"1":"650.60","2":"23.55637","3":"14"},{"1":"661.85","2":"21.46479","3":"14"},{"1":"690.80","2":"19.19101","3":"14"},{"1":"651.15","2":"20.13080","3":"14"},{"1":"673.25","2":"25.80000","3":"14"},{"1":"650.55","2":"18.77774","3":"14"},{"1":"635.45","2":"19.10982","3":"14"},{"1":"654.20","2":"19.70109","3":"14"},{"1":"652.10","2":"18.61594","3":"14"},{"1":"643.70","2":"20.99721","3":"14"},{"1":"662.90","2":"20.00000","3":"14"},{"1":"672.20","2":"20.98325","3":"14"},{"1":"612.50","2":"21.64262","3":"14"},{"1":"634.20","2":"20.02967","3":"14"},{"1":"662.50","2":"19.81140","3":"14"},{"1":"646.55","2":"18.00000","3":"14"},{"1":"660.05","2":"19.35811","3":"14"},{"1":"634.90","2":"20.17912","3":"14"},{"1":"676.85","2":"21.11986","3":"14"},{"1":"656.80","2":"23.38974","3":"14"},{"1":"631.40","2":"22.18182","3":"14"},{"1":"618.30","2":"19.94283","3":"14"},{"1":"666.85","2":"17.78826","3":"14"},{"1":"637.95","2":"14.70588","3":"14"},{"1":"668.90","2":"19.04077","3":"14"},{"1":"657.50","2":"20.89195","3":"14"},{"1":"684.95","2":"19.83851","3":"14"},{"1":"671.90","2":"19.52191","3":"14"},{"1":"677.95","2":"20.68622","3":"14"},{"1":"632.85","2":"18.18182","3":"14"},{"1":"641.10","2":"18.89224","3":"14"},{"1":"678.80","2":"24.88889","3":"14"},{"1":"648.70","2":"18.58064","3":"14"},{"1":"634.70","2":"18.04000","3":"14"},{"1":"655.05","2":"17.73399","3":"14"},{"1":"665.35","2":"21.45455","3":"14"},{"1":"671.95","2":"19.92343","3":"14"},{"1":"626.80","2":"20.33942","3":"14"},{"1":"612.65","2":"22.54608","3":"14"},{"1":"681.30","2":"21.10344","3":"14"},{"1":"671.25","2":"18.19743","3":"14"},{"1":"654.85","2":"20.10768","3":"14"},{"1":"682.15","2":"19.15984","3":"14"},{"1":"679.80","2":"19.54545","3":"14"},{"1":"636.90","2":"20.88889","3":"14"},{"1":"652.10","2":"18.39150","3":"14"},{"1":"632.00","2":"19.17990","3":"14"},{"1":"660.20","2":"19.39771","3":"14"},{"1":"655.20","2":"21.67827","3":"14"},{"1":"665.95","2":"19.28889","3":"14"},{"1":"658.35","2":"20.34927","3":"14"},{"1":"667.20","2":"20.96416","3":"14"},{"1":"635.60","2":"19.46039","3":"14"},{"1":"693.95","2":"19.28572","3":"14"},{"1":"691.90","2":"20.91979","3":"14"},{"1":"625.30","2":"20.90021","3":"14"},{"1":"671.30","2":"20.59575","3":"14"},{"1":"655.75","2":"19.37500","3":"14"},{"1":"662.55","2":"19.95122","3":"14"},{"1":"669.35","2":"18.84973","3":"14"},{"1":"627.30","2":"18.11787","3":"14"},{"1":"646.35","2":"19.18341","3":"14"},{"1":"637.65","2":"22.00000","3":"14"},{"1":"662.40","2":"21.58416","3":"14"},{"1":"669.95","2":"20.38889","3":"14"},{"1":"650.90","2":"16.29310","3":"14"},{"1":"628.40","2":"18.27778","3":"14"},{"1":"668.65","2":"19.37472","3":"14"},{"1":"632.25","2":"18.90909","3":"14"},{"1":"671.75","2":"16.40693","3":"14"},{"1":"651.45","2":"15.59140","3":"14"},{"1":"658.80","2":"18.70694","3":"14"},{"1":"661.45","2":"18.32985","3":"14"},{"1":"644.20","2":"17.90235","3":"14"},{"1":"652.50","2":"18.91157","3":"14"},{"1":"651.20","2":"20.32497","3":"14"},{"1":"645.10","2":"20.02457","3":"14"},{"1":"634.55","2":"24.00000","3":"14"},{"1":"631.05","2":"17.60784","3":"14"},{"1":"640.75","2":"19.34853","3":"14"},{"1":"694.25","2":"19.67846","3":"14"},{"1":"656.65","2":"18.72861","3":"14"},{"1":"621.75","2":"15.88235","3":"14"},{"1":"667.45","2":"20.05491","3":"14"},{"1":"634.10","2":"17.98825","3":"14"},{"1":"654.10","2":"16.96629","3":"14"},{"1":"650.45","2":"19.23937","3":"14"},{"1":"634.10","2":"19.19586","3":"14"},{"1":"628.65","2":"19.59906","3":"14"},{"1":"662.55","2":"20.54348","3":"14"},{"1":"643.60","2":"18.58848","3":"14"},{"1":"698.25","2":"15.60419","3":"14"},{"1":"654.60","2":"15.29304","3":"14"},{"1":"664.40","2":"17.65537","3":"14"},{"1":"666.60","2":"17.57976","3":"14"},{"1":"666.65","2":"22.33333","3":"14"},{"1":"647.60","2":"18.75000","3":"14"},{"1":"634.05","2":"18.10241","3":"14"},{"1":"664.15","2":"20.25641","3":"14"},{"1":"655.30","2":"18.80207","3":"14"},{"1":"627.25","2":"18.77230","3":"14"},{"1":"669.85","2":"20.40521","3":"14"},{"1":"679.40","2":"18.65079","3":"14"},{"1":"639.80","2":"20.70707","3":"14"},{"1":"664.95","2":"22.00000","3":"14"},{"1":"664.15","2":"17.69978","3":"14"},{"1":"655.35","2":"21.48329","3":"14"},{"1":"623.20","2":"16.70103","3":"14"},{"1":"661.85","2":"19.57567","3":"14"},{"1":"653.80","2":"17.25806","3":"14"},{"1":"644.40","2":"17.37526","3":"14"},{"1":"640.10","2":"17.34931","3":"14"},{"1":"643.25","2":"16.26229","3":"14"},{"1":"644.45","2":"17.70045","3":"14"},{"1":"640.15","2":"20.12881","3":"14"},{"1":"661.20","2":"18.26539","3":"14"},{"1":"639.50","2":"14.54214","3":"14"},{"1":"662.45","2":"19.15261","3":"14"},{"1":"644.70","2":"17.36574","3":"14"},{"1":"680.45","2":"15.13898","3":"14"},{"1":"658.00","2":"17.84266","3":"14"},{"1":"663.45","2":"15.40704","3":"14"},{"1":"678.05","2":"18.86534","3":"14"},{"1":"635.05","2":"16.47413","3":"14"},{"1":"631.90","2":"17.86263","3":"14"},{"1":"689.10","2":"21.88586","3":"14"},{"1":"676.60","2":"20.20000","3":"14"},{"1":"645.75","2":"19.03640","3":"14"},{"1":"700.30","2":"17.88991","3":"15"},{"1":"654.20","2":"21.52466","3":"15"},{"1":"646.55","2":"18.69723","3":"15"},{"1":"658.00","2":"17.35714","3":"15"},{"1":"676.60","2":"18.67133","3":"15"},{"1":"694.25","2":"21.40625","3":"15"},{"1":"662.75","2":"19.50000","3":"15"},{"1":"646.70","2":"20.89412","3":"15"},{"1":"664.00","2":"19.94737","3":"15"},{"1":"644.20","2":"20.80556","3":"15"},{"1":"623.45","2":"21.23809","3":"15"},{"1":"664.15","2":"21.00000","3":"15"},{"1":"643.70","2":"20.60000","3":"15"},{"1":"690.80","2":"20.00822","3":"15"},{"1":"627.25","2":"18.02778","3":"15"},{"1":"664.40","2":"20.25196","3":"15"},{"1":"662.55","2":"16.97787","3":"15"},{"1":"643.20","2":"16.50980","3":"15"},{"1":"645.55","2":"22.70402","3":"15"},{"1":"659.40","2":"19.91111","3":"15"},{"1":"699.10","2":"18.33333","3":"15"},{"1":"635.60","2":"22.61905","3":"15"},{"1":"652.85","2":"19.44828","3":"15"},{"1":"676.55","2":"25.05263","3":"15"},{"1":"633.65","2":"20.67544","3":"15"},{"1":"668.95","2":"18.68235","3":"15"},{"1":"678.05","2":"22.84553","3":"15"},{"1":"682.45","2":"19.26667","3":"15"},{"1":"642.75","2":"19.25000","3":"15"},{"1":"624.55","2":"20.54545","3":"15"},{"1":"658.60","2":"20.60697","3":"15"},{"1":"626.80","2":"21.07268","3":"15"},{"1":"659.90","2":"21.53581","3":"15"},{"1":"662.40","2":"19.90400","3":"15"},{"1":"644.50","2":"21.19407","3":"15"},{"1":"682.15","2":"21.86535","3":"15"},{"1":"655.85","2":"18.32965","3":"15"},{"1":"650.60","2":"16.22857","3":"15"},{"1":"654.85","2":"19.17857","3":"15"},{"1":"616.45","2":"20.27737","3":"15"},{"1":"636.90","2":"22.98614","3":"15"},{"1":"659.05","2":"20.44444","3":"15"},{"1":"646.90","2":"19.82085","3":"15"},{"1":"616.30","2":"23.20522","3":"15"},{"1":"645.55","2":"19.26697","3":"15"},{"1":"671.90","2":"23.30189","3":"15"},{"1":"684.80","2":"21.18829","3":"15"},{"1":"669.35","2":"20.87180","3":"15"},{"1":"648.25","2":"19.01749","3":"15"},{"1":"684.30","2":"21.91938","3":"15"},{"1":"665.10","2":"20.10124","3":"15"},{"1":"632.20","2":"21.47651","3":"15"},{"1":"661.60","2":"20.06579","3":"15"},{"1":"664.70","2":"20.37510","3":"15"},{"1":"653.50","2":"22.44648","3":"15"},{"1":"687.55","2":"22.89524","3":"15"},{"1":"621.75","2":"20.49797","3":"15"},{"1":"642.40","2":"20.00000","3":"15"},{"1":"666.70","2":"22.25658","3":"15"},{"1":"663.85","2":"21.56436","3":"15"},{"1":"654.10","2":"19.47737","3":"15"},{"1":"645.25","2":"17.67002","3":"15"},{"1":"643.25","2":"21.94756","3":"15"},{"1":"650.90","2":"21.78339","3":"15"},{"1":"659.15","2":"19.14000","3":"15"},{"1":"660.10","2":"18.11050","3":"15"},{"1":"646.95","2":"20.68242","3":"15"},{"1":"655.70","2":"22.62361","3":"15"},{"1":"686.05","2":"21.78650","3":"15"},{"1":"673.30","2":"18.58293","3":"15"},{"1":"664.95","2":"21.54545","3":"15"},{"1":"639.85","2":"21.15289","3":"15"},{"1":"667.20","2":"16.63333","3":"15"},{"1":"661.85","2":"21.14438","3":"15"},{"1":"632.85","2":"19.78182","3":"15"},{"1":"665.95","2":"18.98373","3":"15"},{"1":"691.35","2":"17.66767","3":"15"},{"1":"649.70","2":"17.75499","3":"15"},{"1":"671.75","2":"15.27273","3":"15"},{"1":"665.35","2":"14.00000","3":"15"},{"1":"640.15","2":"20.59613","3":"15"},{"1":"680.45","2":"16.31169","3":"15"},{"1":"655.35","2":"21.12796","3":"15"},{"1":"679.50","2":"17.48801","3":"15"},{"1":"639.25","2":"17.88679","3":"15"},{"1":"698.45","2":"19.30676","3":"15"},{"1":"706.75","2":"20.89231","3":"15"},{"1":"634.15","2":"21.28684","3":"15"},{"1":"617.35","2":"20.19560","3":"15"},{"1":"645.75","2":"24.95000","3":"15"},{"1":"653.10","2":"18.13043","3":"15"},{"1":"693.95","2":"20.00000","3":"15"},{"1":"661.85","2":"18.72951","3":"15"},{"1":"623.20","2":"18.25000","3":"15"},{"1":"653.85","2":"18.99257","3":"15"},{"1":"682.55","2":"19.88764","3":"15"},{"1":"664.30","2":"19.37895","3":"15"},{"1":"647.60","2":"20.46259","3":"15"},{"1":"648.20","2":"22.29157","3":"15"},{"1":"640.10","2":"20.70474","3":"15"},{"1":"651.40","2":"19.06005","3":"15"},{"1":"643.40","2":"20.23247","3":"15"},{"1":"656.55","2":"19.69012","3":"15"},{"1":"679.80","2":"20.36254","3":"15"},{"1":"659.35","2":"19.75422","3":"15"},{"1":"634.55","2":"19.37977","3":"15"},{"1":"672.70","2":"22.92351","3":"15"},{"1":"681.60","2":"19.37340","3":"15"},{"1":"657.00","2":"19.15516","3":"15"},{"1":"649.30","2":"21.30000","3":"15"},{"1":"636.95","2":"18.30357","3":"15"},{"1":"651.15","2":"21.07926","3":"15"},{"1":"651.45","2":"18.79121","3":"15"},{"1":"616.30","2":"19.62662","3":"15"},{"1":"667.15","2":"19.59016","3":"15"},{"1":"657.80","2":"20.87187","3":"15"},{"1":"639.90","2":"21.11500","3":"15"},{"1":"639.75","2":"20.08452","3":"15"},{"1":"674.25","2":"19.91049","3":"15"},{"1":"641.55","2":"17.81285","3":"15"},{"1":"619.80","2":"18.13333","3":"15"},{"1":"670.00","2":"19.22221","3":"15"},{"1":"673.25","2":"18.66072","3":"15"},{"1":"666.05","2":"19.60000","3":"15"},{"1":"655.20","2":"19.28384","3":"15"},{"1":"649.15","2":"22.81818","3":"15"},{"1":"644.95","2":"18.80922","3":"15"},{"1":"659.80","2":"21.37363","3":"15"},{"1":"645.60","2":"20.02041","3":"15"},{"1":"637.65","2":"21.49862","3":"15"},{"1":"664.45","2":"15.42857","3":"15"},{"1":"652.40","2":"22.40000","3":"15"},{"1":"620.30","2":"20.12709","3":"15"},{"1":"612.50","2":"19.03798","3":"15"},{"1":"655.30","2":"17.34216","3":"15"},{"1":"655.05","2":"17.01863","3":"15"},{"1":"640.85","2":"20.80000","3":"15"},{"1":"665.20","2":"21.15385","3":"15"},{"1":"696.55","2":"18.45833","3":"15"},{"1":"653.95","2":"19.14082","3":"15"},{"1":"626.10","2":"19.40766","3":"15"},{"1":"653.55","2":"19.56896","3":"15"},{"1":"652.50","2":"21.50120","3":"15"},{"1":"655.75","2":"17.52941","3":"15"},{"1":"634.05","2":"16.43017","3":"15"},{"1":"661.85","2":"19.79654","3":"15"},{"1":"618.05","2":"17.18613","3":"15"},{"1":"675.40","2":"17.61589","3":"15"},{"1":"624.15","2":"20.12537","3":"15"},{"1":"637.10","2":"22.16667","3":"15"},{"1":"637.95","2":"19.96154","3":"15"},{"1":"640.75","2":"19.03945","3":"15"},{"1":"621.40","2":"15.22436","3":"15"},{"1":"631.85","2":"21.14475","3":"15"},{"1":"645.75","2":"19.64390","3":"15"},{"1":"673.05","2":"21.04869","3":"15"},{"1":"639.35","2":"20.17544","3":"15"},{"1":"655.05","2":"21.39130","3":"15"},{"1":"670.70","2":"20.00833","3":"15"},{"1":"647.70","2":"20.29137","3":"15"},{"1":"641.45","2":"17.66667","3":"15"},{"1":"672.05","2":"18.22055","3":"15"},{"1":"671.70","2":"20.27100","3":"15"},{"1":"633.90","2":"20.19895","3":"15"},{"1":"679.40","2":"21.38424","3":"15"},{"1":"677.95","2":"20.97368","3":"15"},{"1":"668.65","2":"20.00000","3":"15"},{"1":"656.40","2":"17.15328","3":"15"},{"1":"666.85","2":"22.34977","3":"15"},{"1":"646.40","2":"22.17007","3":"15"},{"1":"658.35","2":"18.18182","3":"15"},{"1":"661.35","2":"18.95714","3":"15"},{"1":"650.55","2":"19.74533","3":"15"},{"1":"644.40","2":"16.42623","3":"15"},{"1":"624.95","2":"16.62540","3":"15"},{"1":"634.90","2":"16.38177","3":"15"},{"1":"660.20","2":"20.07416","3":"15"},{"1":"652.30","2":"17.99544","3":"15"},{"1":"647.60","2":"19.39130","3":"15"},{"1":"668.80","2":"16.42857","3":"15"},{"1":"680.05","2":"16.72949","3":"15"},{"1":"644.70","2":"24.41345","3":"15"},{"1":"634.00","2":"18.26415","3":"15"},{"1":"622.05","2":"18.95504","3":"15"},{"1":"623.60","2":"21.03896","3":"15"},{"1":"665.90","2":"20.74074","3":"15"},{"1":"663.90","2":"18.10000","3":"15"},{"1":"672.45","2":"19.84615","3":"15"},{"1":"661.45","2":"21.60000","3":"15"},{"1":"660.30","2":"22.44242","3":"15"},{"1":"632.25","2":"23.01438","3":"15"},{"1":"615.75","2":"17.74892","3":"15"},{"1":"660.75","2":"18.28664","3":"15"},{"1":"629.80","2":"19.26544","3":"15"},{"1":"638.30","2":"22.66667","3":"15"},{"1":"668.10","2":"19.29412","3":"15"},{"1":"666.60","2":"17.36364","3":"15"},{"1":"671.60","2":"19.82143","3":"15"},{"1":"669.30","2":"20.43378","3":"15"},{"1":"657.75","2":"21.03721","3":"15"},{"1":"628.65","2":"19.92462","3":"15"},{"1":"669.95","2":"19.00986","3":"15"},{"1":"675.70","2":"23.82222","3":"15"},{"1":"694.80","2":"19.36909","3":"15"},{"1":"652.30","2":"19.82857","3":"15"},{"1":"642.20","2":"15.25885","3":"15"},{"1":"634.20","2":"17.16129","3":"15"},{"1":"662.90","2":"21.81333","3":"15"},{"1":"630.35","2":"19.07471","3":"15"},{"1":"634.10","2":"25.78512","3":"15"},{"1":"630.40","2":"18.21261","3":"15"},{"1":"679.75","2":"18.16606","3":"15"},{"1":"648.35","2":"16.97297","3":"15"},{"1":"681.30","2":"21.50087","3":"15"},{"1":"653.80","2":"20.60000","3":"15"},{"1":"646.50","2":"16.99029","3":"15"},{"1":"616.30","2":"20.77954","3":"15"},{"1":"657.90","2":"15.51247","3":"15"},{"1":"623.10","2":"19.88506","3":"15"},{"1":"635.45","2":"21.39882","3":"15"},{"1":"638.70","2":"20.49751","3":"15"},{"1":"681.90","2":"19.36376","3":"15"},{"1":"635.95","2":"17.65957","3":"15"},{"1":"635.60","2":"21.01796","3":"15"},{"1":"656.80","2":"19.05565","3":"15"},{"1":"653.70","2":"22.53846","3":"15"},{"1":"646.00","2":"21.10787","3":"15"},{"1":"683.40","2":"20.05135","3":"15"},{"1":"662.70","2":"14.20176","3":"15"},{"1":"666.85","2":"18.47687","3":"15"},{"1":"634.70","2":"18.63542","3":"15"},{"1":"631.95","2":"20.94595","3":"15"},{"1":"671.60","2":"21.08548","3":"15"},{"1":"633.05","2":"18.69288","3":"15"},{"1":"634.95","2":"20.86808","3":"15"},{"1":"654.60","2":"19.82558","3":"15"},{"1":"677.25","2":"19.75000","3":"15"},{"1":"672.55","2":"19.50000","3":"15"},{"1":"649.85","2":"18.39080","3":"15"},{"1":"639.30","2":"18.78676","3":"15"},{"1":"642.20","2":"19.77018","3":"15"},{"1":"668.90","2":"19.33333","3":"15"},{"1":"660.95","2":"21.46392","3":"15"},{"1":"669.80","2":"23.08492","3":"15"},{"1":"689.10","2":"21.06299","3":"15"},{"1":"657.15","2":"18.68687","3":"15"},{"1":"639.50","2":"20.77024","3":"15"},{"1":"632.45","2":"19.30556","3":"15"},{"1":"641.80","2":"20.13280","3":"15"},{"1":"628.55","2":"20.66964","3":"15"},{"1":"647.30","2":"22.28155","3":"15"},{"1":"669.30","2":"20.60027","3":"15"},{"1":"648.70","2":"20.82734","3":"15"},{"1":"647.25","2":"19.22492","3":"15"},{"1":"631.90","2":"17.65477","3":"15"},{"1":"648.00","2":"17.00000","3":"15"},{"1":"645.00","2":"16.49773","3":"15"},{"1":"667.45","2":"19.78261","3":"15"},{"1":"618.30","2":"22.30216","3":"15"},{"1":"650.90","2":"17.73077","3":"15"},{"1":"662.50","2":"20.44836","3":"15"},{"1":"637.00","2":"20.37169","3":"15"},{"1":"695.30","2":"20.16479","3":"15"},{"1":"691.90","2":"21.61538","3":"15"},{"1":"656.50","2":"20.56143","3":"15"},{"1":"620.50","2":"19.95551","3":"15"},{"1":"663.45","2":"21.18387","3":"15"},{"1":"673.55","2":"18.81042","3":"15"},{"1":"666.65","2":"20.57838","3":"15"},{"1":"663.35","2":"18.32461","3":"15"},{"1":"656.70","2":"18.82063","3":"15"},{"1":"636.50","2":"20.81633","3":"15"},{"1":"643.70","2":"20.00000","3":"15"},{"1":"672.05","2":"19.68182","3":"15"},{"1":"655.80","2":"19.39018","3":"15"},{"1":"654.30","2":"20.92732","3":"15"},{"1":"641.10","2":"19.94437","3":"15"},{"1":"652.10","2":"20.79109","3":"15"},{"1":"664.00","2":"19.20354","3":"15"},{"1":"671.30","2":"19.02439","3":"15"},{"1":"695.20","2":"17.62058","3":"15"},{"1":"662.65","2":"20.23715","3":"15"},{"1":"666.55","2":"19.29374","3":"15"},{"1":"658.80","2":"18.82998","3":"15"},{"1":"655.55","2":"20.33949","3":"15"},{"1":"684.35","2":"19.22900","3":"15"},{"1":"679.65","2":"17.89130","3":"15"},{"1":"644.45","2":"19.51881","3":"15"},{"1":"643.50","2":"19.08451","3":"15"},{"1":"631.40","2":"19.93548","3":"15"},{"1":"672.30","2":"18.87326","3":"15"},{"1":"659.40","2":"20.14178","3":"15"},{"1":"630.55","2":"23.55637","3":"15"},{"1":"654.85","2":"21.46479","3":"15"},{"1":"651.80","2":"19.19101","3":"15"},{"1":"626.90","2":"20.13080","3":"15"},{"1":"656.65","2":"25.80000","3":"15"},{"1":"628.75","2":"18.77774","3":"15"},{"1":"636.10","2":"19.10982","3":"15"},{"1":"638.35","2":"19.70109","3":"15"},{"1":"676.95","2":"18.61594","3":"15"},{"1":"627.10","2":"20.99721","3":"15"},{"1":"698.25","2":"20.00000","3":"15"},{"1":"625.85","2":"20.98325","3":"15"},{"1":"636.70","2":"21.64262","3":"15"},{"1":"643.05","2":"20.02967","3":"15"},{"1":"633.15","2":"19.81140","3":"15"},{"1":"662.55","2":"18.00000","3":"15"},{"1":"657.65","2":"19.35811","3":"15"},{"1":"637.95","2":"20.17912","3":"15"},{"1":"649.50","2":"21.11986","3":"15"},{"1":"671.95","2":"23.38974","3":"15"},{"1":"686.70","2":"22.18182","3":"15"},{"1":"664.15","2":"19.94283","3":"15"},{"1":"639.80","2":"17.78826","3":"15"},{"1":"644.55","2":"14.70588","3":"15"},{"1":"682.65","2":"19.04077","3":"15"},{"1":"671.90","2":"20.89195","3":"15"},{"1":"640.50","2":"19.83851","3":"15"},{"1":"632.00","2":"19.52191","3":"15"},{"1":"630.55","2":"20.68622","3":"15"},{"1":"651.20","2":"18.18182","3":"15"},{"1":"672.20","2":"18.89224","3":"15"},{"1":"661.90","2":"24.88889","3":"15"},{"1":"653.55","2":"18.58064","3":"15"},{"1":"651.35","2":"18.04000","3":"15"},{"1":"676.15","2":"17.73399","3":"15"},{"1":"667.60","2":"21.45455","3":"15"},{"1":"663.50","2":"19.92343","3":"15"},{"1":"666.10","2":"20.33942","3":"15"},{"1":"678.80","2":"22.54608","3":"15"},{"1":"635.05","2":"21.10344","3":"15"},{"1":"662.45","2":"18.19743","3":"15"},{"1":"643.60","2":"20.10768","3":"15"},{"1":"653.40","2":"19.15984","3":"15"},{"1":"638.20","2":"19.54545","3":"15"},{"1":"657.00","2":"20.88889","3":"15"},{"1":"625.30","2":"18.39150","3":"15"},{"1":"654.90","2":"19.17990","3":"15"},{"1":"667.45","2":"19.39771","3":"15"},{"1":"668.00","2":"21.67827","3":"15"},{"1":"666.15","2":"19.28889","3":"15"},{"1":"681.30","2":"20.34927","3":"15"},{"1":"605.55","2":"20.96416","3":"15"},{"1":"622.60","2":"19.46039","3":"15"},{"1":"650.45","2":"19.28572","3":"15"},{"1":"628.25","2":"20.91979","3":"15"},{"1":"634.10","2":"20.90021","3":"15"},{"1":"652.40","2":"20.59575","3":"15"},{"1":"648.95","2":"19.37500","3":"15"},{"1":"666.00","2":"19.95122","3":"15"},{"1":"663.85","2":"18.84973","3":"15"},{"1":"654.20","2":"18.11787","3":"15"},{"1":"640.90","2":"19.18341","3":"15"},{"1":"660.05","2":"22.00000","3":"15"},{"1":"650.65","2":"21.58416","3":"15"},{"1":"661.90","2":"20.38889","3":"15"},{"1":"634.40","2":"16.29310","3":"15"},{"1":"657.40","2":"18.27778","3":"15"},{"1":"661.20","2":"19.37472","3":"15"},{"1":"628.40","2":"18.90909","3":"15"},{"1":"683.35","2":"16.40693","3":"15"},{"1":"612.65","2":"15.59140","3":"15"},{"1":"676.85","2":"18.70694","3":"15"},{"1":"671.25","2":"18.32985","3":"15"},{"1":"671.65","2":"17.90235","3":"15"},{"1":"646.35","2":"18.91157","3":"15"},{"1":"652.00","2":"20.32497","3":"15"},{"1":"669.85","2":"20.02457","3":"15"},{"1":"606.75","2":"24.00000","3":"15"},{"1":"645.10","2":"17.60784","3":"15"},{"1":"704.30","2":"19.34853","3":"15"},{"1":"672.35","2":"19.67846","3":"15"},{"1":"635.75","2":"18.72861","3":"15"},{"1":"665.65","2":"15.88235","3":"15"},{"1":"652.35","2":"20.05491","3":"15"},{"1":"666.45","2":"17.98825","3":"15"},{"1":"666.65","2":"16.96629","3":"15"},{"1":"673.90","2":"19.23937","3":"15"},{"1":"641.45","2":"19.19586","3":"15"},{"1":"644.45","2":"19.59906","3":"15"},{"1":"662.40","2":"20.54348","3":"15"},{"1":"651.90","2":"18.58848","3":"15"},{"1":"655.40","2":"15.60419","3":"15"},{"1":"609.00","2":"15.29304","3":"15"},{"1":"638.55","2":"17.65537","3":"15"},{"1":"655.35","2":"17.57976","3":"15"},{"1":"644.20","2":"22.33333","3":"15"},{"1":"643.40","2":"18.75000","3":"15"},{"1":"661.60","2":"18.10241","3":"15"},{"1":"691.05","2":"20.25641","3":"15"},{"1":"661.95","2":"18.80207","3":"15"},{"1":"638.30","2":"18.77230","3":"15"},{"1":"698.20","2":"20.40521","3":"15"},{"1":"652.10","2":"18.65079","3":"15"},{"1":"668.60","2":"20.70707","3":"15"},{"1":"684.95","2":"22.00000","3":"15"},{"1":"635.20","2":"17.69978","3":"15"},{"1":"646.20","2":"21.48329","3":"15"},{"1":"673.55","2":"16.70103","3":"15"},{"1":"647.05","2":"19.57567","3":"15"},{"1":"651.85","2":"17.25806","3":"15"},{"1":"668.40","2":"17.37526","3":"15"},{"1":"657.50","2":"17.34931","3":"15"},{"1":"636.60","2":"16.26229","3":"15"},{"1":"664.75","2":"17.70045","3":"15"},{"1":"666.15","2":"20.12881","3":"15"},{"1":"627.30","2":"18.26539","3":"15"},{"1":"678.40","2":"14.54214","3":"15"},{"1":"669.10","2":"19.15261","3":"15"},{"1":"637.35","2":"17.36574","3":"15"},{"1":"638.00","2":"15.13898","3":"15"},{"1":"657.55","2":"17.84266","3":"15"},{"1":"656.80","2":"15.40704","3":"15"},{"1":"669.35","2":"18.86534","3":"15"},{"1":"655.05","2":"16.47413","3":"15"},{"1":"643.50","2":"17.86263","3":"15"},{"1":"664.95","2":"21.88586","3":"15"},{"1":"631.05","2":"20.20000","3":"15"},{"1":"632.95","2":"19.03640","3":"15"},{"1":"662.40","2":"17.88991","3":"16"},{"1":"668.80","2":"21.52466","3":"16"},{"1":"631.90","2":"18.69723","3":"16"},{"1":"639.35","2":"17.35714","3":"16"},{"1":"631.95","2":"18.67133","3":"16"},{"1":"632.85","2":"21.40625","3":"16"},{"1":"668.10","2":"19.50000","3":"16"},{"1":"660.75","2":"20.89412","3":"16"},{"1":"647.05","2":"19.94737","3":"16"},{"1":"631.05","2":"20.80556","3":"16"},{"1":"681.30","2":"21.23809","3":"16"},{"1":"650.90","2":"21.00000","3":"16"},{"1":"635.60","2":"20.60000","3":"16"},{"1":"664.15","2":"20.00822","3":"16"},{"1":"673.05","2":"18.02778","3":"16"},{"1":"654.85","2":"20.25196","3":"16"},{"1":"658.60","2":"16.97787","3":"16"},{"1":"643.50","2":"16.50980","3":"16"},{"1":"659.05","2":"22.70402","3":"16"},{"1":"634.40","2":"19.91111","3":"16"},{"1":"672.70","2":"18.33333","3":"16"},{"1":"671.60","2":"22.61905","3":"16"},{"1":"616.45","2":"19.44828","3":"16"},{"1":"666.70","2":"25.05263","3":"16"},{"1":"655.05","2":"20.67544","3":"16"},{"1":"678.80","2":"18.68235","3":"16"},{"1":"640.75","2":"22.84553","3":"16"},{"1":"671.30","2":"19.26667","3":"16"},{"1":"635.95","2":"19.25000","3":"16"},{"1":"646.40","2":"20.54545","3":"16"},{"1":"662.45","2":"20.60697","3":"16"},{"1":"669.95","2":"21.07268","3":"16"},{"1":"678.05","2":"21.53581","3":"16"},{"1":"652.85","2":"19.90400","3":"16"},{"1":"684.35","2":"21.19407","3":"16"},{"1":"652.00","2":"21.86535","3":"16"},{"1":"699.10","2":"18.32965","3":"16"},{"1":"679.40","2":"16.22857","3":"16"},{"1":"672.55","2":"19.17857","3":"16"},{"1":"665.90","2":"20.27737","3":"16"},{"1":"670.70","2":"22.98614","3":"16"},{"1":"630.55","2":"20.44444","3":"16"},{"1":"657.55","2":"19.82085","3":"16"},{"1":"630.40","2":"23.20522","3":"16"},{"1":"616.30","2":"19.26697","3":"16"},{"1":"659.35","2":"23.30189","3":"16"},{"1":"627.30","2":"21.18829","3":"16"},{"1":"654.20","2":"20.87180","3":"16"},{"1":"653.85","2":"19.01749","3":"16"},{"1":"662.40","2":"21.91938","3":"16"},{"1":"648.35","2":"20.10124","3":"16"},{"1":"648.25","2":"21.47651","3":"16"},{"1":"672.20","2":"20.06579","3":"16"},{"1":"621.40","2":"20.37510","3":"16"},{"1":"643.40","2":"22.44648","3":"16"},{"1":"655.20","2":"22.89524","3":"16"},{"1":"656.80","2":"20.49797","3":"16"},{"1":"626.10","2":"20.00000","3":"16"},{"1":"691.90","2":"22.25658","3":"16"},{"1":"683.35","2":"21.56436","3":"16"},{"1":"651.85","2":"19.47737","3":"16"},{"1":"661.45","2":"17.67002","3":"16"},{"1":"651.45","2":"21.94756","3":"16"},{"1":"652.50","2":"21.78339","3":"16"},{"1":"657.75","2":"19.14000","3":"16"},{"1":"681.60","2":"18.11050","3":"16"},{"1":"657.15","2":"20.68242","3":"16"},{"1":"639.75","2":"22.62361","3":"16"},{"1":"666.05","2":"21.78650","3":"16"},{"1":"641.45","2":"18.58293","3":"16"},{"1":"668.95","2":"21.54545","3":"16"},{"1":"660.95","2":"21.15289","3":"16"},{"1":"664.40","2":"16.63333","3":"16"},{"1":"647.60","2":"21.14438","3":"16"},{"1":"635.75","2":"19.78182","3":"16"},{"1":"651.40","2":"18.98373","3":"16"},{"1":"664.00","2":"17.66767","3":"16"},{"1":"638.00","2":"17.75499","3":"16"},{"1":"671.90","2":"15.27273","3":"16"},{"1":"651.80","2":"14.00000","3":"16"},{"1":"634.10","2":"20.59613","3":"16"},{"1":"623.45","2":"16.31169","3":"16"},{"1":"679.80","2":"21.12796","3":"16"},{"1":"669.35","2":"17.48801","3":"16"},{"1":"638.20","2":"17.88679","3":"16"},{"1":"662.70","2":"19.30676","3":"16"},{"1":"668.40","2":"20.89231","3":"16"},{"1":"654.85","2":"21.28684","3":"16"},{"1":"655.35","2":"20.19560","3":"16"},{"1":"638.70","2":"24.95000","3":"16"},{"1":"658.35","2":"18.13043","3":"16"},{"1":"666.00","2":"20.00000","3":"16"},{"1":"653.40","2":"18.72951","3":"16"},{"1":"643.60","2":"18.25000","3":"16"},{"1":"662.90","2":"18.99257","3":"16"},{"1":"657.50","2":"19.88764","3":"16"},{"1":"634.10","2":"19.37895","3":"16"},{"1":"634.55","2":"20.46259","3":"16"},{"1":"653.50","2":"22.29157","3":"16"},{"1":"654.10","2":"20.70474","3":"16"},{"1":"630.55","2":"19.06005","3":"16"},{"1":"695.20","2":"20.23247","3":"16"},{"1":"645.75","2":"19.69012","3":"16"},{"1":"655.35","2":"20.36254","3":"16"},{"1":"661.85","2":"19.75422","3":"16"},{"1":"671.60","2":"19.37977","3":"16"},{"1":"695.30","2":"22.92351","3":"16"},{"1":"650.60","2":"19.37340","3":"16"},{"1":"666.55","2":"19.15516","3":"16"},{"1":"658.80","2":"21.30000","3":"16"},{"1":"684.30","2":"18.30357","3":"16"},{"1":"626.80","2":"21.07926","3":"16"},{"1":"698.45","2":"18.79121","3":"16"},{"1":"662.65","2":"19.62662","3":"16"},{"1":"618.05","2":"19.59016","3":"16"},{"1":"663.50","2":"20.87187","3":"16"},{"1":"645.25","2":"21.11500","3":"16"},{"1":"609.00","2":"20.08452","3":"16"},{"1":"643.50","2":"19.91049","3":"16"},{"1":"661.20","2":"17.81285","3":"16"},{"1":"632.95","2":"18.13333","3":"16"},{"1":"623.20","2":"19.22221","3":"16"},{"1":"652.40","2":"18.66072","3":"16"},{"1":"654.60","2":"19.60000","3":"16"},{"1":"654.20","2":"19.28384","3":"16"},{"1":"636.10","2":"22.81818","3":"16"},{"1":"682.65","2":"18.80922","3":"16"},{"1":"682.15","2":"21.37363","3":"16"},{"1":"643.25","2":"20.02041","3":"16"},{"1":"643.70","2":"21.49862","3":"16"},{"1":"641.10","2":"15.42857","3":"16"},{"1":"654.90","2":"22.40000","3":"16"},{"1":"680.05","2":"20.12709","3":"16"},{"1":"666.85","2":"19.03798","3":"16"},{"1":"640.50","2":"17.34216","3":"16"},{"1":"646.95","2":"17.01863","3":"16"},{"1":"612.50","2":"20.80000","3":"16"},{"1":"671.95","2":"21.15385","3":"16"},{"1":"646.00","2":"18.45833","3":"16"},{"1":"621.75","2":"19.14082","3":"16"},{"1":"666.60","2":"19.40766","3":"16"},{"1":"680.45","2":"19.56896","3":"16"},{"1":"677.95","2":"21.50120","3":"16"},{"1":"625.30","2":"17.52941","3":"16"},{"1":"616.30","2":"16.43017","3":"16"},{"1":"628.65","2":"19.79654","3":"16"},{"1":"661.35","2":"17.18613","3":"16"},{"1":"652.35","2":"17.61589","3":"16"},{"1":"679.65","2":"20.12537","3":"16"},{"1":"638.55","2":"22.16667","3":"16"},{"1":"656.40","2":"19.96154","3":"16"},{"1":"629.80","2":"19.03945","3":"16"},{"1":"675.40","2":"15.22436","3":"16"},{"1":"666.15","2":"21.14475","3":"16"},{"1":"684.95","2":"19.64390","3":"16"},{"1":"657.40","2":"21.04869","3":"16"},{"1":"668.65","2":"20.17544","3":"16"},{"1":"677.25","2":"21.39130","3":"16"},{"1":"687.55","2":"20.00833","3":"16"},{"1":"666.85","2":"20.29137","3":"16"},{"1":"615.75","2":"17.66667","3":"16"},{"1":"666.10","2":"18.22055","3":"16"},{"1":"622.05","2":"20.27100","3":"16"},{"1":"663.90","2":"20.19895","3":"16"},{"1":"645.00","2":"21.38424","3":"16"},{"1":"639.50","2":"20.97368","3":"16"},{"1":"661.60","2":"20.00000","3":"16"},{"1":"669.10","2":"17.15328","3":"16"},{"1":"655.75","2":"22.34977","3":"16"},{"1":"619.80","2":"22.17007","3":"16"},{"1":"693.95","2":"18.18182","3":"16"},{"1":"643.05","2":"18.95714","3":"16"},{"1":"637.35","2":"19.74533","3":"16"},{"1":"639.30","2":"16.42623","3":"16"},{"1":"679.75","2":"16.62540","3":"16"},{"1":"648.00","2":"16.38177","3":"16"},{"1":"642.20","2":"20.07416","3":"16"},{"1":"627.10","2":"17.99544","3":"16"},{"1":"672.35","2":"19.39130","3":"16"},{"1":"659.80","2":"16.42857","3":"16"},{"1":"694.80","2":"16.72949","3":"16"},{"1":"628.55","2":"24.41345","3":"16"},{"1":"691.05","2":"18.26415","3":"16"},{"1":"651.20","2":"18.95504","3":"16"},{"1":"633.15","2":"21.03896","3":"16"},{"1":"646.55","2":"20.74074","3":"16"},{"1":"689.10","2":"18.10000","3":"16"},{"1":"647.30","2":"19.84615","3":"16"},{"1":"664.95","2":"21.60000","3":"16"},{"1":"632.00","2":"22.44242","3":"16"},{"1":"637.65","2":"23.01438","3":"16"},{"1":"670.00","2":"17.74892","3":"16"},{"1":"662.50","2":"18.28664","3":"16"},{"1":"651.15","2":"19.26544","3":"16"},{"1":"664.95","2":"22.66667","3":"16"},{"1":"682.55","2":"19.29412","3":"16"},{"1":"668.60","2":"17.36364","3":"16"},{"1":"649.70","2":"19.82143","3":"16"},{"1":"655.70","2":"20.43378","3":"16"},{"1":"620.30","2":"21.03721","3":"16"},{"1":"645.60","2":"19.92462","3":"16"},{"1":"633.90","2":"19.00986","3":"16"},{"1":"634.20","2":"23.82222","3":"16"},{"1":"656.50","2":"19.36909","3":"16"},{"1":"617.35","2":"19.82857","3":"16"},{"1":"645.55","2":"15.25885","3":"16"},{"1":"638.35","2":"17.16129","3":"16"},{"1":"635.05","2":"21.81333","3":"16"},{"1":"676.55","2":"19.07471","3":"16"},{"1":"651.90","2":"25.78512","3":"16"},{"1":"676.95","2":"18.21261","3":"16"},{"1":"691.35","2":"18.16606","3":"16"},{"1":"666.15","2":"16.97297","3":"16"},{"1":"669.30","2":"21.50087","3":"16"},{"1":"655.85","2":"20.60000","3":"16"},{"1":"633.65","2":"16.99029","3":"16"},{"1":"656.80","2":"20.77954","3":"16"},{"1":"673.90","2":"15.51247","3":"16"},{"1":"631.40","2":"19.88506","3":"16"},{"1":"640.90","2":"21.39882","3":"16"},{"1":"654.30","2":"20.49751","3":"16"},{"1":"665.10","2":"19.36376","3":"16"},{"1":"635.60","2":"17.65957","3":"16"},{"1":"650.65","2":"21.01796","3":"16"},{"1":"648.70","2":"19.05565","3":"16"},{"1":"672.45","2":"22.53846","3":"16"},{"1":"656.55","2":"21.10787","3":"16"},{"1":"663.85","2":"20.05135","3":"16"},{"1":"652.10","2":"14.20176","3":"16"},{"1":"645.10","2":"18.47687","3":"16"},{"1":"644.95","2":"18.63542","3":"16"},{"1":"684.80","2":"20.94595","3":"16"},{"1":"667.15","2":"21.08548","3":"16"},{"1":"647.25","2":"18.69288","3":"16"},{"1":"623.60","2":"20.86808","3":"16"},{"1":"661.95","2":"19.82558","3":"16"},{"1":"690.80","2":"19.75000","3":"16"},{"1":"635.45","2":"19.50000","3":"16"},{"1":"667.45","2":"18.39080","3":"16"},{"1":"649.30","2":"18.78676","3":"16"},{"1":"641.55","2":"19.77018","3":"16"},{"1":"639.25","2":"19.33333","3":"16"},{"1":"660.05","2":"21.46392","3":"16"},{"1":"665.95","2":"23.08492","3":"16"},{"1":"639.80","2":"21.06299","3":"16"},{"1":"669.35","2":"18.68687","3":"16"},{"1":"683.40","2":"20.77024","3":"16"},{"1":"649.50","2":"19.30556","3":"16"},{"1":"652.30","2":"20.13280","3":"16"},{"1":"671.65","2":"20.66964","3":"16"},{"1":"636.95","2":"22.28155","3":"16"},{"1":"658.00","2":"20.60027","3":"16"},{"1":"637.95","2":"20.82734","3":"16"},{"1":"661.90","2":"19.22492","3":"16"},{"1":"612.65","2":"17.65477","3":"16"},{"1":"653.55","2":"17.00000","3":"16"},{"1":"671.90","2":"16.49773","3":"16"},{"1":"643.40","2":"19.78261","3":"16"},{"1":"635.20","2":"22.30216","3":"16"},{"1":"661.85","2":"17.73077","3":"16"},{"1":"630.35","2":"20.44836","3":"16"},{"1":"664.00","2":"20.37169","3":"16"},{"1":"649.15","2":"20.16479","3":"16"},{"1":"662.75","2":"21.61538","3":"16"},{"1":"637.00","2":"20.56143","3":"16"},{"1":"624.95","2":"19.95551","3":"16"},{"1":"650.90","2":"21.18387","3":"16"},{"1":"632.20","2":"18.81042","3":"16"},{"1":"673.55","2":"20.57838","3":"16"},{"1":"657.90","2":"18.32461","3":"16"},{"1":"664.75","2":"18.82063","3":"16"},{"1":"671.75","2":"20.81633","3":"16"},{"1":"632.25","2":"20.00000","3":"16"},{"1":"661.85","2":"19.68182","3":"16"},{"1":"655.40","2":"19.39018","3":"16"},{"1":"660.30","2":"20.92732","3":"16"},{"1":"657.00","2":"19.94437","3":"16"},{"1":"706.75","2":"20.79109","3":"16"},{"1":"667.20","2":"19.20354","3":"16"},{"1":"634.15","2":"19.02439","3":"16"},{"1":"664.45","2":"17.62058","3":"16"},{"1":"694.25","2":"20.23715","3":"16"},{"1":"618.30","2":"19.29374","3":"16"},{"1":"650.55","2":"18.82998","3":"16"},{"1":"649.85","2":"20.33949","3":"16"},{"1":"638.30","2":"19.22900","3":"16"},{"1":"647.60","2":"17.89130","3":"16"},{"1":"648.95","2":"19.51881","3":"16"},{"1":"676.85","2":"19.08451","3":"16"},{"1":"676.60","2":"19.93548","3":"16"},{"1":"656.65","2":"18.87326","3":"16"},{"1":"662.55","2":"20.14178","3":"16"},{"1":"653.70","2":"23.55637","3":"16"},{"1":"636.50","2":"21.46479","3":"16"},{"1":"655.30","2":"19.19101","3":"16"},{"1":"640.15","2":"20.13080","3":"16"},{"1":"666.65","2":"25.80000","3":"16"},{"1":"666.65","2":"18.77774","3":"16"},{"1":"698.25","2":"19.10982","3":"16"},{"1":"657.65","2":"19.70109","3":"16"},{"1":"648.20","2":"18.61594","3":"16"},{"1":"627.25","2":"20.99721","3":"16"},{"1":"704.30","2":"20.00000","3":"16"},{"1":"634.70","2":"20.98325","3":"16"},{"1":"655.55","2":"21.64262","3":"16"},{"1":"644.20","2":"20.02967","3":"16"},{"1":"637.95","2":"19.81140","3":"16"},{"1":"678.40","2":"18.00000","3":"16"},{"1":"664.70","2":"19.35811","3":"16"},{"1":"657.00","2":"20.17912","3":"16"},{"1":"605.55","2":"21.11986","3":"16"},{"1":"660.10","2":"23.38974","3":"16"},{"1":"652.30","2":"22.18182","3":"16"},{"1":"664.15","2":"19.94283","3":"16"},{"1":"647.70","2":"17.78826","3":"16"},{"1":"659.40","2":"14.70588","3":"16"},{"1":"652.40","2":"19.04077","3":"16"},{"1":"644.55","2":"20.89195","3":"16"},{"1":"616.30","2":"19.83851","3":"16"},{"1":"672.30","2":"19.52191","3":"16"},{"1":"682.45","2":"20.68622","3":"16"},{"1":"639.85","2":"18.18182","3":"16"},{"1":"650.45","2":"18.89224","3":"16"},{"1":"686.05","2":"24.88889","3":"16"},{"1":"643.70","2":"18.58064","3":"16"},{"1":"626.90","2":"18.04000","3":"16"},{"1":"646.50","2":"17.73399","3":"16"},{"1":"620.50","2":"21.45455","3":"16"},{"1":"653.80","2":"19.92343","3":"16"},{"1":"686.70","2":"20.33942","3":"16"},{"1":"642.75","2":"22.54608","3":"16"},{"1":"668.00","2":"21.10344","3":"16"},{"1":"636.90","2":"18.19743","3":"16"},{"1":"624.55","2":"20.10768","3":"16"},{"1":"643.20","2":"19.15984","3":"16"},{"1":"642.40","2":"19.54545","3":"16"},{"1":"634.90","2":"20.88889","3":"16"},{"1":"636.70","2":"18.39150","3":"16"},{"1":"664.30","2":"19.17990","3":"16"},{"1":"651.35","2":"19.39771","3":"16"},{"1":"657.80","2":"21.67827","3":"16"},{"1":"644.50","2":"19.28889","3":"16"},{"1":"642.20","2":"20.34927","3":"16"},{"1":"653.95","2":"20.96416","3":"16"},{"1":"673.55","2":"19.46039","3":"16"},{"1":"628.75","2":"19.28572","3":"16"},{"1":"644.45","2":"20.91979","3":"16"},{"1":"656.70","2":"20.90021","3":"16"},{"1":"662.55","2":"20.59575","3":"16"},{"1":"669.30","2":"19.37500","3":"16"},{"1":"661.60","2":"19.95122","3":"16"},{"1":"700.30","2":"18.84973","3":"16"},{"1":"663.45","2":"18.11787","3":"16"},{"1":"622.60","2":"19.18341","3":"16"},{"1":"644.45","2":"22.00000","3":"16"},{"1":"628.40","2":"21.58416","3":"16"},{"1":"606.75","2":"20.38889","3":"16"},{"1":"672.05","2":"16.29310","3":"16"},{"1":"641.80","2":"18.27778","3":"16"},{"1":"673.25","2":"19.37472","3":"16"},{"1":"679.50","2":"18.90909","3":"16"},{"1":"671.25","2":"16.40693","3":"16"},{"1":"637.10","2":"15.59140","3":"16"},{"1":"681.90","2":"18.70694","3":"16"},{"1":"638.30","2":"18.32985","3":"16"},{"1":"644.20","2":"17.90235","3":"16"},{"1":"673.30","2":"18.91157","3":"16"},{"1":"665.20","2":"20.32497","3":"16"},{"1":"644.70","2":"20.02457","3":"16"},{"1":"659.40","2":"24.00000","3":"16"},{"1":"639.90","2":"17.60784","3":"16"},{"1":"663.35","2":"19.34853","3":"16"},{"1":"646.20","2":"19.67846","3":"16"},{"1":"634.05","2":"18.72861","3":"16"},{"1":"644.40","2":"15.88235","3":"16"},{"1":"659.15","2":"20.05491","3":"16"},{"1":"671.70","2":"17.98825","3":"16"},{"1":"669.80","2":"16.96629","3":"16"},{"1":"653.55","2":"19.23937","3":"16"},{"1":"633.05","2":"19.19586","3":"16"},{"1":"675.70","2":"19.59906","3":"16"},{"1":"646.90","2":"20.54348","3":"16"},{"1":"646.70","2":"18.58848","3":"16"},{"1":"631.85","2":"15.60419","3":"16"},{"1":"667.45","2":"15.29304","3":"16"},{"1":"674.25","2":"17.65537","3":"16"},{"1":"667.60","2":"17.57976","3":"16"},{"1":"660.20","2":"22.33333","3":"16"},{"1":"634.00","2":"18.75000","3":"16"},{"1":"659.90","2":"18.10241","3":"16"},{"1":"652.10","2":"20.25641","3":"16"},{"1":"668.90","2":"18.80207","3":"16"},{"1":"696.55","2":"18.77230","3":"16"},{"1":"640.10","2":"20.40521","3":"16"},{"1":"645.75","2":"18.65079","3":"16"},{"1":"640.85","2":"20.70707","3":"16"},{"1":"676.15","2":"22.00000","3":"16"},{"1":"698.20","2":"17.69978","3":"16"},{"1":"665.65","2":"21.48329","3":"16"},{"1":"669.85","2":"16.70103","3":"16"},{"1":"666.45","2":"19.57567","3":"16"},{"1":"655.80","2":"17.25806","3":"16"},{"1":"681.30","2":"17.37526","3":"16"},{"1":"661.90","2":"17.34931","3":"16"},{"1":"641.45","2":"16.26229","3":"16"},{"1":"628.25","2":"17.70045","3":"16"},{"1":"645.55","2":"20.12881","3":"16"},{"1":"663.85","2":"18.26539","3":"16"},{"1":"634.95","2":"14.54214","3":"16"},{"1":"646.35","2":"19.15261","3":"16"},{"1":"665.35","2":"17.36574","3":"16"},{"1":"636.60","2":"15.13898","3":"16"},{"1":"625.85","2":"17.84266","3":"16"},{"1":"623.10","2":"15.40704","3":"16"},{"1":"672.05","2":"18.86534","3":"16"},{"1":"624.15","2":"16.47413","3":"16"},{"1":"632.45","2":"17.86263","3":"16"},{"1":"653.10","2":"21.88586","3":"16"},{"1":"655.05","2":"20.20000","3":"16"},{"1":"655.05","2":"19.03640","3":"16"},{"1":"639.75","2":"17.88991","3":"17"},{"1":"634.10","2":"21.52466","3":"17"},{"1":"661.90","2":"18.69723","3":"17"},{"1":"642.20","2":"17.35714","3":"17"},{"1":"665.35","2":"18.67133","3":"17"},{"1":"669.35","2":"21.40625","3":"17"},{"1":"661.60","2":"19.50000","3":"17"},{"1":"658.00","2":"20.89412","3":"17"},{"1":"673.55","2":"19.94737","3":"17"},{"1":"649.70","2":"20.80556","3":"17"},{"1":"624.55","2":"21.23809","3":"17"},{"1":"681.30","2":"21.00000","3":"17"},{"1":"647.30","2":"20.60000","3":"17"},{"1":"646.35","2":"20.00822","3":"17"},{"1":"666.85","2":"18.02778","3":"17"},{"1":"655.05","2":"20.25196","3":"17"},{"1":"638.30","2":"16.97787","3":"17"},{"1":"635.20","2":"16.50980","3":"17"},{"1":"664.40","2":"22.70402","3":"17"},{"1":"696.55","2":"19.91111","3":"17"},{"1":"662.55","2":"18.33333","3":"17"},{"1":"689.10","2":"22.61905","3":"17"},{"1":"656.55","2":"19.44828","3":"17"},{"1":"648.95","2":"25.05263","3":"17"},{"1":"636.90","2":"20.67544","3":"17"},{"1":"664.30","2":"18.68235","3":"17"},{"1":"637.00","2":"22.84553","3":"17"},{"1":"621.40","2":"19.26667","3":"17"},{"1":"632.85","2":"19.25000","3":"17"},{"1":"664.95","2":"20.54545","3":"17"},{"1":"679.80","2":"20.60697","3":"17"},{"1":"653.85","2":"21.07268","3":"17"},{"1":"639.35","2":"21.53581","3":"17"},{"1":"653.80","2":"19.90400","3":"17"},{"1":"643.70","2":"21.19407","3":"17"},{"1":"628.25","2":"21.86535","3":"17"},{"1":"646.70","2":"18.32965","3":"17"},{"1":"648.25","2":"16.22857","3":"17"},{"1":"660.20","2":"19.17857","3":"17"},{"1":"646.40","2":"20.27737","3":"17"},{"1":"652.40","2":"22.98614","3":"17"},{"1":"657.90","2":"20.44444","3":"17"},{"1":"630.35","2":"19.82085","3":"17"},{"1":"644.20","2":"23.20522","3":"17"},{"1":"661.60","2":"19.26697","3":"17"},{"1":"651.45","2":"23.30189","3":"17"},{"1":"684.30","2":"21.18829","3":"17"},{"1":"657.80","2":"20.87180","3":"17"},{"1":"644.55","2":"19.01749","3":"17"},{"1":"655.40","2":"21.91938","3":"17"},{"1":"690.80","2":"20.10124","3":"17"},{"1":"673.30","2":"21.47651","3":"17"},{"1":"637.65","2":"20.06579","3":"17"},{"1":"677.25","2":"20.37510","3":"17"},{"1":"645.55","2":"22.44648","3":"17"},{"1":"605.55","2":"22.89524","3":"17"},{"1":"661.35","2":"20.49797","3":"17"},{"1":"659.80","2":"20.00000","3":"17"},{"1":"652.50","2":"22.25658","3":"17"},{"1":"653.10","2":"21.56436","3":"17"},{"1":"661.85","2":"19.47737","3":"17"},{"1":"666.15","2":"17.67002","3":"17"},{"1":"677.95","2":"21.94756","3":"17"},{"1":"626.90","2":"21.78339","3":"17"},{"1":"670.00","2":"19.14000","3":"17"},{"1":"617.35","2":"18.11050","3":"17"},{"1":"657.50","2":"20.68242","3":"17"},{"1":"682.15","2":"22.62361","3":"17"},{"1":"645.55","2":"21.78650","3":"17"},{"1":"631.90","2":"18.58293","3":"17"},{"1":"634.70","2":"21.54545","3":"17"},{"1":"647.60","2":"21.15289","3":"17"},{"1":"623.20","2":"16.63333","3":"17"},{"1":"661.95","2":"21.14438","3":"17"},{"1":"653.55","2":"19.78182","3":"17"},{"1":"631.95","2":"18.98373","3":"17"},{"1":"621.75","2":"17.66767","3":"17"},{"1":"706.75","2":"17.75499","3":"17"},{"1":"691.35","2":"15.27273","3":"17"},{"1":"639.80","2":"14.00000","3":"17"},{"1":"641.45","2":"20.59613","3":"17"},{"1":"666.05","2":"16.31169","3":"17"},{"1":"662.45","2":"21.12796","3":"17"},{"1":"646.95","2":"17.48801","3":"17"},{"1":"684.35","2":"17.88679","3":"17"},{"1":"660.95","2":"19.30676","3":"17"},{"1":"628.75","2":"20.89231","3":"17"},{"1":"655.70","2":"21.28684","3":"17"},{"1":"650.45","2":"20.19560","3":"17"},{"1":"616.45","2":"24.95000","3":"17"},{"1":"645.00","2":"18.13043","3":"17"},{"1":"649.30","2":"20.00000","3":"17"},{"1":"638.55","2":"18.72951","3":"17"},{"1":"672.05","2":"18.25000","3":"17"},{"1":"652.85","2":"18.99257","3":"17"},{"1":"656.70","2":"19.88764","3":"17"},{"1":"639.50","2":"19.37895","3":"17"},{"1":"665.65","2":"20.46259","3":"17"},{"1":"649.85","2":"22.29157","3":"17"},{"1":"647.05","2":"20.70474","3":"17"},{"1":"628.65","2":"19.06005","3":"17"},{"1":"691.90","2":"20.23247","3":"17"},{"1":"667.45","2":"19.69012","3":"17"},{"1":"633.90","2":"20.36254","3":"17"},{"1":"609.00","2":"19.75422","3":"17"},{"1":"647.60","2":"19.37977","3":"17"},{"1":"669.80","2":"22.92351","3":"17"},{"1":"675.40","2":"19.37340","3":"17"},{"1":"643.70","2":"19.15516","3":"17"},{"1":"648.70","2":"21.30000","3":"17"},{"1":"641.10","2":"18.30357","3":"17"},{"1":"660.30","2":"21.07926","3":"17"},{"1":"661.85","2":"18.79121","3":"17"},{"1":"652.30","2":"19.62662","3":"17"},{"1":"643.25","2":"19.59016","3":"17"},{"1":"635.60","2":"20.87187","3":"17"},{"1":"662.40","2":"21.11500","3":"17"},{"1":"636.10","2":"20.08452","3":"17"},{"1":"679.50","2":"19.91049","3":"17"},{"1":"630.40","2":"17.81285","3":"17"},{"1":"673.90","2":"18.13333","3":"17"},{"1":"612.65","2":"19.22221","3":"17"},{"1":"642.75","2":"18.66072","3":"17"},{"1":"660.05","2":"19.60000","3":"17"},{"1":"666.10","2":"19.28384","3":"17"},{"1":"665.10","2":"22.81818","3":"17"},{"1":"667.45","2":"18.80922","3":"17"},{"1":"666.55","2":"21.37363","3":"17"},{"1":"616.30","2":"20.02041","3":"17"},{"1":"646.00","2":"21.49862","3":"17"},{"1":"663.85","2":"15.42857","3":"17"},{"1":"634.40","2":"22.40000","3":"17"},{"1":"655.30","2":"20.12709","3":"17"},{"1":"640.15","2":"19.03798","3":"17"},{"1":"662.50","2":"17.34216","3":"17"},{"1":"622.05","2":"17.01863","3":"17"},{"1":"693.95","2":"20.80000","3":"17"},{"1":"634.55","2":"21.15385","3":"17"},{"1":"655.35","2":"18.45833","3":"17"},{"1":"669.30","2":"19.14082","3":"17"},{"1":"625.85","2":"19.40766","3":"17"},{"1":"635.95","2":"19.56896","3":"17"},{"1":"626.10","2":"21.50120","3":"17"},{"1":"615.75","2":"17.52941","3":"17"},{"1":"668.00","2":"16.43017","3":"17"},{"1":"643.60","2":"19.79654","3":"17"},{"1":"655.80","2":"17.18613","3":"17"},{"1":"673.05","2":"17.61589","3":"17"},{"1":"664.15","2":"20.12537","3":"17"},{"1":"662.90","2":"22.16667","3":"17"},{"1":"627.25","2":"19.96154","3":"17"},{"1":"653.50","2":"19.03945","3":"17"},{"1":"659.40","2":"15.22436","3":"17"},{"1":"680.05","2":"21.14475","3":"17"},{"1":"639.30","2":"19.64390","3":"17"},{"1":"698.20","2":"21.04869","3":"17"},{"1":"651.20","2":"20.17544","3":"17"},{"1":"666.00","2":"21.39130","3":"17"},{"1":"612.50","2":"20.00833","3":"17"},{"1":"646.90","2":"20.29137","3":"17"},{"1":"657.75","2":"17.66667","3":"17"},{"1":"636.70","2":"18.22055","3":"17"},{"1":"665.20","2":"20.27100","3":"17"},{"1":"638.00","2":"20.19895","3":"17"},{"1":"639.25","2":"21.38424","3":"17"},{"1":"625.30","2":"20.97368","3":"17"},{"1":"676.85","2":"20.00000","3":"17"},{"1":"684.95","2":"17.15328","3":"17"},{"1":"632.00","2":"22.34977","3":"17"},{"1":"660.75","2":"22.17007","3":"17"},{"1":"671.90","2":"18.18182","3":"17"},{"1":"672.55","2":"18.95714","3":"17"},{"1":"646.55","2":"19.74533","3":"17"},{"1":"679.75","2":"16.42623","3":"17"},{"1":"651.35","2":"16.62540","3":"17"},{"1":"651.90","2":"16.38177","3":"17"},{"1":"636.95","2":"20.07416","3":"17"},{"1":"654.90","2":"17.99544","3":"17"},{"1":"658.80","2":"19.39130","3":"17"},{"1":"657.15","2":"16.42857","3":"17"},{"1":"669.35","2":"16.72949","3":"17"},{"1":"679.65","2":"24.41345","3":"17"},{"1":"678.40","2":"18.26415","3":"17"},{"1":"649.15","2":"18.95504","3":"17"},{"1":"673.55","2":"21.03896","3":"17"},{"1":"644.50","2":"20.74074","3":"17"},{"1":"664.15","2":"18.10000","3":"17"},{"1":"651.40","2":"19.84615","3":"17"},{"1":"669.85","2":"21.60000","3":"17"},{"1":"622.60","2":"22.44242","3":"17"},{"1":"683.35","2":"23.01438","3":"17"},{"1":"699.10","2":"17.74892","3":"17"},{"1":"637.35","2":"18.28664","3":"17"},{"1":"656.50","2":"19.26544","3":"17"},{"1":"657.00","2":"22.66667","3":"17"},{"1":"676.95","2":"19.29412","3":"17"},{"1":"630.55","2":"17.36364","3":"17"},{"1":"664.95","2":"19.82143","3":"17"},{"1":"650.90","2":"20.43378","3":"17"},{"1":"668.95","2":"21.03721","3":"17"},{"1":"666.45","2":"19.92462","3":"17"},{"1":"653.70","2":"19.00986","3":"17"},{"1":"627.30","2":"23.82222","3":"17"},{"1":"648.35","2":"19.36909","3":"17"},{"1":"679.40","2":"19.82857","3":"17"},{"1":"633.65","2":"15.25885","3":"17"},{"1":"678.80","2":"17.16129","3":"17"},{"1":"664.00","2":"21.81333","3":"17"},{"1":"666.60","2":"19.07471","3":"17"},{"1":"668.80","2":"25.78512","3":"17"},{"1":"694.80","2":"18.21261","3":"17"},{"1":"632.45","2":"18.16606","3":"17"},{"1":"637.10","2":"16.97297","3":"17"},{"1":"656.80","2":"21.50087","3":"17"},{"1":"671.25","2":"20.60000","3":"17"},{"1":"674.25","2":"16.99029","3":"17"},{"1":"653.40","2":"20.77954","3":"17"},{"1":"686.70","2":"15.51247","3":"17"},{"1":"667.60","2":"19.88506","3":"17"},{"1":"631.40","2":"21.39882","3":"17"},{"1":"657.55","2":"20.49751","3":"17"},{"1":"669.30","2":"19.36376","3":"17"},{"1":"655.20","2":"17.65957","3":"17"},{"1":"643.50","2":"21.01796","3":"17"},{"1":"654.20","2":"19.05565","3":"17"},{"1":"666.85","2":"22.53846","3":"17"},{"1":"659.35","2":"21.10787","3":"17"},{"1":"644.45","2":"20.05135","3":"17"},{"1":"662.70","2":"14.20176","3":"17"},{"1":"640.85","2":"18.47687","3":"17"},{"1":"670.70","2":"18.63542","3":"17"},{"1":"643.50","2":"20.94595","3":"17"},{"1":"672.45","2":"21.08548","3":"17"},{"1":"641.55","2":"18.69288","3":"17"},{"1":"671.70","2":"20.86808","3":"17"},{"1":"672.20","2":"19.82558","3":"17"},{"1":"634.05","2":"19.75000","3":"17"},{"1":"663.50","2":"19.50000","3":"17"},{"1":"649.50","2":"18.39080","3":"17"},{"1":"620.50","2":"18.78676","3":"17"},{"1":"675.70","2":"19.77018","3":"17"},{"1":"661.85","2":"19.33333","3":"17"},{"1":"663.85","2":"21.46392","3":"17"},{"1":"666.15","2":"23.08492","3":"17"},{"1":"623.60","2":"21.06299","3":"17"},{"1":"624.15","2":"18.68687","3":"17"},{"1":"672.30","2":"20.77024","3":"17"},{"1":"644.95","2":"19.30556","3":"17"},{"1":"629.80","2":"20.13280","3":"17"},{"1":"666.70","2":"20.66964","3":"17"},{"1":"664.00","2":"22.28155","3":"17"},{"1":"663.90","2":"20.60027","3":"17"},{"1":"662.75","2":"20.82734","3":"17"},{"1":"652.30","2":"19.22492","3":"17"},{"1":"640.75","2":"17.65477","3":"17"},{"1":"645.75","2":"17.00000","3":"17"},{"1":"678.05","2":"16.49773","3":"17"},{"1":"664.70","2":"19.78261","3":"17"},{"1":"655.35","2":"22.30216","3":"17"},{"1":"654.60","2":"17.73077","3":"17"},{"1":"680.45","2":"20.44836","3":"17"},{"1":"655.05","2":"20.37169","3":"17"},{"1":"652.40","2":"20.16479","3":"17"},{"1":"652.10","2":"21.61538","3":"17"},{"1":"656.80","2":"20.56143","3":"17"},{"1":"634.95","2":"19.95551","3":"17"},{"1":"668.60","2":"21.18387","3":"17"},{"1":"655.85","2":"18.81042","3":"17"},{"1":"636.60","2":"20.57838","3":"17"},{"1":"654.10","2":"18.32461","3":"17"},{"1":"643.20","2":"18.82063","3":"17"},{"1":"638.70","2":"20.81633","3":"17"},{"1":"645.10","2":"20.00000","3":"17"},{"1":"628.40","2":"19.68182","3":"17"},{"1":"671.65","2":"19.39018","3":"17"},{"1":"652.00","2":"20.92732","3":"17"},{"1":"616.30","2":"19.94437","3":"17"},{"1":"645.75","2":"20.79109","3":"17"},{"1":"635.45","2":"19.20354","3":"17"},{"1":"654.30","2":"19.02439","3":"17"},{"1":"643.05","2":"17.62058","3":"17"},{"1":"668.90","2":"20.23715","3":"17"},{"1":"664.75","2":"19.29374","3":"17"},{"1":"632.95","2":"18.82998","3":"17"},{"1":"651.80","2":"20.33949","3":"17"},{"1":"654.85","2":"19.22900","3":"17"},{"1":"659.05","2":"17.89130","3":"17"},{"1":"647.70","2":"19.51881","3":"17"},{"1":"666.65","2":"19.08451","3":"17"},{"1":"646.50","2":"19.93548","3":"17"},{"1":"664.45","2":"18.87326","3":"17"},{"1":"655.75","2":"20.14178","3":"17"},{"1":"671.30","2":"23.55637","3":"17"},{"1":"662.40","2":"21.46479","3":"17"},{"1":"658.60","2":"19.19101","3":"17"},{"1":"657.65","2":"20.13080","3":"17"},{"1":"698.45","2":"25.80000","3":"17"},{"1":"653.55","2":"18.77774","3":"17"},{"1":"668.10","2":"19.10982","3":"17"},{"1":"694.25","2":"19.70109","3":"17"},{"1":"640.90","2":"18.61594","3":"17"},{"1":"641.45","2":"20.99721","3":"17"},{"1":"633.05","2":"20.00000","3":"17"},{"1":"635.60","2":"20.98325","3":"17"},{"1":"661.20","2":"21.64262","3":"17"},{"1":"650.60","2":"20.02967","3":"17"},{"1":"667.20","2":"19.81140","3":"17"},{"1":"672.05","2":"18.00000","3":"17"},{"1":"669.95","2":"19.35811","3":"17"},{"1":"652.35","2":"20.17912","3":"17"},{"1":"637.95","2":"21.11986","3":"17"},{"1":"643.40","2":"23.38974","3":"17"},{"1":"695.30","2":"22.18182","3":"17"},{"1":"645.60","2":"19.94283","3":"17"},{"1":"642.20","2":"17.78826","3":"17"},{"1":"644.45","2":"14.70588","3":"17"},{"1":"660.10","2":"19.04077","3":"17"},{"1":"618.05","2":"20.89195","3":"17"},{"1":"661.45","2":"19.83851","3":"17"},{"1":"638.30","2":"19.52191","3":"17"},{"1":"669.10","2":"20.68622","3":"17"},{"1":"651.85","2":"18.18182","3":"17"},{"1":"687.55","2":"18.89224","3":"17"},{"1":"639.90","2":"24.88889","3":"17"},{"1":"676.15","2":"18.58064","3":"17"},{"1":"638.20","2":"18.04000","3":"17"},{"1":"634.10","2":"17.73399","3":"17"},{"1":"666.65","2":"21.45455","3":"17"},{"1":"657.40","2":"19.92343","3":"17"},{"1":"704.30","2":"20.33942","3":"17"},{"1":"631.85","2":"22.54608","3":"17"},{"1":"639.85","2":"21.10344","3":"17"},{"1":"656.65","2":"18.19743","3":"17"},{"1":"663.35","2":"20.10768","3":"17"},{"1":"668.40","2":"19.15984","3":"17"},{"1":"682.65","2":"19.54545","3":"17"},{"1":"628.55","2":"20.88889","3":"17"},{"1":"637.95","2":"18.39150","3":"17"},{"1":"681.30","2":"19.17990","3":"17"},{"1":"665.95","2":"19.39771","3":"17"},{"1":"634.90","2":"21.67827","3":"17"},{"1":"630.55","2":"19.28889","3":"17"},{"1":"651.15","2":"20.34927","3":"17"},{"1":"663.45","2":"20.96416","3":"17"},{"1":"668.65","2":"19.46039","3":"17"},{"1":"681.90","2":"19.28572","3":"17"},{"1":"681.60","2":"20.91979","3":"17"},{"1":"698.25","2":"20.90021","3":"17"},{"1":"700.30","2":"20.59575","3":"17"},{"1":"661.90","2":"19.37500","3":"17"},{"1":"667.15","2":"19.95122","3":"17"},{"1":"650.65","2":"18.84973","3":"17"},{"1":"691.05","2":"18.11787","3":"17"},{"1":"633.15","2":"19.18341","3":"17"},{"1":"671.75","2":"22.00000","3":"17"},{"1":"648.20","2":"21.58416","3":"17"},{"1":"672.70","2":"20.38889","3":"17"},{"1":"623.45","2":"16.29310","3":"17"},{"1":"659.40","2":"18.27778","3":"17"},{"1":"648.00","2":"19.37472","3":"17"},{"1":"619.80","2":"18.90909","3":"17"},{"1":"647.25","2":"16.40693","3":"17"},{"1":"638.35","2":"15.59140","3":"17"},{"1":"644.20","2":"18.70694","3":"17"},{"1":"632.25","2":"18.32985","3":"17"},{"1":"606.75","2":"17.90235","3":"17"},{"1":"626.80","2":"18.91157","3":"17"},{"1":"656.40","2":"20.32497","3":"17"},{"1":"634.15","2":"20.02457","3":"17"},{"1":"671.90","2":"24.00000","3":"17"},{"1":"682.55","2":"17.60784","3":"17"},{"1":"650.90","2":"19.34853","3":"17"},{"1":"686.05","2":"19.67846","3":"17"},{"1":"635.05","2":"18.72861","3":"17"},{"1":"634.20","2":"15.88235","3":"17"},{"1":"632.20","2":"20.05491","3":"17"},{"1":"682.45","2":"17.98825","3":"17"},{"1":"624.95","2":"16.96629","3":"17"},{"1":"662.65","2":"19.23937","3":"17"},{"1":"616.30","2":"19.19586","3":"17"},{"1":"673.25","2":"19.59906","3":"17"},{"1":"644.40","2":"20.54348","3":"17"},{"1":"665.90","2":"18.58848","3":"17"},{"1":"645.25","2":"15.60419","3":"17"},{"1":"641.80","2":"15.29304","3":"17"},{"1":"659.90","2":"17.65537","3":"17"},{"1":"676.60","2":"17.57976","3":"17"},{"1":"640.50","2":"22.33333","3":"17"},{"1":"655.55","2":"18.75000","3":"17"},{"1":"671.95","2":"18.10241","3":"17"},{"1":"652.10","2":"20.25641","3":"17"},{"1":"618.30","2":"18.80207","3":"17"},{"1":"650.55","2":"18.77230","3":"17"},{"1":"671.60","2":"20.40521","3":"17"},{"1":"634.00","2":"18.65079","3":"17"},{"1":"627.10","2":"20.70707","3":"17"},{"1":"654.20","2":"22.00000","3":"17"},{"1":"657.00","2":"17.69978","3":"17"},{"1":"623.10","2":"21.48329","3":"17"},{"1":"662.55","2":"16.70103","3":"17"},{"1":"631.05","2":"19.57567","3":"17"},{"1":"646.20","2":"17.25806","3":"17"},{"1":"636.50","2":"17.37526","3":"17"},{"1":"642.40","2":"17.34931","3":"17"},{"1":"644.70","2":"16.26229","3":"17"},{"1":"659.15","2":"17.70045","3":"17"},{"1":"653.95","2":"20.12881","3":"17"},{"1":"658.35","2":"18.26539","3":"17"},{"1":"676.55","2":"14.54214","3":"17"},{"1":"672.35","2":"19.15261","3":"17"},{"1":"643.40","2":"17.36574","3":"17"},{"1":"684.80","2":"15.13898","3":"17"},{"1":"683.40","2":"17.84266","3":"17"},{"1":"620.30","2":"15.40704","3":"17"},{"1":"671.60","2":"18.86534","3":"17"},{"1":"655.05","2":"16.47413","3":"17"},{"1":"654.85","2":"17.86263","3":"17"},{"1":"635.75","2":"21.88586","3":"17"},{"1":"640.10","2":"20.20000","3":"17"},{"1":"695.20","2":"19.03640","3":"17"},{"1":"687.55","2":"17.88991","3":"18"},{"1":"649.15","2":"21.52466","3":"18"},{"1":"649.70","2":"18.69723","3":"18"},{"1":"642.75","2":"17.35714","3":"18"},{"1":"636.70","2":"18.67133","3":"18"},{"1":"666.05","2":"21.40625","3":"18"},{"1":"661.60","2":"19.50000","3":"18"},{"1":"682.55","2":"20.89412","3":"18"},{"1":"700.30","2":"19.94737","3":"18"},{"1":"606.75","2":"20.80556","3":"18"},{"1":"661.45","2":"21.23809","3":"18"},{"1":"645.75","2":"21.00000","3":"18"},{"1":"639.85","2":"20.60000","3":"18"},{"1":"673.30","2":"20.00822","3":"18"},{"1":"695.30","2":"18.02778","3":"18"},{"1":"646.70","2":"20.25196","3":"18"},{"1":"646.90","2":"16.97787","3":"18"},{"1":"664.15","2":"16.50980","3":"18"},{"1":"671.60","2":"22.70402","3":"18"},{"1":"656.80","2":"19.91111","3":"18"},{"1":"666.65","2":"18.33333","3":"18"},{"1":"641.10","2":"22.61905","3":"18"},{"1":"617.35","2":"19.44828","3":"18"},{"1":"647.60","2":"25.05263","3":"18"},{"1":"661.90","2":"20.67544","3":"18"},{"1":"682.65","2":"18.68235","3":"18"},{"1":"663.85","2":"22.84553","3":"18"},{"1":"665.35","2":"19.26667","3":"18"},{"1":"640.10","2":"19.25000","3":"18"},{"1":"659.15","2":"20.54545","3":"18"},{"1":"633.05","2":"20.60697","3":"18"},{"1":"659.90","2":"21.07268","3":"18"},{"1":"624.15","2":"21.53581","3":"18"},{"1":"637.95","2":"19.90400","3":"18"},{"1":"673.55","2":"21.19407","3":"18"},{"1":"657.00","2":"21.86535","3":"18"},{"1":"650.90","2":"18.32965","3":"18"},{"1":"637.65","2":"16.22857","3":"18"},{"1":"659.05","2":"19.17857","3":"18"},{"1":"664.45","2":"20.27737","3":"18"},{"1":"669.30","2":"22.98614","3":"18"},{"1":"691.35","2":"20.44444","3":"18"},{"1":"645.55","2":"19.82085","3":"18"},{"1":"684.30","2":"23.20522","3":"18"},{"1":"661.85","2":"19.26697","3":"18"},{"1":"651.40","2":"23.30189","3":"18"},{"1":"612.65","2":"21.18829","3":"18"},{"1":"651.15","2":"20.87180","3":"18"},{"1":"638.55","2":"19.01749","3":"18"},{"1":"640.50","2":"21.91938","3":"18"},{"1":"655.85","2":"20.10124","3":"18"},{"1":"662.65","2":"21.47651","3":"18"},{"1":"634.15","2":"20.06579","3":"18"},{"1":"620.30","2":"20.37510","3":"18"},{"1":"639.75","2":"22.44648","3":"18"},{"1":"672.05","2":"22.89524","3":"18"},{"1":"632.45","2":"20.49797","3":"18"},{"1":"667.60","2":"20.00000","3":"18"},{"1":"668.40","2":"22.25658","3":"18"},{"1":"652.30","2":"21.56436","3":"18"},{"1":"658.60","2":"19.47737","3":"18"},{"1":"651.45","2":"17.67002","3":"18"},{"1":"659.40","2":"21.94756","3":"18"},{"1":"657.65","2":"21.78339","3":"18"},{"1":"653.55","2":"19.14000","3":"18"},{"1":"657.80","2":"18.11050","3":"18"},{"1":"649.30","2":"20.68242","3":"18"},{"1":"639.30","2":"22.62361","3":"18"},{"1":"616.30","2":"21.78650","3":"18"},{"1":"634.90","2":"18.58293","3":"18"},{"1":"667.20","2":"21.54545","3":"18"},{"1":"684.80","2":"21.15289","3":"18"},{"1":"676.15","2":"16.63333","3":"18"},{"1":"661.60","2":"21.14438","3":"18"},{"1":"669.30","2":"19.78182","3":"18"},{"1":"635.95","2":"18.98373","3":"18"},{"1":"640.15","2":"17.66767","3":"18"},{"1":"656.40","2":"17.75499","3":"18"},{"1":"679.75","2":"15.27273","3":"18"},{"1":"622.60","2":"14.00000","3":"18"},{"1":"683.40","2":"20.59613","3":"18"},{"1":"657.50","2":"16.31169","3":"18"},{"1":"668.90","2":"21.12796","3":"18"},{"1":"654.60","2":"17.48801","3":"18"},{"1":"643.40","2":"17.88679","3":"18"},{"1":"646.35","2":"19.30676","3":"18"},{"1":"619.80","2":"20.89231","3":"18"},{"1":"630.40","2":"21.28684","3":"18"},{"1":"652.10","2":"20.19560","3":"18"},{"1":"646.40","2":"24.95000","3":"18"},{"1":"631.85","2":"18.13043","3":"18"},{"1":"671.90","2":"20.00000","3":"18"},{"1":"643.05","2":"18.72951","3":"18"},{"1":"649.85","2":"18.25000","3":"18"},{"1":"671.70","2":"18.99257","3":"18"},{"1":"672.55","2":"19.88764","3":"18"},{"1":"644.40","2":"19.37895","3":"18"},{"1":"667.45","2":"20.46259","3":"18"},{"1":"660.75","2":"22.29157","3":"18"},{"1":"652.10","2":"20.70474","3":"18"},{"1":"635.75","2":"19.06005","3":"18"},{"1":"659.40","2":"20.23247","3":"18"},{"1":"666.85","2":"19.69012","3":"18"},{"1":"664.75","2":"20.36254","3":"18"},{"1":"638.30","2":"19.75422","3":"18"},{"1":"657.15","2":"19.37977","3":"18"},{"1":"694.25","2":"22.92351","3":"18"},{"1":"661.90","2":"19.37340","3":"18"},{"1":"633.90","2":"19.15516","3":"18"},{"1":"671.60","2":"21.30000","3":"18"},{"1":"696.55","2":"18.30357","3":"18"},{"1":"638.30","2":"21.07926","3":"18"},{"1":"635.20","2":"18.79121","3":"18"},{"1":"651.80","2":"19.62662","3":"18"},{"1":"646.00","2":"19.59016","3":"18"},{"1":"673.25","2":"20.87187","3":"18"},{"1":"665.10","2":"21.11500","3":"18"},{"1":"672.20","2":"20.08452","3":"18"},{"1":"684.35","2":"19.91049","3":"18"},{"1":"625.85","2":"17.81285","3":"18"},{"1":"662.40","2":"18.13333","3":"18"},{"1":"640.75","2":"19.22221","3":"18"},{"1":"663.50","2":"18.66072","3":"18"},{"1":"671.95","2":"19.60000","3":"18"},{"1":"672.35","2":"19.28384","3":"18"},{"1":"633.65","2":"22.81818","3":"18"},{"1":"641.45","2":"18.80922","3":"18"},{"1":"666.00","2":"21.37363","3":"18"},{"1":"618.05","2":"20.02041","3":"18"},{"1":"627.30","2":"21.49862","3":"18"},{"1":"655.35","2":"15.42857","3":"18"},{"1":"648.25","2":"22.40000","3":"18"},{"1":"664.15","2":"20.12709","3":"18"},{"1":"639.25","2":"19.03798","3":"18"},{"1":"652.30","2":"17.34216","3":"18"},{"1":"623.10","2":"17.01863","3":"18"},{"1":"698.20","2":"20.80000","3":"18"},{"1":"686.05","2":"21.15385","3":"18"},{"1":"672.70","2":"18.45833","3":"18"},{"1":"643.70","2":"19.14082","3":"18"},{"1":"640.85","2":"19.40766","3":"18"},{"1":"634.20","2":"19.56896","3":"18"},{"1":"630.55","2":"21.50120","3":"18"},{"1":"654.85","2":"17.52941","3":"18"},{"1":"665.65","2":"16.43017","3":"18"},{"1":"624.55","2":"19.79654","3":"18"},{"1":"679.80","2":"17.18613","3":"18"},{"1":"662.55","2":"17.61589","3":"18"},{"1":"666.60","2":"20.12537","3":"18"},{"1":"662.70","2":"22.16667","3":"18"},{"1":"628.75","2":"19.96154","3":"18"},{"1":"621.40","2":"19.03945","3":"18"},{"1":"645.75","2":"15.22436","3":"18"},{"1":"636.60","2":"21.14475","3":"18"},{"1":"666.15","2":"19.64390","3":"18"},{"1":"694.80","2":"21.04869","3":"18"},{"1":"664.40","2":"20.17544","3":"18"},{"1":"661.35","2":"21.39130","3":"18"},{"1":"637.00","2":"20.00833","3":"18"},{"1":"651.20","2":"20.29137","3":"18"},{"1":"631.05","2":"17.66667","3":"18"},{"1":"644.50","2":"18.22055","3":"18"},{"1":"652.85","2":"20.27100","3":"18"},{"1":"658.80","2":"20.19895","3":"18"},{"1":"626.10","2":"21.38424","3":"18"},{"1":"644.55","2":"20.97368","3":"18"},{"1":"672.30","2":"20.00000","3":"18"},{"1":"634.40","2":"17.15328","3":"18"},{"1":"647.25","2":"22.34977","3":"18"},{"1":"609.00","2":"22.17007","3":"18"},{"1":"632.95","2":"18.18182","3":"18"},{"1":"657.40","2":"18.95714","3":"18"},{"1":"668.10","2":"19.74533","3":"18"},{"1":"698.25","2":"16.42623","3":"18"},{"1":"660.10","2":"16.62540","3":"18"},{"1":"616.30","2":"16.38177","3":"18"},{"1":"636.50","2":"20.07416","3":"18"},{"1":"644.45","2":"17.99544","3":"18"},{"1":"656.70","2":"19.39130","3":"18"},{"1":"690.80","2":"16.42857","3":"18"},{"1":"650.65","2":"16.72949","3":"18"},{"1":"648.20","2":"24.41345","3":"18"},{"1":"680.45","2":"18.26415","3":"18"},{"1":"623.60","2":"18.95504","3":"18"},{"1":"656.65","2":"21.03896","3":"18"},{"1":"622.05","2":"20.74074","3":"18"},{"1":"669.85","2":"18.10000","3":"18"},{"1":"657.90","2":"19.84615","3":"18"},{"1":"656.50","2":"21.60000","3":"18"},{"1":"634.10","2":"22.44242","3":"18"},{"1":"683.35","2":"23.01438","3":"18"},{"1":"655.70","2":"17.74892","3":"18"},{"1":"639.50","2":"18.28664","3":"18"},{"1":"659.80","2":"19.26544","3":"18"},{"1":"657.75","2":"22.66667","3":"18"},{"1":"658.00","2":"19.29412","3":"18"},{"1":"673.90","2":"17.36364","3":"18"},{"1":"664.95","2":"19.82143","3":"18"},{"1":"639.90","2":"20.43378","3":"18"},{"1":"615.75","2":"21.03721","3":"18"},{"1":"653.55","2":"19.92462","3":"18"},{"1":"647.05","2":"19.00986","3":"18"},{"1":"628.55","2":"23.82222","3":"18"},{"1":"671.75","2":"19.36909","3":"18"},{"1":"689.10","2":"19.82857","3":"18"},{"1":"638.35","2":"15.25885","3":"18"},{"1":"655.30","2":"17.16129","3":"18"},{"1":"641.55","2":"21.81333","3":"18"},{"1":"650.45","2":"19.07471","3":"18"},{"1":"653.50","2":"25.78512","3":"18"},{"1":"628.65","2":"18.21261","3":"18"},{"1":"668.60","2":"18.16606","3":"18"},{"1":"643.50","2":"16.97297","3":"18"},{"1":"669.95","2":"21.50087","3":"18"},{"1":"616.45","2":"20.60000","3":"18"},{"1":"669.35","2":"16.99029","3":"18"},{"1":"637.10","2":"20.77954","3":"18"},{"1":"655.20","2":"15.51247","3":"18"},{"1":"626.80","2":"19.88506","3":"18"},{"1":"656.55","2":"21.39882","3":"18"},{"1":"632.25","2":"20.49751","3":"18"},{"1":"642.20","2":"19.36376","3":"18"},{"1":"643.40","2":"17.65957","3":"18"},{"1":"662.45","2":"21.01796","3":"18"},{"1":"646.50","2":"19.05565","3":"18"},{"1":"640.90","2":"22.53846","3":"18"},{"1":"682.45","2":"21.10787","3":"18"},{"1":"679.40","2":"20.05135","3":"18"},{"1":"635.05","2":"14.20176","3":"18"},{"1":"663.90","2":"18.47687","3":"18"},{"1":"679.50","2":"18.63542","3":"18"},{"1":"620.50","2":"20.94595","3":"18"},{"1":"628.40","2":"21.08548","3":"18"},{"1":"691.05","2":"18.69288","3":"18"},{"1":"630.35","2":"20.86808","3":"18"},{"1":"652.40","2":"19.82558","3":"18"},{"1":"681.30","2":"19.75000","3":"18"},{"1":"706.75","2":"19.50000","3":"18"},{"1":"636.95","2":"18.39080","3":"18"},{"1":"642.40","2":"18.78676","3":"18"},{"1":"676.60","2":"19.77018","3":"18"},{"1":"641.45","2":"19.33333","3":"18"},{"1":"655.05","2":"21.46392","3":"18"},{"1":"645.00","2":"23.08492","3":"18"},{"1":"660.95","2":"21.06299","3":"18"},{"1":"658.35","2":"18.68687","3":"18"},{"1":"673.05","2":"20.77024","3":"18"},{"1":"678.80","2":"19.30556","3":"18"},{"1":"631.40","2":"20.13280","3":"18"},{"1":"671.25","2":"20.66964","3":"18"},{"1":"652.50","2":"22.28155","3":"18"},{"1":"678.05","2":"20.60027","3":"18"},{"1":"631.90","2":"20.82734","3":"18"},{"1":"644.70","2":"19.22492","3":"18"},{"1":"647.30","2":"17.65477","3":"18"},{"1":"672.05","2":"17.00000","3":"18"},{"1":"634.05","2":"16.49773","3":"18"},{"1":"643.60","2":"19.78261","3":"18"},{"1":"665.90","2":"22.30216","3":"18"},{"1":"699.10","2":"17.73077","3":"18"},{"1":"673.55","2":"20.44836","3":"18"},{"1":"653.70","2":"20.37169","3":"18"},{"1":"652.40","2":"20.16479","3":"18"},{"1":"655.40","2":"21.61538","3":"18"},{"1":"672.45","2":"20.56143","3":"18"},{"1":"651.90","2":"19.95551","3":"18"},{"1":"667.15","2":"21.18387","3":"18"},{"1":"643.70","2":"18.81042","3":"18"},{"1":"632.85","2":"20.57838","3":"18"},{"1":"653.95","2":"18.32461","3":"18"},{"1":"662.40","2":"18.82063","3":"18"},{"1":"664.70","2":"20.81633","3":"18"},{"1":"624.95","2":"20.00000","3":"18"},{"1":"664.30","2":"19.68182","3":"18"},{"1":"655.75","2":"19.39018","3":"18"},{"1":"627.10","2":"20.92732","3":"18"},{"1":"654.90","2":"19.94437","3":"18"},{"1":"638.00","2":"20.79109","3":"18"},{"1":"623.45","2":"19.20354","3":"18"},{"1":"653.40","2":"19.02439","3":"18"},{"1":"634.10","2":"17.62058","3":"18"},{"1":"675.70","2":"20.23715","3":"18"},{"1":"666.45","2":"19.29374","3":"18"},{"1":"662.50","2":"18.82998","3":"18"},{"1":"684.95","2":"20.33949","3":"18"},{"1":"650.55","2":"19.22900","3":"18"},{"1":"704.30","2":"17.89130","3":"18"},{"1":"668.00","2":"19.51881","3":"18"},{"1":"652.35","2":"19.08451","3":"18"},{"1":"637.95","2":"19.93548","3":"18"},{"1":"664.95","2":"18.87326","3":"18"},{"1":"634.00","2":"20.14178","3":"18"},{"1":"659.35","2":"23.55637","3":"18"},{"1":"678.40","2":"21.46479","3":"18"},{"1":"634.70","2":"19.19101","3":"18"},{"1":"670.00","2":"20.13080","3":"18"},{"1":"671.90","2":"25.80000","3":"18"},{"1":"654.20","2":"18.77774","3":"18"},{"1":"681.30","2":"19.10982","3":"18"},{"1":"632.20","2":"19.70109","3":"18"},{"1":"634.55","2":"18.61594","3":"18"},{"1":"653.85","2":"20.99721","3":"18"},{"1":"663.85","2":"20.00000","3":"18"},{"1":"661.20","2":"20.98325","3":"18"},{"1":"676.85","2":"21.64262","3":"18"},{"1":"657.00","2":"20.02967","3":"18"},{"1":"641.80","2":"19.81140","3":"18"},{"1":"676.55","2":"18.00000","3":"18"},{"1":"653.10","2":"19.35811","3":"18"},{"1":"655.55","2":"20.17912","3":"18"},{"1":"655.35","2":"21.11986","3":"18"},{"1":"691.90","2":"23.38974","3":"18"},{"1":"660.05","2":"22.18182","3":"18"},{"1":"693.95","2":"19.94283","3":"18"},{"1":"651.85","2":"17.78826","3":"18"},{"1":"644.45","2":"14.70588","3":"18"},{"1":"698.45","2":"19.04077","3":"18"},{"1":"635.45","2":"20.89195","3":"18"},{"1":"636.10","2":"19.83851","3":"18"},{"1":"664.00","2":"19.52191","3":"18"},{"1":"623.20","2":"20.68622","3":"18"},{"1":"642.20","2":"18.18182","3":"18"},{"1":"654.20","2":"18.89224","3":"18"},{"1":"681.90","2":"24.88889","3":"18"},{"1":"665.20","2":"18.58064","3":"18"},{"1":"647.70","2":"18.04000","3":"18"},{"1":"648.35","2":"17.73399","3":"18"},{"1":"652.00","2":"21.45455","3":"18"},{"1":"635.60","2":"19.92343","3":"18"},{"1":"663.35","2":"20.33942","3":"18"},{"1":"664.00","2":"22.54608","3":"18"},{"1":"669.80","2":"21.10344","3":"18"},{"1":"666.85","2":"18.19743","3":"18"},{"1":"628.25","2":"20.10768","3":"18"},{"1":"663.45","2":"19.15984","3":"18"},{"1":"645.25","2":"19.54545","3":"18"},{"1":"644.95","2":"20.88889","3":"18"},{"1":"654.30","2":"18.39150","3":"18"},{"1":"679.65","2":"19.17990","3":"18"},{"1":"680.05","2":"19.39771","3":"18"},{"1":"682.15","2":"21.67827","3":"18"},{"1":"665.95","2":"19.28889","3":"18"},{"1":"626.90","2":"20.34927","3":"18"},{"1":"644.20","2":"20.96416","3":"18"},{"1":"660.20","2":"19.46039","3":"18"},{"1":"668.65","2":"19.28572","3":"18"},{"1":"612.50","2":"20.91979","3":"18"},{"1":"676.95","2":"20.90021","3":"18"},{"1":"662.55","2":"20.59575","3":"18"},{"1":"661.85","2":"19.37500","3":"18"},{"1":"647.60","2":"19.95122","3":"18"},{"1":"649.50","2":"18.84973","3":"18"},{"1":"643.25","2":"18.11787","3":"18"},{"1":"666.10","2":"19.18341","3":"18"},{"1":"662.90","2":"22.00000","3":"18"},{"1":"621.75","2":"21.58416","3":"18"},{"1":"646.95","2":"20.38889","3":"18"},{"1":"651.35","2":"16.29310","3":"18"},{"1":"636.90","2":"18.27778","3":"18"},{"1":"670.70","2":"19.37472","3":"18"},{"1":"630.55","2":"18.90909","3":"18"},{"1":"643.20","2":"16.40693","3":"18"},{"1":"660.30","2":"15.59140","3":"18"},{"1":"648.95","2":"18.70694","3":"18"},{"1":"667.45","2":"18.32985","3":"18"},{"1":"638.70","2":"17.90235","3":"18"},{"1":"686.70","2":"18.91157","3":"18"},{"1":"671.30","2":"20.32497","3":"18"},{"1":"677.95","2":"20.02457","3":"18"},{"1":"648.70","2":"24.00000","3":"18"},{"1":"646.20","2":"17.60784","3":"18"},{"1":"627.25","2":"19.34853","3":"18"},{"1":"653.80","2":"19.67846","3":"18"},{"1":"639.80","2":"18.72861","3":"18"},{"1":"657.55","2":"15.88235","3":"18"},{"1":"644.20","2":"20.05491","3":"18"},{"1":"668.80","2":"17.98825","3":"18"},{"1":"645.10","2":"16.96629","3":"18"},{"1":"635.60","2":"19.23937","3":"18"},{"1":"674.25","2":"19.19586","3":"18"},{"1":"669.10","2":"19.59906","3":"18"},{"1":"661.95","2":"20.54348","3":"18"},{"1":"638.20","2":"18.58848","3":"18"},{"1":"666.70","2":"15.60419","3":"18"},{"1":"639.35","2":"15.29304","3":"18"},{"1":"645.60","2":"17.65537","3":"18"},{"1":"655.05","2":"17.57976","3":"18"},{"1":"616.30","2":"22.33333","3":"18"},{"1":"666.15","2":"18.75000","3":"18"},{"1":"646.55","2":"18.10241","3":"18"},{"1":"666.55","2":"20.25641","3":"18"},{"1":"661.85","2":"18.80207","3":"18"},{"1":"650.90","2":"18.77230","3":"18"},{"1":"605.55","2":"20.40521","3":"18"},{"1":"669.35","2":"18.65079","3":"18"},{"1":"695.20","2":"20.70707","3":"18"},{"1":"618.30","2":"22.00000","3":"18"},{"1":"655.80","2":"17.69978","3":"18"},{"1":"625.30","2":"21.48329","3":"18"},{"1":"655.05","2":"16.70103","3":"18"},{"1":"681.60","2":"19.57567","3":"18"},{"1":"654.10","2":"17.25806","3":"18"},{"1":"632.00","2":"17.37526","3":"18"},{"1":"675.40","2":"17.34931","3":"18"},{"1":"643.50","2":"16.26229","3":"18"},{"1":"656.80","2":"17.70045","3":"18"},{"1":"629.80","2":"20.12881","3":"18"},{"1":"654.85","2":"18.26539","3":"18"},{"1":"634.95","2":"14.54214","3":"18"},{"1":"677.25","2":"19.15261","3":"18"},{"1":"645.55","2":"17.36574","3":"18"},{"1":"671.65","2":"15.13898","3":"18"},{"1":"631.95","2":"17.84266","3":"18"},{"1":"633.15","2":"15.40704","3":"18"},{"1":"668.95","2":"18.86534","3":"18"},{"1":"662.75","2":"16.47413","3":"18"},{"1":"637.35","2":"17.86263","3":"18"},{"1":"666.65","2":"21.88586","3":"18"},{"1":"650.60","2":"20.20000","3":"18"},{"1":"648.00","2":"19.03640","3":"18"},{"1":"655.85","2":"17.88991","3":"19"},{"1":"621.40","2":"21.52466","3":"19"},{"1":"634.90","2":"18.69723","3":"19"},{"1":"623.20","2":"17.35714","3":"19"},{"1":"662.90","2":"18.67133","3":"19"},{"1":"656.55","2":"21.40625","3":"19"},{"1":"634.55","2":"19.50000","3":"19"},{"1":"676.60","2":"20.89412","3":"19"},{"1":"666.65","2":"19.94737","3":"19"},{"1":"639.35","2":"20.80556","3":"19"},{"1":"667.20","2":"21.23809","3":"19"},{"1":"676.55","2":"21.00000","3":"19"},{"1":"664.70","2":"20.60000","3":"19"},{"1":"643.50","2":"20.00822","3":"19"},{"1":"644.50","2":"18.02778","3":"19"},{"1":"634.10","2":"20.25196","3":"19"},{"1":"634.10","2":"16.97787","3":"19"},{"1":"669.85","2":"16.50980","3":"19"},{"1":"626.80","2":"22.70402","3":"19"},{"1":"617.35","2":"19.91111","3":"19"},{"1":"660.20","2":"18.33333","3":"19"},{"1":"698.20","2":"22.61905","3":"19"},{"1":"646.20","2":"19.44828","3":"19"},{"1":"628.75","2":"25.05263","3":"19"},{"1":"623.45","2":"20.67544","3":"19"},{"1":"669.30","2":"18.68235","3":"19"},{"1":"673.05","2":"22.84553","3":"19"},{"1":"671.90","2":"19.26667","3":"19"},{"1":"634.40","2":"19.25000","3":"19"},{"1":"659.90","2":"20.54545","3":"19"},{"1":"679.40","2":"20.60697","3":"19"},{"1":"638.55","2":"21.07268","3":"19"},{"1":"667.45","2":"21.53581","3":"19"},{"1":"656.65","2":"19.90400","3":"19"},{"1":"664.95","2":"21.19407","3":"19"},{"1":"666.10","2":"21.86535","3":"19"},{"1":"649.85","2":"18.32965","3":"19"},{"1":"646.90","2":"16.22857","3":"19"},{"1":"638.35","2":"19.17857","3":"19"},{"1":"647.70","2":"20.27737","3":"19"},{"1":"662.40","2":"22.98614","3":"19"},{"1":"655.40","2":"20.44444","3":"19"},{"1":"663.50","2":"19.82085","3":"19"},{"1":"666.60","2":"23.20522","3":"19"},{"1":"643.40","2":"19.26697","3":"19"},{"1":"655.35","2":"23.30189","3":"19"},{"1":"645.55","2":"21.18829","3":"19"},{"1":"643.05","2":"20.87180","3":"19"},{"1":"695.30","2":"19.01749","3":"19"},{"1":"656.40","2":"21.91938","3":"19"},{"1":"657.90","2":"20.10124","3":"19"},{"1":"641.45","2":"21.47651","3":"19"},{"1":"620.50","2":"20.06579","3":"19"},{"1":"645.75","2":"20.37510","3":"19"},{"1":"650.90","2":"22.44648","3":"19"},{"1":"678.40","2":"22.89524","3":"19"},{"1":"627.30","2":"20.49797","3":"19"},{"1":"605.55","2":"20.00000","3":"19"},{"1":"664.40","2":"22.25658","3":"19"},{"1":"632.00","2":"21.56436","3":"19"},{"1":"653.55","2":"19.47737","3":"19"},{"1":"663.35","2":"17.67002","3":"19"},{"1":"631.90","2":"21.94756","3":"19"},{"1":"672.35","2":"21.78339","3":"19"},{"1":"640.10","2":"19.14000","3":"19"},{"1":"652.10","2":"18.11050","3":"19"},{"1":"644.40","2":"20.68242","3":"19"},{"1":"668.60","2":"22.62361","3":"19"},{"1":"630.40","2":"21.78650","3":"19"},{"1":"647.30","2":"18.58293","3":"19"},{"1":"638.20","2":"21.54545","3":"19"},{"1":"662.70","2":"21.15289","3":"19"},{"1":"640.50","2":"16.63333","3":"19"},{"1":"639.50","2":"21.14438","3":"19"},{"1":"629.80","2":"19.78182","3":"19"},{"1":"643.40","2":"18.98373","3":"19"},{"1":"671.25","2":"17.66767","3":"19"},{"1":"654.20","2":"17.75499","3":"19"},{"1":"616.30","2":"15.27273","3":"19"},{"1":"636.90","2":"14.00000","3":"19"},{"1":"635.45","2":"20.59613","3":"19"},{"1":"639.80","2":"16.31169","3":"19"},{"1":"654.30","2":"21.12796","3":"19"},{"1":"633.05","2":"17.48801","3":"19"},{"1":"666.15","2":"17.88679","3":"19"},{"1":"649.30","2":"19.30676","3":"19"},{"1":"700.30","2":"20.89231","3":"19"},{"1":"672.05","2":"21.28684","3":"19"},{"1":"672.45","2":"20.19560","3":"19"},{"1":"647.60","2":"24.95000","3":"19"},{"1":"665.20","2":"18.13043","3":"19"},{"1":"652.10","2":"20.00000","3":"19"},{"1":"661.95","2":"18.72951","3":"19"},{"1":"643.60","2":"18.25000","3":"19"},{"1":"626.10","2":"18.99257","3":"19"},{"1":"609.00","2":"19.88764","3":"19"},{"1":"612.65","2":"19.37895","3":"19"},{"1":"648.95","2":"20.46259","3":"19"},{"1":"691.05","2":"22.29157","3":"19"},{"1":"620.30","2":"20.70474","3":"19"},{"1":"647.60","2":"19.06005","3":"19"},{"1":"667.60","2":"20.23247","3":"19"},{"1":"672.55","2":"19.69012","3":"19"},{"1":"628.40","2":"20.36254","3":"19"},{"1":"676.15","2":"19.75422","3":"19"},{"1":"693.95","2":"19.37977","3":"19"},{"1":"649.70","2":"22.92351","3":"19"},{"1":"635.75","2":"19.37340","3":"19"},{"1":"650.65","2":"19.15516","3":"19"},{"1":"666.85","2":"21.30000","3":"19"},{"1":"672.70","2":"18.30357","3":"19"},{"1":"639.30","2":"21.07926","3":"19"},{"1":"657.40","2":"18.79121","3":"19"},{"1":"635.60","2":"19.62662","3":"19"},{"1":"669.30","2":"19.59016","3":"19"},{"1":"671.95","2":"20.87187","3":"19"},{"1":"672.20","2":"21.11500","3":"19"},{"1":"666.05","2":"20.08452","3":"19"},{"1":"660.30","2":"19.91049","3":"19"},{"1":"664.00","2":"17.81285","3":"19"},{"1":"678.05","2":"18.13333","3":"19"},{"1":"680.45","2":"19.22221","3":"19"},{"1":"661.85","2":"18.66072","3":"19"},{"1":"645.10","2":"19.60000","3":"19"},{"1":"622.60","2":"19.28384","3":"19"},{"1":"631.05","2":"22.81818","3":"19"},{"1":"635.05","2":"18.80922","3":"19"},{"1":"663.45","2":"21.37363","3":"19"},{"1":"640.90","2":"20.02041","3":"19"},{"1":"634.95","2":"21.49862","3":"19"},{"1":"628.65","2":"15.42857","3":"19"},{"1":"650.90","2":"22.40000","3":"19"},{"1":"662.65","2":"20.12709","3":"19"},{"1":"658.60","2":"19.03798","3":"19"},{"1":"657.15","2":"17.34216","3":"19"},{"1":"668.95","2":"17.01863","3":"19"},{"1":"669.95","2":"20.80000","3":"19"},{"1":"637.35","2":"21.15385","3":"19"},{"1":"669.35","2":"18.45833","3":"19"},{"1":"652.40","2":"19.14082","3":"19"},{"1":"638.30","2":"19.40766","3":"19"},{"1":"706.75","2":"19.56896","3":"19"},{"1":"654.10","2":"21.50120","3":"19"},{"1":"690.80","2":"17.52941","3":"19"},{"1":"630.35","2":"16.43017","3":"19"},{"1":"650.45","2":"19.79654","3":"19"},{"1":"606.75","2":"17.18613","3":"19"},{"1":"642.20","2":"17.61589","3":"19"},{"1":"618.05","2":"20.12537","3":"19"},{"1":"638.30","2":"22.16667","3":"19"},{"1":"652.30","2":"19.96154","3":"19"},{"1":"641.55","2":"19.03945","3":"19"},{"1":"653.55","2":"15.22436","3":"19"},{"1":"643.70","2":"21.14475","3":"19"},{"1":"645.75","2":"19.64390","3":"19"},{"1":"695.20","2":"21.04869","3":"19"},{"1":"691.35","2":"20.17544","3":"19"},{"1":"668.10","2":"21.39130","3":"19"},{"1":"664.00","2":"20.00833","3":"19"},{"1":"673.30","2":"20.29137","3":"19"},{"1":"612.50","2":"17.66667","3":"19"},{"1":"660.05","2":"18.22055","3":"19"},{"1":"643.25","2":"20.27100","3":"19"},{"1":"650.55","2":"20.19895","3":"19"},{"1":"662.55","2":"21.38424","3":"19"},{"1":"644.20","2":"20.97368","3":"19"},{"1":"647.25","2":"20.00000","3":"19"},{"1":"646.35","2":"17.15328","3":"19"},{"1":"666.70","2":"22.34977","3":"19"},{"1":"686.70","2":"22.17007","3":"19"},{"1":"671.65","2":"18.18182","3":"19"},{"1":"659.40","2":"18.95714","3":"19"},{"1":"672.05","2":"19.74533","3":"19"},{"1":"666.00","2":"16.42623","3":"19"},{"1":"662.40","2":"16.62540","3":"19"},{"1":"644.45","2":"16.38177","3":"19"},{"1":"651.20","2":"20.07416","3":"19"},{"1":"633.90","2":"17.99544","3":"19"},{"1":"659.40","2":"19.39130","3":"19"},{"1":"661.35","2":"16.42857","3":"19"},{"1":"653.10","2":"16.72949","3":"19"},{"1":"676.95","2":"24.41345","3":"19"},{"1":"657.55","2":"18.26415","3":"19"},{"1":"668.90","2":"18.95504","3":"19"},{"1":"657.00","2":"21.03896","3":"19"},{"1":"663.90","2":"20.74074","3":"19"},{"1":"623.60","2":"18.10000","3":"19"},{"1":"648.20","2":"19.84615","3":"19"},{"1":"668.40","2":"21.60000","3":"19"},{"1":"646.55","2":"22.44242","3":"19"},{"1":"657.00","2":"23.01438","3":"19"},{"1":"661.90","2":"17.74892","3":"19"},{"1":"651.40","2":"18.28664","3":"19"},{"1":"648.25","2":"19.26544","3":"19"},{"1":"661.90","2":"22.66667","3":"19"},{"1":"664.95","2":"19.29412","3":"19"},{"1":"669.80","2":"17.36364","3":"19"},{"1":"664.15","2":"19.82143","3":"19"},{"1":"644.45","2":"20.43378","3":"19"},{"1":"632.95","2":"21.03721","3":"19"},{"1":"631.95","2":"19.92462","3":"19"},{"1":"649.50","2":"19.00986","3":"19"},{"1":"633.65","2":"23.82222","3":"19"},{"1":"655.75","2":"19.36909","3":"19"},{"1":"657.75","2":"19.82857","3":"19"},{"1":"637.10","2":"15.25885","3":"19"},{"1":"642.75","2":"17.16129","3":"19"},{"1":"618.30","2":"21.81333","3":"19"},{"1":"681.60","2":"19.07471","3":"19"},{"1":"632.25","2":"25.78512","3":"19"},{"1":"616.45","2":"18.21261","3":"19"},{"1":"673.25","2":"18.16606","3":"19"},{"1":"653.70","2":"16.97297","3":"19"},{"1":"646.70","2":"21.50087","3":"19"},{"1":"665.10","2":"20.60000","3":"19"},{"1":"624.95","2":"16.99029","3":"19"},{"1":"670.00","2":"20.77954","3":"19"},{"1":"627.10","2":"15.51247","3":"19"},{"1":"654.85","2":"19.88506","3":"19"},{"1":"687.55","2":"21.39882","3":"19"},{"1":"668.00","2":"20.49751","3":"19"},{"1":"616.30","2":"19.36376","3":"19"},{"1":"664.45","2":"17.65957","3":"19"},{"1":"656.50","2":"21.01796","3":"19"},{"1":"655.05","2":"19.05565","3":"19"},{"1":"624.15","2":"22.53846","3":"19"},{"1":"668.65","2":"21.10787","3":"19"},{"1":"640.85","2":"20.05135","3":"19"},{"1":"653.50","2":"14.20176","3":"19"},{"1":"684.35","2":"18.47687","3":"19"},{"1":"631.40","2":"18.63542","3":"19"},{"1":"646.95","2":"20.94595","3":"19"},{"1":"661.20","2":"21.08548","3":"19"},{"1":"641.80","2":"18.69288","3":"19"},{"1":"678.80","2":"20.86808","3":"19"},{"1":"665.95","2":"19.82558","3":"19"},{"1":"634.70","2":"19.75000","3":"19"},{"1":"681.90","2":"19.50000","3":"19"},{"1":"623.10","2":"18.39080","3":"19"},{"1":"679.75","2":"18.78676","3":"19"},{"1":"633.15","2":"19.77018","3":"19"},{"1":"696.55","2":"19.33333","3":"19"},{"1":"660.75","2":"21.46392","3":"19"},{"1":"651.90","2":"23.08492","3":"19"},{"1":"694.80","2":"21.06299","3":"19"},{"1":"682.55","2":"18.68687","3":"19"},{"1":"638.00","2":"20.77024","3":"19"},{"1":"648.00","2":"19.30556","3":"19"},{"1":"662.55","2":"20.13280","3":"19"},{"1":"651.85","2":"20.66964","3":"19"},{"1":"636.60","2":"22.28155","3":"19"},{"1":"662.50","2":"20.60027","3":"19"},{"1":"634.05","2":"20.82734","3":"19"},{"1":"656.80","2":"19.22492","3":"19"},{"1":"666.55","2":"17.65477","3":"19"},{"1":"657.80","2":"17.00000","3":"19"},{"1":"670.70","2":"16.49773","3":"19"},{"1":"661.60","2":"19.78261","3":"19"},{"1":"641.10","2":"22.30216","3":"19"},{"1":"679.80","2":"17.73077","3":"19"},{"1":"637.00","2":"20.44836","3":"19"},{"1":"647.05","2":"20.37169","3":"19"},{"1":"659.15","2":"20.16479","3":"19"},{"1":"624.55","2":"21.61538","3":"19"},{"1":"640.15","2":"20.56143","3":"19"},{"1":"626.90","2":"19.95551","3":"19"},{"1":"655.55","2":"21.18387","3":"19"},{"1":"683.40","2":"18.81042","3":"19"},{"1":"666.65","2":"20.57838","3":"19"},{"1":"698.25","2":"18.32461","3":"19"},{"1":"665.35","2":"18.82063","3":"19"},{"1":"684.30","2":"20.81633","3":"19"},{"1":"682.15","2":"20.00000","3":"19"},{"1":"639.25","2":"19.68182","3":"19"},{"1":"673.55","2":"19.39018","3":"19"},{"1":"646.50","2":"20.92732","3":"19"},{"1":"651.80","2":"19.94437","3":"19"},{"1":"669.35","2":"20.79109","3":"19"},{"1":"691.90","2":"19.20354","3":"19"},{"1":"632.45","2":"19.02439","3":"19"},{"1":"664.30","2":"17.62058","3":"19"},{"1":"637.95","2":"20.23715","3":"19"},{"1":"658.80","2":"19.29374","3":"19"},{"1":"652.00","2":"18.82998","3":"19"},{"1":"643.50","2":"20.33949","3":"19"},{"1":"679.65","2":"19.22900","3":"19"},{"1":"663.85","2":"17.89130","3":"19"},{"1":"684.95","2":"19.51881","3":"19"},{"1":"677.25","2":"19.08451","3":"19"},{"1":"682.65","2":"19.93548","3":"19"},{"1":"682.45","2":"18.87326","3":"19"},{"1":"638.70","2":"20.14178","3":"19"},{"1":"681.30","2":"23.55637","3":"19"},{"1":"659.35","2":"21.46479","3":"19"},{"1":"654.20","2":"19.19101","3":"19"},{"1":"673.90","2":"20.13080","3":"19"},{"1":"616.30","2":"25.80000","3":"19"},{"1":"667.45","2":"18.77774","3":"19"},{"1":"640.75","2":"19.10982","3":"19"},{"1":"676.85","2":"19.70109","3":"19"},{"1":"648.70","2":"18.61594","3":"19"},{"1":"645.55","2":"20.99721","3":"19"},{"1":"636.50","2":"20.00000","3":"19"},{"1":"689.10","2":"20.98325","3":"19"},{"1":"630.55","2":"21.64262","3":"19"},{"1":"654.90","2":"20.02967","3":"19"},{"1":"619.80","2":"19.81140","3":"19"},{"1":"658.00","2":"18.00000","3":"19"},{"1":"675.40","2":"19.35811","3":"19"},{"1":"637.95","2":"20.17912","3":"19"},{"1":"671.90","2":"21.11986","3":"19"},{"1":"632.85","2":"23.38974","3":"19"},{"1":"655.05","2":"22.18182","3":"19"},{"1":"656.80","2":"19.94283","3":"19"},{"1":"641.45","2":"17.78826","3":"19"},{"1":"694.25","2":"14.70588","3":"19"},{"1":"679.50","2":"19.04077","3":"19"},{"1":"665.90","2":"20.89195","3":"19"},{"1":"659.80","2":"19.83851","3":"19"},{"1":"664.75","2":"19.52191","3":"19"},{"1":"632.20","2":"20.68622","3":"19"},{"1":"653.40","2":"18.18182","3":"19"},{"1":"646.00","2":"18.89224","3":"19"},{"1":"634.15","2":"24.88889","3":"19"},{"1":"667.15","2":"18.58064","3":"19"},{"1":"660.10","2":"18.04000","3":"19"},{"1":"645.25","2":"17.73399","3":"19"},{"1":"671.70","2":"21.45455","3":"19"},{"1":"660.95","2":"19.92343","3":"19"},{"1":"645.00","2":"20.33942","3":"19"},{"1":"677.95","2":"22.54608","3":"19"},{"1":"675.70","2":"21.10344","3":"19"},{"1":"671.30","2":"18.19743","3":"19"},{"1":"663.85","2":"20.10768","3":"19"},{"1":"655.20","2":"19.15984","3":"19"},{"1":"639.90","2":"19.54545","3":"19"},{"1":"636.95","2":"20.88889","3":"19"},{"1":"672.30","2":"18.39150","3":"19"},{"1":"625.30","2":"19.17990","3":"19"},{"1":"674.25","2":"19.39771","3":"19"},{"1":"630.55","2":"21.67827","3":"19"},{"1":"671.60","2":"19.28889","3":"19"},{"1":"635.95","2":"20.34927","3":"19"},{"1":"664.15","2":"20.96416","3":"19"},{"1":"644.55","2":"19.46039","3":"19"},{"1":"643.70","2":"19.28572","3":"19"},{"1":"662.45","2":"20.91979","3":"19"},{"1":"704.30","2":"20.90021","3":"19"},{"1":"628.25","2":"20.59575","3":"19"},{"1":"666.85","2":"19.37500","3":"19"},{"1":"635.20","2":"19.95122","3":"19"},{"1":"656.70","2":"18.84973","3":"19"},{"1":"646.40","2":"18.11787","3":"19"},{"1":"636.10","2":"19.18341","3":"19"},{"1":"650.60","2":"22.00000","3":"19"},{"1":"645.60","2":"21.58416","3":"19"},{"1":"639.75","2":"20.38889","3":"19"},{"1":"681.30","2":"16.29310","3":"19"},{"1":"666.15","2":"18.27778","3":"19"},{"1":"615.75","2":"19.37472","3":"19"},{"1":"644.20","2":"18.90909","3":"19"},{"1":"652.50","2":"16.40693","3":"19"},{"1":"628.55","2":"15.59140","3":"19"},{"1":"659.05","2":"18.70694","3":"19"},{"1":"686.05","2":"18.32985","3":"19"},{"1":"661.60","2":"17.90235","3":"19"},{"1":"665.65","2":"18.91157","3":"19"},{"1":"634.00","2":"20.32497","3":"19"},{"1":"634.20","2":"20.02457","3":"19"},{"1":"627.25","2":"24.00000","3":"19"},{"1":"654.60","2":"17.60784","3":"19"},{"1":"653.85","2":"19.34853","3":"19"},{"1":"654.85","2":"19.67846","3":"19"},{"1":"652.30","2":"18.72861","3":"19"},{"1":"657.65","2":"15.88235","3":"19"},{"1":"657.50","2":"20.05491","3":"19"},{"1":"661.45","2":"17.98825","3":"19"},{"1":"643.20","2":"16.96629","3":"19"},{"1":"662.75","2":"19.23937","3":"19"},{"1":"651.35","2":"19.19586","3":"19"},{"1":"653.80","2":"19.59906","3":"19"},{"1":"652.35","2":"20.54348","3":"19"},{"1":"671.75","2":"18.58848","3":"19"},{"1":"639.85","2":"15.60419","3":"19"},{"1":"621.75","2":"15.29304","3":"19"},{"1":"625.85","2":"17.65537","3":"19"},{"1":"698.45","2":"17.57976","3":"19"},{"1":"655.35","2":"22.33333","3":"19"},{"1":"680.05","2":"18.75000","3":"19"},{"1":"636.70","2":"18.10241","3":"19"},{"1":"652.40","2":"20.25641","3":"19"},{"1":"642.20","2":"18.80207","3":"19"},{"1":"655.05","2":"18.77230","3":"19"},{"1":"699.10","2":"20.40521","3":"19"},{"1":"652.85","2":"18.65079","3":"19"},{"1":"637.65","2":"20.70707","3":"19"},{"1":"655.30","2":"22.00000","3":"19"},{"1":"673.55","2":"17.69978","3":"19"},{"1":"668.80","2":"21.48329","3":"19"},{"1":"666.45","2":"16.70103","3":"19"},{"1":"655.70","2":"19.57567","3":"19"},{"1":"684.80","2":"17.25806","3":"19"},{"1":"635.60","2":"17.37526","3":"19"},{"1":"648.35","2":"17.34931","3":"19"},{"1":"671.60","2":"16.26229","3":"19"},{"1":"653.95","2":"17.70045","3":"19"},{"1":"658.35","2":"20.12881","3":"19"},{"1":"651.45","2":"18.26539","3":"19"},{"1":"669.10","2":"14.54214","3":"19"},{"1":"642.40","2":"19.15261","3":"19"},{"1":"661.85","2":"17.36574","3":"19"},{"1":"644.95","2":"15.13898","3":"19"},{"1":"644.70","2":"17.84266","3":"19"},{"1":"661.85","2":"15.40704","3":"19"},{"1":"655.80","2":"18.86534","3":"19"},{"1":"651.15","2":"16.47413","3":"19"},{"1":"631.85","2":"17.86263","3":"19"},{"1":"622.05","2":"21.88586","3":"19"},{"1":"683.35","2":"20.20000","3":"19"},{"1":"649.15","2":"19.03640","3":"19"},{"1":"686.05","2":"17.88991","3":"20"},{"1":"663.85","2":"21.52466","3":"20"},{"1":"645.75","2":"18.69723","3":"20"},{"1":"646.70","2":"17.35714","3":"20"},{"1":"657.80","2":"18.67133","3":"20"},{"1":"639.25","2":"21.40625","3":"20"},{"1":"639.30","2":"19.50000","3":"20"},{"1":"671.60","2":"20.89412","3":"20"},{"1":"657.65","2":"19.94737","3":"20"},{"1":"662.45","2":"20.80556","3":"20"},{"1":"680.45","2":"21.23809","3":"20"},{"1":"691.90","2":"21.00000","3":"20"},{"1":"649.70","2":"20.60000","3":"20"},{"1":"650.60","2":"20.00822","3":"20"},{"1":"668.60","2":"18.02778","3":"20"},{"1":"648.35","2":"20.25196","3":"20"},{"1":"684.95","2":"16.97787","3":"20"},{"1":"635.95","2":"16.50980","3":"20"},{"1":"668.00","2":"22.70402","3":"20"},{"1":"653.50","2":"19.91111","3":"20"},{"1":"649.50","2":"18.33333","3":"20"},{"1":"665.10","2":"22.61905","3":"20"},{"1":"639.35","2":"19.44828","3":"20"},{"1":"653.80","2":"25.05263","3":"20"},{"1":"652.40","2":"20.67544","3":"20"},{"1":"654.90","2":"18.68235","3":"20"},{"1":"631.90","2":"22.84553","3":"20"},{"1":"635.75","2":"19.26667","3":"20"},{"1":"656.80","2":"19.25000","3":"20"},{"1":"651.80","2":"20.54545","3":"20"},{"1":"666.85","2":"20.60697","3":"20"},{"1":"647.70","2":"21.07268","3":"20"},{"1":"645.75","2":"21.53581","3":"20"},{"1":"668.10","2":"19.90400","3":"20"},{"1":"624.55","2":"21.19407","3":"20"},{"1":"628.25","2":"21.86535","3":"20"},{"1":"671.25","2":"18.32965","3":"20"},{"1":"632.95","2":"16.22857","3":"20"},{"1":"662.65","2":"19.17857","3":"20"},{"1":"642.75","2":"20.27737","3":"20"},{"1":"662.55","2":"22.98614","3":"20"},{"1":"634.10","2":"20.44444","3":"20"},{"1":"643.25","2":"19.82085","3":"20"},{"1":"672.55","2":"23.20522","3":"20"},{"1":"645.00","2":"19.26697","3":"20"},{"1":"676.15","2":"23.30189","3":"20"},{"1":"662.90","2":"21.18829","3":"20"},{"1":"669.35","2":"20.87180","3":"20"},{"1":"657.40","2":"19.01749","3":"20"},{"1":"651.85","2":"21.91938","3":"20"},{"1":"647.25","2":"20.10124","3":"20"},{"1":"681.30","2":"21.47651","3":"20"},{"1":"653.95","2":"20.06579","3":"20"},{"1":"679.50","2":"20.37510","3":"20"},{"1":"653.85","2":"22.44648","3":"20"},{"1":"652.85","2":"22.89524","3":"20"},{"1":"634.05","2":"20.49797","3":"20"},{"1":"615.75","2":"20.00000","3":"20"},{"1":"643.70","2":"22.25658","3":"20"},{"1":"654.30","2":"21.56436","3":"20"},{"1":"661.85","2":"19.47737","3":"20"},{"1":"687.55","2":"17.67002","3":"20"},{"1":"646.90","2":"21.94756","3":"20"},{"1":"636.10","2":"21.78339","3":"20"},{"1":"652.40","2":"19.14000","3":"20"},{"1":"671.70","2":"18.11050","3":"20"},{"1":"684.80","2":"20.68242","3":"20"},{"1":"623.10","2":"22.62361","3":"20"},{"1":"643.05","2":"21.78650","3":"20"},{"1":"676.95","2":"18.58293","3":"20"},{"1":"660.30","2":"21.54545","3":"20"},{"1":"698.20","2":"21.15289","3":"20"},{"1":"638.55","2":"16.63333","3":"20"},{"1":"636.50","2":"21.14438","3":"20"},{"1":"660.75","2":"19.78182","3":"20"},{"1":"656.70","2":"18.98373","3":"20"},{"1":"649.15","2":"17.66767","3":"20"},{"1":"651.90","2":"17.75499","3":"20"},{"1":"659.40","2":"15.27273","3":"20"},{"1":"655.80","2":"14.00000","3":"20"},{"1":"655.55","2":"20.59613","3":"20"},{"1":"622.05","2":"16.31169","3":"20"},{"1":"673.30","2":"21.12796","3":"20"},{"1":"704.30","2":"17.48801","3":"20"},{"1":"639.80","2":"17.88679","3":"20"},{"1":"684.30","2":"19.30676","3":"20"},{"1":"676.60","2":"20.89231","3":"20"},{"1":"666.05","2":"21.28684","3":"20"},{"1":"652.00","2":"20.19560","3":"20"},{"1":"661.90","2":"24.95000","3":"20"},{"1":"627.25","2":"18.13043","3":"20"},{"1":"659.90","2":"20.00000","3":"20"},{"1":"634.10","2":"18.72951","3":"20"},{"1":"623.45","2":"18.25000","3":"20"},{"1":"647.60","2":"18.99257","3":"20"},{"1":"636.60","2":"19.88764","3":"20"},{"1":"647.60","2":"19.37895","3":"20"},{"1":"654.10","2":"20.46259","3":"20"},{"1":"661.85","2":"22.29157","3":"20"},{"1":"653.40","2":"20.70474","3":"20"},{"1":"664.30","2":"19.06005","3":"20"},{"1":"632.85","2":"20.23247","3":"20"},{"1":"669.30","2":"19.69012","3":"20"},{"1":"682.45","2":"20.36254","3":"20"},{"1":"662.55","2":"19.75422","3":"20"},{"1":"653.70","2":"19.37977","3":"20"},{"1":"630.35","2":"22.92351","3":"20"},{"1":"652.10","2":"19.37340","3":"20"},{"1":"631.95","2":"19.15516","3":"20"},{"1":"651.40","2":"21.30000","3":"20"},{"1":"673.55","2":"18.30357","3":"20"},{"1":"670.70","2":"21.07926","3":"20"},{"1":"668.80","2":"18.79121","3":"20"},{"1":"700.30","2":"19.62662","3":"20"},{"1":"666.45","2":"19.59016","3":"20"},{"1":"643.40","2":"20.87187","3":"20"},{"1":"637.00","2":"21.11500","3":"20"},{"1":"657.00","2":"20.08452","3":"20"},{"1":"668.65","2":"19.91049","3":"20"},{"1":"691.05","2":"17.81285","3":"20"},{"1":"652.10","2":"18.13333","3":"20"},{"1":"691.35","2":"19.22221","3":"20"},{"1":"648.00","2":"18.66072","3":"20"},{"1":"644.50","2":"19.60000","3":"20"},{"1":"654.20","2":"19.28384","3":"20"},{"1":"660.05","2":"22.81818","3":"20"},{"1":"616.30","2":"18.80922","3":"20"},{"1":"648.70","2":"21.37363","3":"20"},{"1":"666.70","2":"20.02041","3":"20"},{"1":"664.15","2":"21.49862","3":"20"},{"1":"653.10","2":"15.42857","3":"20"},{"1":"660.20","2":"22.40000","3":"20"},{"1":"651.20","2":"20.12709","3":"20"},{"1":"647.30","2":"19.03798","3":"20"},{"1":"662.40","2":"17.34216","3":"20"},{"1":"628.65","2":"17.01863","3":"20"},{"1":"686.70","2":"20.80000","3":"20"},{"1":"706.75","2":"21.15385","3":"20"},{"1":"642.40","2":"18.45833","3":"20"},{"1":"672.45","2":"19.14082","3":"20"},{"1":"627.30","2":"19.40766","3":"20"},{"1":"623.20","2":"19.56896","3":"20"},{"1":"666.65","2":"21.50120","3":"20"},{"1":"606.75","2":"17.52941","3":"20"},{"1":"658.35","2":"16.43017","3":"20"},{"1":"695.20","2":"19.79654","3":"20"},{"1":"634.20","2":"17.18613","3":"20"},{"1":"636.95","2":"17.61589","3":"20"},{"1":"634.70","2":"20.12537","3":"20"},{"1":"651.15","2":"22.16667","3":"20"},{"1":"659.80","2":"19.96154","3":"20"},{"1":"684.35","2":"19.03945","3":"20"},{"1":"646.55","2":"15.22436","3":"20"},{"1":"645.25","2":"21.14475","3":"20"},{"1":"645.60","2":"19.64390","3":"20"},{"1":"656.80","2":"21.04869","3":"20"},{"1":"636.70","2":"20.17544","3":"20"},{"1":"640.85","2":"21.39130","3":"20"},{"1":"616.45","2":"20.00833","3":"20"},{"1":"659.40","2":"20.29137","3":"20"},{"1":"646.00","2":"17.66667","3":"20"},{"1":"676.85","2":"18.22055","3":"20"},{"1":"694.25","2":"20.27100","3":"20"},{"1":"661.60","2":"20.19895","3":"20"},{"1":"672.30","2":"21.38424","3":"20"},{"1":"667.45","2":"20.97368","3":"20"},{"1":"646.35","2":"20.00000","3":"20"},{"1":"640.50","2":"17.15328","3":"20"},{"1":"633.65","2":"22.34977","3":"20"},{"1":"618.30","2":"22.17007","3":"20"},{"1":"641.55","2":"18.18182","3":"20"},{"1":"665.20","2":"18.95714","3":"20"},{"1":"654.20","2":"19.74533","3":"20"},{"1":"656.55","2":"16.42623","3":"20"},{"1":"609.00","2":"16.62540","3":"20"},{"1":"637.35","2":"16.38177","3":"20"},{"1":"657.50","2":"20.07416","3":"20"},{"1":"643.50","2":"17.99544","3":"20"},{"1":"634.15","2":"19.39130","3":"20"},{"1":"640.15","2":"16.42857","3":"20"},{"1":"666.85","2":"16.72949","3":"20"},{"1":"633.90","2":"24.41345","3":"20"},{"1":"661.60","2":"18.26415","3":"20"},{"1":"667.15","2":"18.95504","3":"20"},{"1":"673.90","2":"21.03896","3":"20"},{"1":"658.80","2":"20.74074","3":"20"},{"1":"662.40","2":"18.10000","3":"20"},{"1":"699.10","2":"19.84615","3":"20"},{"1":"643.40","2":"21.60000","3":"20"},{"1":"662.50","2":"22.44242","3":"20"},{"1":"654.60","2":"23.01438","3":"20"},{"1":"620.50","2":"17.74892","3":"20"},{"1":"653.55","2":"18.28664","3":"20"},{"1":"632.20","2":"19.26544","3":"20"},{"1":"643.20","2":"22.66667","3":"20"},{"1":"644.20","2":"19.29412","3":"20"},{"1":"637.10","2":"17.36364","3":"20"},{"1":"661.85","2":"19.82143","3":"20"},{"1":"653.55","2":"20.43378","3":"20"},{"1":"639.50","2":"21.03721","3":"20"},{"1":"666.60","2":"19.92462","3":"20"},{"1":"655.05","2":"19.00986","3":"20"},{"1":"626.80","2":"23.82222","3":"20"},{"1":"671.60","2":"19.36909","3":"20"},{"1":"678.80","2":"19.82857","3":"20"},{"1":"638.20","2":"15.25885","3":"20"},{"1":"638.70","2":"17.16129","3":"20"},{"1":"635.05","2":"21.81333","3":"20"},{"1":"628.55","2":"19.07471","3":"20"},{"1":"624.15","2":"25.78512","3":"20"},{"1":"657.00","2":"18.21261","3":"20"},{"1":"632.00","2":"18.16606","3":"20"},{"1":"605.55","2":"16.97297","3":"20"},{"1":"630.40","2":"21.50087","3":"20"},{"1":"681.90","2":"20.60000","3":"20"},{"1":"636.90","2":"16.99029","3":"20"},{"1":"628.40","2":"20.77954","3":"20"},{"1":"652.30","2":"15.51247","3":"20"},{"1":"635.20","2":"19.88506","3":"20"},{"1":"662.70","2":"21.39882","3":"20"},{"1":"675.40","2":"20.49751","3":"20"},{"1":"666.15","2":"19.36376","3":"20"},{"1":"666.10","2":"17.65957","3":"20"},{"1":"646.40","2":"21.01796","3":"20"},{"1":"616.30","2":"19.05565","3":"20"},{"1":"671.75","2":"22.53846","3":"20"},{"1":"667.45","2":"21.10787","3":"20"},{"1":"646.50","2":"20.05135","3":"20"},{"1":"641.45","2":"14.20176","3":"20"},{"1":"627.10","2":"18.47687","3":"20"},{"1":"641.10","2":"18.63542","3":"20"},{"1":"631.05","2":"20.94595","3":"20"},{"1":"633.05","2":"21.08548","3":"20"},{"1":"678.40","2":"18.69288","3":"20"},{"1":"661.35","2":"20.86808","3":"20"},{"1":"634.40","2":"19.82558","3":"20"},{"1":"659.35","2":"19.75000","3":"20"},{"1":"625.30","2":"19.50000","3":"20"},{"1":"655.05","2":"18.39080","3":"20"},{"1":"664.95","2":"18.78676","3":"20"},{"1":"660.10","2":"19.77018","3":"20"},{"1":"656.50","2":"19.33333","3":"20"},{"1":"638.35","2":"21.46392","3":"20"},{"1":"639.75","2":"23.08492","3":"20"},{"1":"639.90","2":"21.06299","3":"20"},{"1":"635.45","2":"18.68687","3":"20"},{"1":"667.20","2":"20.77024","3":"20"},{"1":"696.55","2":"19.30556","3":"20"},{"1":"641.45","2":"20.13280","3":"20"},{"1":"624.95","2":"20.66964","3":"20"},{"1":"664.95","2":"22.28155","3":"20"},{"1":"672.70","2":"20.60027","3":"20"},{"1":"659.05","2":"20.82734","3":"20"},{"1":"642.20","2":"19.22492","3":"20"},{"1":"667.60","2":"17.65477","3":"20"},{"1":"664.45","2":"17.00000","3":"20"},{"1":"658.00","2":"16.49773","3":"20"},{"1":"659.15","2":"19.78261","3":"20"},{"1":"655.85","2":"22.30216","3":"20"},{"1":"634.95","2":"17.73077","3":"20"},{"1":"654.85","2":"20.44836","3":"20"},{"1":"657.55","2":"20.37169","3":"20"},{"1":"681.30","2":"20.16479","3":"20"},{"1":"645.10","2":"21.61538","3":"20"},{"1":"650.65","2":"20.56143","3":"20"},{"1":"679.80","2":"19.95551","3":"20"},{"1":"634.90","2":"21.18387","3":"20"},{"1":"655.30","2":"18.81042","3":"20"},{"1":"672.05","2":"20.57838","3":"20"},{"1":"661.45","2":"18.32461","3":"20"},{"1":"665.35","2":"18.82063","3":"20"},{"1":"631.40","2":"20.81633","3":"20"},{"1":"631.85","2":"20.00000","3":"20"},{"1":"650.55","2":"19.68182","3":"20"},{"1":"679.65","2":"19.39018","3":"20"},{"1":"644.45","2":"20.92732","3":"20"},{"1":"671.90","2":"19.94437","3":"20"},{"1":"698.25","2":"20.79109","3":"20"},{"1":"683.40","2":"19.20354","3":"20"},{"1":"646.20","2":"19.02439","3":"20"},{"1":"656.65","2":"17.62058","3":"20"},{"1":"666.65","2":"20.23715","3":"20"},{"1":"666.15","2":"19.29374","3":"20"},{"1":"637.95","2":"18.82998","3":"20"},{"1":"621.40","2":"20.33949","3":"20"},{"1":"665.90","2":"19.22900","3":"20"},{"1":"673.05","2":"17.89130","3":"20"},{"1":"677.95","2":"19.51881","3":"20"},{"1":"671.30","2":"19.08451","3":"20"},{"1":"640.75","2":"19.93548","3":"20"},{"1":"689.10","2":"18.87326","3":"20"},{"1":"693.95","2":"20.14178","3":"20"},{"1":"671.95","2":"23.55637","3":"20"},{"1":"643.60","2":"21.46479","3":"20"},{"1":"638.00","2":"19.19101","3":"20"},{"1":"669.10","2":"20.13080","3":"20"},{"1":"622.60","2":"25.80000","3":"20"},{"1":"634.00","2":"18.77774","3":"20"},{"1":"694.80","2":"19.10982","3":"20"},{"1":"640.90","2":"19.70109","3":"20"},{"1":"672.35","2":"18.61594","3":"20"},{"1":"644.95","2":"20.99721","3":"20"},{"1":"669.95","2":"20.00000","3":"20"},{"1":"651.45","2":"20.98325","3":"20"},{"1":"675.70","2":"21.64262","3":"20"},{"1":"644.45","2":"20.02967","3":"20"},{"1":"635.60","2":"19.81140","3":"20"},{"1":"652.50","2":"18.00000","3":"20"},{"1":"668.95","2":"19.35811","3":"20"},{"1":"676.55","2":"20.17912","3":"20"},{"1":"639.85","2":"21.11986","3":"20"},{"1":"645.55","2":"23.38974","3":"20"},{"1":"655.75","2":"22.18182","3":"20"},{"1":"650.90","2":"19.94283","3":"20"},{"1":"664.70","2":"17.78826","3":"20"},{"1":"618.05","2":"14.70588","3":"20"},{"1":"663.85","2":"19.04077","3":"20"},{"1":"626.10","2":"20.89195","3":"20"},{"1":"668.40","2":"19.83851","3":"20"},{"1":"690.80","2":"19.52191","3":"20"},{"1":"637.65","2":"20.68622","3":"20"},{"1":"677.25","2":"18.18182","3":"20"},{"1":"664.40","2":"18.89224","3":"20"},{"1":"661.20","2":"24.88889","3":"20"},{"1":"679.40","2":"18.58064","3":"20"},{"1":"655.35","2":"18.04000","3":"20"},{"1":"683.35","2":"17.73399","3":"20"},{"1":"629.80","2":"21.45455","3":"20"},{"1":"650.45","2":"19.92343","3":"20"},{"1":"682.65","2":"20.33942","3":"20"},{"1":"630.55","2":"22.54608","3":"20"},{"1":"669.30","2":"21.10344","3":"20"},{"1":"654.85","2":"18.19743","3":"20"},{"1":"664.00","2":"20.10768","3":"20"},{"1":"669.80","2":"19.15984","3":"20"},{"1":"646.95","2":"19.54545","3":"20"},{"1":"621.75","2":"20.88889","3":"20"},{"1":"612.65","2":"18.39150","3":"20"},{"1":"635.60","2":"19.17990","3":"20"},{"1":"628.75","2":"19.39771","3":"20"},{"1":"678.05","2":"21.67827","3":"20"},{"1":"648.95","2":"19.28889","3":"20"},{"1":"655.40","2":"20.34927","3":"20"},{"1":"666.00","2":"20.96416","3":"20"},{"1":"643.70","2":"19.46039","3":"20"},{"1":"657.90","2":"19.28572","3":"20"},{"1":"623.60","2":"20.91979","3":"20"},{"1":"619.80","2":"20.90021","3":"20"},{"1":"642.20","2":"20.59575","3":"20"},{"1":"661.95","2":"19.37500","3":"20"},{"1":"655.05","2":"19.95122","3":"20"},{"1":"663.90","2":"18.84973","3":"20"},{"1":"665.95","2":"18.11787","3":"20"},{"1":"695.30","2":"19.18341","3":"20"},{"1":"669.85","2":"22.00000","3":"20"},{"1":"679.75","2":"21.58416","3":"20"},{"1":"656.40","2":"20.38889","3":"20"},{"1":"644.55","2":"16.29310","3":"20"},{"1":"673.55","2":"18.27778","3":"20"},{"1":"651.35","2":"19.37472","3":"20"},{"1":"674.25","2":"18.90909","3":"20"},{"1":"660.95","2":"16.40693","3":"20"},{"1":"655.70","2":"15.59140","3":"20"},{"1":"644.70","2":"18.70694","3":"20"},{"1":"644.20","2":"18.32985","3":"20"},{"1":"638.30","2":"17.90235","3":"20"},{"1":"680.05","2":"18.91157","3":"20"},{"1":"698.45","2":"20.32497","3":"20"},{"1":"644.40","2":"20.02457","3":"20"},{"1":"657.15","2":"24.00000","3":"20"},{"1":"612.50","2":"17.60784","3":"20"},{"1":"638.30","2":"19.34853","3":"20"},{"1":"662.75","2":"19.67846","3":"20"},{"1":"658.60","2":"18.72861","3":"20"},{"1":"671.65","2":"15.88235","3":"20"},{"1":"663.45","2":"20.05491","3":"20"},{"1":"657.75","2":"17.98825","3":"20"},{"1":"634.55","2":"16.96629","3":"20"},{"1":"664.15","2":"19.23937","3":"20"},{"1":"648.20","2":"19.19586","3":"20"},{"1":"669.35","2":"19.59906","3":"20"},{"1":"663.35","2":"20.54348","3":"20"},{"1":"673.25","2":"18.58848","3":"20"},{"1":"663.50","2":"15.60419","3":"20"},{"1":"664.00","2":"15.29304","3":"20"},{"1":"664.75","2":"17.65537","3":"20"},{"1":"650.90","2":"17.57976","3":"20"},{"1":"666.55","2":"22.33333","3":"20"},{"1":"617.35","2":"18.75000","3":"20"},{"1":"652.35","2":"18.10241","3":"20"},{"1":"616.30","2":"20.25641","3":"20"},{"1":"630.55","2":"18.80207","3":"20"},{"1":"655.20","2":"18.77230","3":"20"},{"1":"671.90","2":"20.40521","3":"20"},{"1":"672.20","2":"18.65079","3":"20"},{"1":"652.30","2":"20.70707","3":"20"},{"1":"649.30","2":"22.00000","3":"20"},{"1":"645.55","2":"17.69978","3":"20"},{"1":"648.25","2":"21.48329","3":"20"},{"1":"640.10","2":"16.70103","3":"20"},{"1":"641.80","2":"19.57567","3":"20"},{"1":"632.25","2":"17.25806","3":"20"},{"1":"633.15","2":"17.37526","3":"20"},{"1":"647.05","2":"17.34931","3":"20"},{"1":"620.30","2":"16.26229","3":"20"},{"1":"625.85","2":"17.70045","3":"20"},{"1":"649.85","2":"20.12881","3":"20"},{"1":"681.60","2":"18.26539","3":"20"},{"1":"643.50","2":"14.54214","3":"20"},{"1":"632.45","2":"19.15261","3":"20"},{"1":"672.05","2":"17.36574","3":"20"},{"1":"655.35","2":"15.13898","3":"20"},{"1":"665.65","2":"17.84266","3":"20"},{"1":"668.90","2":"15.40704","3":"20"},{"1":"682.15","2":"18.86534","3":"20"},{"1":"637.95","2":"16.47413","3":"20"},{"1":"626.90","2":"17.86263","3":"20"},{"1":"661.90","2":"21.88586","3":"20"},{"1":"670.00","2":"20.20000","3":"20"},{"1":"682.55","2":"19.03640","3":"20"},{"1":"642.20","2":"17.88991","3":"21"},{"1":"650.45","2":"21.52466","3":"21"},{"1":"653.10","2":"18.69723","3":"21"},{"1":"646.70","2":"17.35714","3":"21"},{"1":"699.10","2":"18.67133","3":"21"},{"1":"661.90","2":"21.40625","3":"21"},{"1":"660.30","2":"19.50000","3":"21"},{"1":"665.10","2":"20.89412","3":"21"},{"1":"652.85","2":"19.94737","3":"21"},{"1":"643.25","2":"20.80556","3":"21"},{"1":"671.70","2":"21.23809","3":"21"},{"1":"619.80","2":"21.00000","3":"21"},{"1":"668.40","2":"20.60000","3":"21"},{"1":"642.75","2":"20.00822","3":"21"},{"1":"644.95","2":"18.02778","3":"21"},{"1":"665.90","2":"20.25196","3":"21"},{"1":"684.80","2":"16.97787","3":"21"},{"1":"671.30","2":"16.50980","3":"21"},{"1":"655.35","2":"22.70402","3":"21"},{"1":"680.45","2":"19.91111","3":"21"},{"1":"659.35","2":"18.33333","3":"21"},{"1":"662.70","2":"22.61905","3":"21"},{"1":"615.75","2":"19.44828","3":"21"},{"1":"643.05","2":"25.05263","3":"21"},{"1":"673.55","2":"20.67544","3":"21"},{"1":"655.30","2":"18.68235","3":"21"},{"1":"655.05","2":"22.84553","3":"21"},{"1":"634.05","2":"19.26667","3":"21"},{"1":"657.00","2":"19.25000","3":"21"},{"1":"669.35","2":"20.54545","3":"21"},{"1":"664.95","2":"20.60697","3":"21"},{"1":"635.60","2":"21.07268","3":"21"},{"1":"668.60","2":"21.53581","3":"21"},{"1":"628.25","2":"19.90400","3":"21"},{"1":"656.50","2":"21.19407","3":"21"},{"1":"634.00","2":"21.86535","3":"21"},{"1":"633.90","2":"18.32965","3":"21"},{"1":"653.50","2":"16.22857","3":"21"},{"1":"648.00","2":"19.17857","3":"21"},{"1":"660.05","2":"20.27737","3":"21"},{"1":"653.40","2":"22.98614","3":"21"},{"1":"635.60","2":"20.44444","3":"21"},{"1":"665.65","2":"19.82085","3":"21"},{"1":"661.95","2":"23.20522","3":"21"},{"1":"606.75","2":"19.26697","3":"21"},{"1":"634.95","2":"23.30189","3":"21"},{"1":"612.65","2":"21.18829","3":"21"},{"1":"664.15","2":"20.87180","3":"21"},{"1":"631.05","2":"19.01749","3":"21"},{"1":"638.30","2":"21.91938","3":"21"},{"1":"623.20","2":"20.10124","3":"21"},{"1":"641.45","2":"21.47651","3":"21"},{"1":"644.45","2":"20.06579","3":"21"},{"1":"674.25","2":"20.37510","3":"21"},{"1":"647.25","2":"22.44648","3":"21"},{"1":"653.95","2":"22.89524","3":"21"},{"1":"666.65","2":"20.49797","3":"21"},{"1":"681.30","2":"20.00000","3":"21"},{"1":"664.70","2":"22.25658","3":"21"},{"1":"633.15","2":"21.56436","3":"21"},{"1":"616.30","2":"19.47737","3":"21"},{"1":"668.90","2":"17.67002","3":"21"},{"1":"628.55","2":"21.94756","3":"21"},{"1":"640.15","2":"21.78339","3":"21"},{"1":"620.30","2":"19.14000","3":"21"},{"1":"644.50","2":"18.11050","3":"21"},{"1":"668.95","2":"20.68242","3":"21"},{"1":"676.95","2":"22.62361","3":"21"},{"1":"698.25","2":"21.78650","3":"21"},{"1":"627.10","2":"18.58293","3":"21"},{"1":"643.70","2":"21.54545","3":"21"},{"1":"632.95","2":"21.15289","3":"21"},{"1":"668.10","2":"16.63333","3":"21"},{"1":"679.80","2":"21.14438","3":"21"},{"1":"628.65","2":"19.78182","3":"21"},{"1":"678.05","2":"18.98373","3":"21"},{"1":"666.65","2":"17.66767","3":"21"},{"1":"679.40","2":"17.75499","3":"21"},{"1":"664.40","2":"15.27273","3":"21"},{"1":"666.60","2":"14.00000","3":"21"},{"1":"625.85","2":"20.59613","3":"21"},{"1":"691.90","2":"16.31169","3":"21"},{"1":"645.75","2":"21.12796","3":"21"},{"1":"652.40","2":"17.48801","3":"21"},{"1":"633.05","2":"17.88679","3":"21"},{"1":"640.85","2":"19.30676","3":"21"},{"1":"671.90","2":"20.89231","3":"21"},{"1":"669.10","2":"21.28684","3":"21"},{"1":"626.10","2":"20.19560","3":"21"},{"1":"633.65","2":"24.95000","3":"21"},{"1":"659.40","2":"18.13043","3":"21"},{"1":"654.20","2":"20.00000","3":"21"},{"1":"671.25","2":"18.72951","3":"21"},{"1":"706.75","2":"18.25000","3":"21"},{"1":"682.65","2":"18.99257","3":"21"},{"1":"635.95","2":"19.88764","3":"21"},{"1":"649.30","2":"19.37895","3":"21"},{"1":"666.85","2":"20.46259","3":"21"},{"1":"639.50","2":"22.29157","3":"21"},{"1":"660.95","2":"20.70474","3":"21"},{"1":"661.45","2":"19.06005","3":"21"},{"1":"666.05","2":"20.23247","3":"21"},{"1":"642.20","2":"19.69012","3":"21"},{"1":"658.80","2":"20.36254","3":"21"},{"1":"634.40","2":"19.75422","3":"21"},{"1":"664.00","2":"19.37977","3":"21"},{"1":"679.65","2":"22.92351","3":"21"},{"1":"646.40","2":"19.37340","3":"21"},{"1":"652.10","2":"19.15516","3":"21"},{"1":"643.50","2":"21.30000","3":"21"},{"1":"631.90","2":"18.30357","3":"21"},{"1":"637.95","2":"21.07926","3":"21"},{"1":"656.65","2":"18.79121","3":"21"},{"1":"671.75","2":"19.62662","3":"21"},{"1":"646.35","2":"19.59016","3":"21"},{"1":"637.65","2":"20.87187","3":"21"},{"1":"677.25","2":"21.11500","3":"21"},{"1":"666.85","2":"20.08452","3":"21"},{"1":"691.05","2":"19.91049","3":"21"},{"1":"660.20","2":"17.81285","3":"21"},{"1":"632.20","2":"18.13333","3":"21"},{"1":"662.90","2":"19.22221","3":"21"},{"1":"652.30","2":"18.66072","3":"21"},{"1":"635.45","2":"19.60000","3":"21"},{"1":"626.90","2":"19.28384","3":"21"},{"1":"672.70","2":"22.81818","3":"21"},{"1":"673.05","2":"18.80922","3":"21"},{"1":"676.15","2":"21.37363","3":"21"},{"1":"660.75","2":"20.02041","3":"21"},{"1":"664.95","2":"21.49862","3":"21"},{"1":"657.55","2":"15.42857","3":"21"},{"1":"657.75","2":"22.40000","3":"21"},{"1":"647.05","2":"20.12709","3":"21"},{"1":"624.55","2":"19.03798","3":"21"},{"1":"671.65","2":"17.34216","3":"21"},{"1":"623.45","2":"17.01863","3":"21"},{"1":"677.95","2":"20.80000","3":"21"},{"1":"673.90","2":"21.15385","3":"21"},{"1":"646.90","2":"18.45833","3":"21"},{"1":"669.80","2":"19.14082","3":"21"},{"1":"634.10","2":"19.40766","3":"21"},{"1":"663.50","2":"19.56896","3":"21"},{"1":"622.60","2":"21.50120","3":"21"},{"1":"644.70","2":"17.52941","3":"21"},{"1":"623.60","2":"16.43017","3":"21"},{"1":"695.20","2":"19.79654","3":"21"},{"1":"678.80","2":"17.18613","3":"21"},{"1":"662.40","2":"17.61589","3":"21"},{"1":"665.35","2":"20.12537","3":"21"},{"1":"646.20","2":"22.16667","3":"21"},{"1":"668.80","2":"19.96154","3":"21"},{"1":"664.75","2":"19.03945","3":"21"},{"1":"689.10","2":"15.22436","3":"21"},{"1":"622.05","2":"21.14475","3":"21"},{"1":"655.70","2":"19.64390","3":"21"},{"1":"645.75","2":"21.04869","3":"21"},{"1":"668.65","2":"20.17544","3":"21"},{"1":"651.85","2":"21.39130","3":"21"},{"1":"646.00","2":"20.00833","3":"21"},{"1":"669.30","2":"20.29137","3":"21"},{"1":"632.00","2":"17.66667","3":"21"},{"1":"655.35","2":"18.22055","3":"21"},{"1":"694.80","2":"20.27100","3":"21"},{"1":"672.35","2":"20.19895","3":"21"},{"1":"640.90","2":"21.38424","3":"21"},{"1":"643.20","2":"20.97368","3":"21"},{"1":"698.20","2":"20.00000","3":"21"},{"1":"618.05","2":"17.15328","3":"21"},{"1":"684.30","2":"22.34977","3":"21"},{"1":"644.55","2":"22.17007","3":"21"},{"1":"684.95","2":"18.18182","3":"21"},{"1":"654.85","2":"18.95714","3":"21"},{"1":"646.55","2":"19.74533","3":"21"},{"1":"651.15","2":"16.42623","3":"21"},{"1":"651.90","2":"16.62540","3":"21"},{"1":"652.30","2":"16.38177","3":"21"},{"1":"616.45","2":"20.07416","3":"21"},{"1":"656.40","2":"17.99544","3":"21"},{"1":"644.20","2":"19.39130","3":"21"},{"1":"634.70","2":"16.42857","3":"21"},{"1":"639.85","2":"16.72949","3":"21"},{"1":"649.85","2":"24.41345","3":"21"},{"1":"672.30","2":"18.26415","3":"21"},{"1":"645.00","2":"18.95504","3":"21"},{"1":"630.55","2":"21.03896","3":"21"},{"1":"665.95","2":"20.74074","3":"21"},{"1":"650.55","2":"18.10000","3":"21"},{"1":"700.30","2":"19.84615","3":"21"},{"1":"673.55","2":"21.60000","3":"21"},{"1":"639.90","2":"22.44242","3":"21"},{"1":"631.85","2":"23.01438","3":"21"},{"1":"657.50","2":"17.74892","3":"21"},{"1":"637.35","2":"18.28664","3":"21"},{"1":"638.35","2":"19.26544","3":"21"},{"1":"626.80","2":"22.66667","3":"21"},{"1":"666.15","2":"19.29412","3":"21"},{"1":"666.10","2":"17.36364","3":"21"},{"1":"655.40","2":"19.82143","3":"21"},{"1":"659.90","2":"20.43378","3":"21"},{"1":"657.40","2":"21.03721","3":"21"},{"1":"658.60","2":"19.92462","3":"21"},{"1":"630.35","2":"19.00986","3":"21"},{"1":"678.40","2":"23.82222","3":"21"},{"1":"663.45","2":"19.36909","3":"21"},{"1":"635.20","2":"19.82857","3":"21"},{"1":"636.95","2":"15.25885","3":"21"},{"1":"671.60","2":"17.16129","3":"21"},{"1":"653.55","2":"21.81333","3":"21"},{"1":"659.05","2":"19.07471","3":"21"},{"1":"696.55","2":"25.78512","3":"21"},{"1":"669.35","2":"18.21261","3":"21"},{"1":"661.60","2":"18.16606","3":"21"},{"1":"662.55","2":"16.97297","3":"21"},{"1":"658.00","2":"21.50087","3":"21"},{"1":"666.45","2":"20.60000","3":"21"},{"1":"666.70","2":"16.99029","3":"21"},{"1":"653.80","2":"20.77954","3":"21"},{"1":"690.80","2":"15.51247","3":"21"},{"1":"651.20","2":"19.88506","3":"21"},{"1":"659.40","2":"21.39882","3":"21"},{"1":"655.75","2":"20.49751","3":"21"},{"1":"635.05","2":"19.36376","3":"21"},{"1":"652.50","2":"17.65957","3":"21"},{"1":"670.00","2":"21.01796","3":"21"},{"1":"644.40","2":"19.05565","3":"21"},{"1":"681.60","2":"22.53846","3":"21"},{"1":"636.10","2":"21.10787","3":"21"},{"1":"645.55","2":"20.05135","3":"21"},{"1":"686.70","2":"14.20176","3":"21"},{"1":"652.00","2":"18.47687","3":"21"},{"1":"666.15","2":"18.63542","3":"21"},{"1":"662.50","2":"20.94595","3":"21"},{"1":"647.60","2":"21.08548","3":"21"},{"1":"676.55","2":"18.69288","3":"21"},{"1":"698.45","2":"20.86808","3":"21"},{"1":"627.25","2":"19.82558","3":"21"},{"1":"648.95","2":"19.75000","3":"21"},{"1":"640.10","2":"19.50000","3":"21"},{"1":"655.05","2":"18.39080","3":"21"},{"1":"654.20","2":"18.78676","3":"21"},{"1":"671.60","2":"19.77018","3":"21"},{"1":"657.15","2":"19.33333","3":"21"},{"1":"638.55","2":"21.46392","3":"21"},{"1":"694.25","2":"23.08492","3":"21"},{"1":"643.60","2":"21.06299","3":"21"},{"1":"667.20","2":"18.68687","3":"21"},{"1":"682.45","2":"20.77024","3":"21"},{"1":"676.60","2":"19.30556","3":"21"},{"1":"662.65","2":"20.13280","3":"21"},{"1":"616.30","2":"20.66964","3":"21"},{"1":"655.80","2":"22.28155","3":"21"},{"1":"638.00","2":"20.60027","3":"21"},{"1":"682.15","2":"20.82734","3":"21"},{"1":"651.40","2":"19.22492","3":"21"},{"1":"641.55","2":"17.65477","3":"21"},{"1":"621.40","2":"17.00000","3":"21"},{"1":"648.25","2":"16.49773","3":"21"},{"1":"644.45","2":"19.78261","3":"21"},{"1":"643.70","2":"22.30216","3":"21"},{"1":"636.60","2":"17.73077","3":"21"},{"1":"652.35","2":"20.44836","3":"21"},{"1":"663.85","2":"20.37169","3":"21"},{"1":"634.55","2":"20.16479","3":"21"},{"1":"631.40","2":"21.61538","3":"21"},{"1":"643.40","2":"20.56143","3":"21"},{"1":"664.30","2":"19.95551","3":"21"},{"1":"663.90","2":"21.18387","3":"21"},{"1":"657.80","2":"18.81042","3":"21"},{"1":"669.95","2":"20.57838","3":"21"},{"1":"630.40","2":"18.32461","3":"21"},{"1":"664.15","2":"18.82063","3":"21"},{"1":"624.95","2":"20.81633","3":"21"},{"1":"664.45","2":"20.00000","3":"21"},{"1":"675.40","2":"19.68182","3":"21"},{"1":"691.35","2":"19.39018","3":"21"},{"1":"634.15","2":"20.92732","3":"21"},{"1":"676.85","2":"19.94437","3":"21"},{"1":"643.40","2":"20.79109","3":"21"},{"1":"635.75","2":"19.20354","3":"21"},{"1":"671.95","2":"19.02439","3":"21"},{"1":"650.90","2":"17.62058","3":"21"},{"1":"672.05","2":"20.23715","3":"21"},{"1":"649.15","2":"19.29374","3":"21"},{"1":"654.85","2":"18.82998","3":"21"},{"1":"665.20","2":"20.33949","3":"21"},{"1":"704.30","2":"19.22900","3":"21"},{"1":"637.00","2":"17.89130","3":"21"},{"1":"647.30","2":"19.51881","3":"21"},{"1":"661.35","2":"19.08451","3":"21"},{"1":"680.05","2":"19.93548","3":"21"},{"1":"652.10","2":"18.87326","3":"21"},{"1":"661.85","2":"20.14178","3":"21"},{"1":"646.95","2":"23.55637","3":"21"},{"1":"609.00","2":"21.46479","3":"21"},{"1":"660.10","2":"19.19101","3":"21"},{"1":"644.20","2":"20.13080","3":"21"},{"1":"636.90","2":"25.80000","3":"21"},{"1":"632.25","2":"18.77774","3":"21"},{"1":"632.45","2":"19.10982","3":"21"},{"1":"661.85","2":"19.70109","3":"21"},{"1":"640.75","2":"18.61594","3":"21"},{"1":"662.55","2":"20.99721","3":"21"},{"1":"672.05","2":"20.00000","3":"21"},{"1":"693.95","2":"20.98325","3":"21"},{"1":"681.30","2":"21.64262","3":"21"},{"1":"669.85","2":"20.02967","3":"21"},{"1":"658.35","2":"19.81140","3":"21"},{"1":"670.70","2":"18.00000","3":"21"},{"1":"686.05","2":"19.35811","3":"21"},{"1":"655.55","2":"20.17912","3":"21"},{"1":"656.55","2":"21.11986","3":"21"},{"1":"682.55","2":"23.38974","3":"21"},{"1":"612.50","2":"22.18182","3":"21"},{"1":"639.35","2":"19.94283","3":"21"},{"1":"641.10","2":"17.78826","3":"21"},{"1":"654.90","2":"14.70588","3":"21"},{"1":"661.20","2":"19.04077","3":"21"},{"1":"653.55","2":"20.89195","3":"21"},{"1":"637.10","2":"19.83851","3":"21"},{"1":"651.45","2":"19.52191","3":"21"},{"1":"672.55","2":"20.68622","3":"21"},{"1":"662.75","2":"18.18182","3":"21"},{"1":"655.20","2":"18.89224","3":"21"},{"1":"655.05","2":"24.88889","3":"21"},{"1":"639.30","2":"18.58064","3":"21"},{"1":"669.30","2":"18.04000","3":"21"},{"1":"661.85","2":"17.73399","3":"21"},{"1":"663.35","2":"21.45455","3":"21"},{"1":"655.85","2":"19.92343","3":"21"},{"1":"638.70","2":"20.33942","3":"21"},{"1":"653.85","2":"22.54608","3":"21"},{"1":"632.85","2":"21.10344","3":"21"},{"1":"621.75","2":"18.19743","3":"21"},{"1":"656.70","2":"20.10768","3":"21"},{"1":"667.15","2":"19.15984","3":"21"},{"1":"648.35","2":"19.54545","3":"21"},{"1":"639.75","2":"20.88889","3":"21"},{"1":"654.30","2":"18.39150","3":"21"},{"1":"647.60","2":"19.17990","3":"21"},{"1":"684.35","2":"19.39771","3":"21"},{"1":"681.90","2":"21.67827","3":"21"},{"1":"672.45","2":"19.28889","3":"21"},{"1":"645.55","2":"20.34927","3":"21"},{"1":"656.80","2":"20.96416","3":"21"},{"1":"638.30","2":"19.46039","3":"21"},{"1":"650.90","2":"19.28572","3":"21"},{"1":"645.60","2":"20.91979","3":"21"},{"1":"605.55","2":"20.90021","3":"21"},{"1":"656.80","2":"20.59575","3":"21"},{"1":"652.40","2":"19.37500","3":"21"},{"1":"687.55","2":"19.95122","3":"21"},{"1":"659.15","2":"18.84973","3":"21"},{"1":"675.70","2":"18.11787","3":"21"},{"1":"646.50","2":"19.18341","3":"21"},{"1":"683.35","2":"22.00000","3":"21"},{"1":"630.55","2":"21.58416","3":"21"},{"1":"654.60","2":"20.38889","3":"21"},{"1":"643.50","2":"16.29310","3":"21"},{"1":"673.25","2":"18.27778","3":"21"},{"1":"657.90","2":"19.37472","3":"21"},{"1":"636.50","2":"18.90909","3":"21"},{"1":"695.30","2":"16.40693","3":"21"},{"1":"657.65","2":"15.59140","3":"21"},{"1":"634.10","2":"18.70694","3":"21"},{"1":"651.35","2":"18.32985","3":"21"},{"1":"653.70","2":"17.90235","3":"21"},{"1":"659.80","2":"18.91157","3":"21"},{"1":"618.30","2":"20.32497","3":"21"},{"1":"672.20","2":"20.02457","3":"21"},{"1":"627.30","2":"24.00000","3":"21"},{"1":"636.70","2":"17.60784","3":"21"},{"1":"641.45","2":"19.34853","3":"21"},{"1":"663.85","2":"19.67846","3":"21"},{"1":"639.25","2":"18.72861","3":"21"},{"1":"664.00","2":"15.88235","3":"21"},{"1":"623.10","2":"20.05491","3":"21"},{"1":"631.95","2":"17.98825","3":"21"},{"1":"628.40","2":"16.96629","3":"21"},{"1":"650.65","2":"19.23937","3":"21"},{"1":"640.50","2":"19.19586","3":"21"},{"1":"647.70","2":"19.59906","3":"21"},{"1":"642.40","2":"20.54348","3":"21"},{"1":"662.45","2":"18.58848","3":"21"},{"1":"639.80","2":"15.60419","3":"21"},{"1":"667.60","2":"15.29304","3":"21"},{"1":"673.30","2":"17.65537","3":"21"},{"1":"661.90","2":"17.57976","3":"21"},{"1":"628.75","2":"22.33333","3":"21"},{"1":"651.80","2":"18.75000","3":"21"},{"1":"668.00","2":"18.10241","3":"21"},{"1":"679.75","2":"20.25641","3":"21"},{"1":"666.55","2":"18.80207","3":"21"},{"1":"638.20","2":"18.77230","3":"21"},{"1":"617.35","2":"20.40521","3":"21"},{"1":"620.50","2":"18.65079","3":"21"},{"1":"661.60","2":"20.70707","3":"21"},{"1":"654.10","2":"22.00000","3":"21"},{"1":"648.70","2":"17.69978","3":"21"},{"1":"634.90","2":"21.48329","3":"21"},{"1":"645.10","2":"16.70103","3":"21"},{"1":"649.50","2":"19.57567","3":"21"},{"1":"625.30","2":"17.25806","3":"21"},{"1":"671.90","2":"17.37526","3":"21"},{"1":"648.20","2":"17.34931","3":"21"},{"1":"616.30","2":"16.26229","3":"21"},{"1":"657.00","2":"17.70045","3":"21"},{"1":"683.40","2":"20.12881","3":"21"},{"1":"662.40","2":"18.26539","3":"21"},{"1":"629.80","2":"14.54214","3":"21"},{"1":"645.25","2":"19.15261","3":"21"},{"1":"666.00","2":"17.36574","3":"21"},{"1":"624.15","2":"15.13898","3":"21"},{"1":"641.80","2":"17.84266","3":"21"},{"1":"649.70","2":"15.40704","3":"21"},{"1":"679.50","2":"18.86534","3":"21"},{"1":"650.60","2":"16.47413","3":"21"},{"1":"637.95","2":"17.86263","3":"21"},{"1":"634.20","2":"21.88586","3":"21"},{"1":"667.45","2":"20.20000","3":"21"},{"1":"667.45","2":"19.03640","3":"21"},{"1":"639.25","2":"17.88991","3":"22"},{"1":"651.80","2":"21.52466","3":"22"},{"1":"671.90","2":"18.69723","3":"22"},{"1":"645.25","2":"17.35714","3":"22"},{"1":"691.35","2":"18.67133","3":"22"},{"1":"660.20","2":"21.40625","3":"22"},{"1":"676.15","2":"19.50000","3":"22"},{"1":"677.25","2":"20.89412","3":"22"},{"1":"671.90","2":"19.94737","3":"22"},{"1":"671.60","2":"20.80556","3":"22"},{"1":"650.90","2":"21.23809","3":"22"},{"1":"655.75","2":"21.00000","3":"22"},{"1":"693.95","2":"20.60000","3":"22"},{"1":"644.95","2":"20.00822","3":"22"},{"1":"655.05","2":"18.02778","3":"22"},{"1":"637.35","2":"20.25196","3":"22"},{"1":"645.75","2":"16.97787","3":"22"},{"1":"666.45","2":"16.50980","3":"22"},{"1":"696.55","2":"22.70402","3":"22"},{"1":"633.65","2":"19.91111","3":"22"},{"1":"636.60","2":"18.33333","3":"22"},{"1":"698.45","2":"22.61905","3":"22"},{"1":"637.95","2":"19.44828","3":"22"},{"1":"678.40","2":"25.05263","3":"22"},{"1":"634.55","2":"20.67544","3":"22"},{"1":"673.55","2":"18.68235","3":"22"},{"1":"667.45","2":"22.84553","3":"22"},{"1":"660.75","2":"19.26667","3":"22"},{"1":"617.35","2":"19.25000","3":"22"},{"1":"668.10","2":"20.54545","3":"22"},{"1":"663.35","2":"20.60697","3":"22"},{"1":"669.10","2":"21.07268","3":"22"},{"1":"681.30","2":"21.53581","3":"22"},{"1":"682.45","2":"19.90400","3":"22"},{"1":"659.80","2":"21.19407","3":"22"},{"1":"666.85","2":"21.86535","3":"22"},{"1":"645.60","2":"18.32965","3":"22"},{"1":"636.95","2":"16.22857","3":"22"},{"1":"635.05","2":"19.17857","3":"22"},{"1":"657.00","2":"20.27737","3":"22"},{"1":"641.10","2":"22.98614","3":"22"},{"1":"624.95","2":"20.44444","3":"22"},{"1":"652.00","2":"19.82085","3":"22"},{"1":"657.50","2":"23.20522","3":"22"},{"1":"653.50","2":"19.26697","3":"22"},{"1":"646.40","2":"23.30189","3":"22"},{"1":"671.65","2":"21.18829","3":"22"},{"1":"655.05","2":"20.87180","3":"22"},{"1":"651.35","2":"19.01749","3":"22"},{"1":"637.95","2":"21.91938","3":"22"},{"1":"631.85","2":"20.10124","3":"22"},{"1":"641.45","2":"21.47651","3":"22"},{"1":"666.15","2":"20.06579","3":"22"},{"1":"657.75","2":"20.37510","3":"22"},{"1":"648.20","2":"22.44648","3":"22"},{"1":"630.55","2":"22.89524","3":"22"},{"1":"680.45","2":"20.49797","3":"22"},{"1":"659.15","2":"20.00000","3":"22"},{"1":"657.80","2":"22.25658","3":"22"},{"1":"672.20","2":"21.56436","3":"22"},{"1":"654.85","2":"19.47737","3":"22"},{"1":"638.55","2":"17.67002","3":"22"},{"1":"632.95","2":"21.94756","3":"22"},{"1":"691.90","2":"21.78339","3":"22"},{"1":"638.35","2":"19.14000","3":"22"},{"1":"646.50","2":"18.11050","3":"22"},{"1":"661.20","2":"20.68242","3":"22"},{"1":"647.60","2":"22.62361","3":"22"},{"1":"639.30","2":"21.78650","3":"22"},{"1":"628.75","2":"18.58293","3":"22"},{"1":"668.95","2":"21.54545","3":"22"},{"1":"626.80","2":"21.15289","3":"22"},{"1":"665.20","2":"16.63333","3":"22"},{"1":"635.20","2":"21.14438","3":"22"},{"1":"657.65","2":"19.78182","3":"22"},{"1":"663.45","2":"18.98373","3":"22"},{"1":"662.65","2":"17.66767","3":"22"},{"1":"654.85","2":"17.75499","3":"22"},{"1":"662.70","2":"15.27273","3":"22"},{"1":"695.20","2":"14.00000","3":"22"},{"1":"639.35","2":"20.59613","3":"22"},{"1":"629.80","2":"16.31169","3":"22"},{"1":"661.85","2":"21.12796","3":"22"},{"1":"690.80","2":"17.48801","3":"22"},{"1":"660.30","2":"17.88679","3":"22"},{"1":"622.60","2":"19.30676","3":"22"},{"1":"653.95","2":"20.89231","3":"22"},{"1":"673.05","2":"21.28684","3":"22"},{"1":"634.10","2":"20.19560","3":"22"},{"1":"655.55","2":"24.95000","3":"22"},{"1":"662.90","2":"18.13043","3":"22"},{"1":"655.20","2":"20.00000","3":"22"},{"1":"663.85","2":"18.72951","3":"22"},{"1":"678.80","2":"18.25000","3":"22"},{"1":"683.40","2":"18.99257","3":"22"},{"1":"605.55","2":"19.88764","3":"22"},{"1":"616.45","2":"19.37895","3":"22"},{"1":"649.70","2":"20.46259","3":"22"},{"1":"645.55","2":"22.29157","3":"22"},{"1":"655.05","2":"20.70474","3":"22"},{"1":"649.50","2":"19.06005","3":"22"},{"1":"640.15","2":"20.23247","3":"22"},{"1":"622.05","2":"19.69012","3":"22"},{"1":"664.40","2":"20.36254","3":"22"},{"1":"615.75","2":"19.75422","3":"22"},{"1":"658.00","2":"19.37977","3":"22"},{"1":"635.45","2":"22.92351","3":"22"},{"1":"630.35","2":"19.37340","3":"22"},{"1":"636.90","2":"19.15516","3":"22"},{"1":"675.40","2":"21.30000","3":"22"},{"1":"674.25","2":"18.30357","3":"22"},{"1":"628.25","2":"21.07926","3":"22"},{"1":"646.70","2":"18.79121","3":"22"},{"1":"653.55","2":"19.62662","3":"22"},{"1":"620.30","2":"19.59016","3":"22"},{"1":"695.30","2":"20.87187","3":"22"},{"1":"651.40","2":"21.11500","3":"22"},{"1":"654.60","2":"20.08452","3":"22"},{"1":"647.70","2":"19.91049","3":"22"},{"1":"606.75","2":"17.81285","3":"22"},{"1":"627.25","2":"18.13333","3":"22"},{"1":"634.10","2":"19.22221","3":"22"},{"1":"643.50","2":"18.66072","3":"22"},{"1":"643.50","2":"19.60000","3":"22"},{"1":"644.20","2":"19.28384","3":"22"},{"1":"659.40","2":"22.81818","3":"22"},{"1":"700.30","2":"18.80922","3":"22"},{"1":"678.05","2":"21.37363","3":"22"},{"1":"666.05","2":"20.02041","3":"22"},{"1":"664.95","2":"21.49862","3":"22"},{"1":"632.85","2":"15.42857","3":"22"},{"1":"655.85","2":"22.40000","3":"22"},{"1":"647.25","2":"20.12709","3":"22"},{"1":"648.00","2":"19.03798","3":"22"},{"1":"665.10","2":"17.34216","3":"22"},{"1":"662.40","2":"17.01863","3":"22"},{"1":"664.95","2":"20.80000","3":"22"},{"1":"669.80","2":"21.15385","3":"22"},{"1":"628.40","2":"18.45833","3":"22"},{"1":"670.00","2":"19.14082","3":"22"},{"1":"618.30","2":"19.40766","3":"22"},{"1":"654.20","2":"19.56896","3":"22"},{"1":"634.05","2":"21.50120","3":"22"},{"1":"652.40","2":"17.52941","3":"22"},{"1":"632.00","2":"16.43017","3":"22"},{"1":"624.15","2":"19.79654","3":"22"},{"1":"631.05","2":"17.18613","3":"22"},{"1":"679.40","2":"17.61589","3":"22"},{"1":"672.30","2":"20.12537","3":"22"},{"1":"667.45","2":"22.16667","3":"22"},{"1":"657.90","2":"19.96154","3":"22"},{"1":"663.85","2":"19.03945","3":"22"},{"1":"644.50","2":"15.22436","3":"22"},{"1":"641.80","2":"21.14475","3":"22"},{"1":"672.05","2":"19.64390","3":"22"},{"1":"624.55","2":"21.04869","3":"22"},{"1":"621.40","2":"20.17544","3":"22"},{"1":"620.50","2":"21.39130","3":"22"},{"1":"654.30","2":"20.00833","3":"22"},{"1":"680.05","2":"20.29137","3":"22"},{"1":"650.65","2":"17.66667","3":"22"},{"1":"630.55","2":"18.22055","3":"22"},{"1":"623.60","2":"20.27100","3":"22"},{"1":"673.90","2":"20.19895","3":"22"},{"1":"653.85","2":"21.38424","3":"22"},{"1":"644.20","2":"20.97368","3":"22"},{"1":"664.30","2":"20.00000","3":"22"},{"1":"671.70","2":"17.15328","3":"22"},{"1":"694.80","2":"22.34977","3":"22"},{"1":"650.45","2":"22.17007","3":"22"},{"1":"689.10","2":"18.18182","3":"22"},{"1":"682.15","2":"18.95714","3":"22"},{"1":"664.75","2":"19.74533","3":"22"},{"1":"638.30","2":"16.42623","3":"22"},{"1":"612.65","2":"16.62540","3":"22"},{"1":"660.10","2":"16.38177","3":"22"},{"1":"642.20","2":"20.07416","3":"22"},{"1":"639.90","2":"17.99544","3":"22"},{"1":"684.80","2":"19.39130","3":"22"},{"1":"644.70","2":"16.42857","3":"22"},{"1":"651.90","2":"16.72949","3":"22"},{"1":"639.80","2":"24.41345","3":"22"},{"1":"666.65","2":"18.26415","3":"22"},{"1":"679.80","2":"18.95504","3":"22"},{"1":"682.65","2":"21.03896","3":"22"},{"1":"652.10","2":"20.74074","3":"22"},{"1":"616.30","2":"18.10000","3":"22"},{"1":"643.25","2":"19.84615","3":"22"},{"1":"671.30","2":"21.60000","3":"22"},{"1":"634.15","2":"22.44242","3":"22"},{"1":"664.15","2":"23.01438","3":"22"},{"1":"637.65","2":"17.74892","3":"22"},{"1":"658.35","2":"18.28664","3":"22"},{"1":"660.05","2":"19.26544","3":"22"},{"1":"633.90","2":"22.66667","3":"22"},{"1":"661.60","2":"19.29412","3":"22"},{"1":"646.00","2":"17.36364","3":"22"},{"1":"672.70","2":"19.82143","3":"22"},{"1":"664.15","2":"20.43378","3":"22"},{"1":"666.65","2":"21.03721","3":"22"},{"1":"646.35","2":"19.92462","3":"22"},{"1":"655.30","2":"19.00986","3":"22"},{"1":"644.45","2":"23.82222","3":"22"},{"1":"665.95","2":"19.36909","3":"22"},{"1":"651.45","2":"19.82857","3":"22"},{"1":"632.45","2":"15.25885","3":"22"},{"1":"691.05","2":"17.16129","3":"22"},{"1":"679.75","2":"21.81333","3":"22"},{"1":"656.70","2":"19.07471","3":"22"},{"1":"668.90","2":"25.78512","3":"22"},{"1":"662.45","2":"18.21261","3":"22"},{"1":"656.80","2":"18.16606","3":"22"},{"1":"640.50","2":"16.97297","3":"22"},{"1":"671.95","2":"21.50087","3":"22"},{"1":"639.85","2":"20.60000","3":"22"},{"1":"661.85","2":"16.99029","3":"22"},{"1":"632.20","2":"20.77954","3":"22"},{"1":"628.65","2":"15.51247","3":"22"},{"1":"675.70","2":"19.88506","3":"22"},{"1":"661.45","2":"21.39882","3":"22"},{"1":"669.85","2":"20.49751","3":"22"},{"1":"650.55","2":"19.36376","3":"22"},{"1":"655.35","2":"17.65957","3":"22"},{"1":"669.35","2":"21.01796","3":"22"},{"1":"656.80","2":"19.05565","3":"22"},{"1":"669.30","2":"22.53846","3":"22"},{"1":"640.90","2":"21.10787","3":"22"},{"1":"662.55","2":"20.05135","3":"22"},{"1":"653.80","2":"14.20176","3":"22"},{"1":"681.60","2":"18.47687","3":"22"},{"1":"637.00","2":"18.63542","3":"22"},{"1":"699.10","2":"20.94595","3":"22"},{"1":"651.85","2":"21.08548","3":"22"},{"1":"652.35","2":"18.69288","3":"22"},{"1":"704.30","2":"20.86808","3":"22"},{"1":"656.50","2":"19.82558","3":"22"},{"1":"665.35","2":"19.75000","3":"22"},{"1":"644.40","2":"19.50000","3":"22"},{"1":"664.00","2":"18.39080","3":"22"},{"1":"650.60","2":"18.78676","3":"22"},{"1":"652.85","2":"19.77018","3":"22"},{"1":"653.55","2":"19.33333","3":"22"},{"1":"609.00","2":"21.46392","3":"22"},{"1":"683.35","2":"23.08492","3":"22"},{"1":"679.50","2":"21.06299","3":"22"},{"1":"647.30","2":"18.68687","3":"22"},{"1":"666.15","2":"20.77024","3":"22"},{"1":"634.95","2":"19.30556","3":"22"},{"1":"676.85","2":"20.13280","3":"22"},{"1":"698.25","2":"20.66964","3":"22"},{"1":"636.70","2":"22.28155","3":"22"},{"1":"632.25","2":"20.60027","3":"22"},{"1":"661.90","2":"20.82734","3":"22"},{"1":"663.50","2":"19.22492","3":"22"},{"1":"652.40","2":"17.65477","3":"22"},{"1":"656.65","2":"17.00000","3":"22"},{"1":"646.90","2":"16.49773","3":"22"},{"1":"635.95","2":"19.78261","3":"22"},{"1":"652.10","2":"22.30216","3":"22"},{"1":"647.60","2":"17.73077","3":"22"},{"1":"665.65","2":"20.44836","3":"22"},{"1":"642.20","2":"20.37169","3":"22"},{"1":"672.35","2":"20.16479","3":"22"},{"1":"652.30","2":"21.61538","3":"22"},{"1":"659.40","2":"20.56143","3":"22"},{"1":"643.60","2":"19.95551","3":"22"},{"1":"656.55","2":"21.18387","3":"22"},{"1":"669.95","2":"18.81042","3":"22"},{"1":"646.20","2":"20.57838","3":"22"},{"1":"649.30","2":"18.32461","3":"22"},{"1":"672.55","2":"18.82063","3":"22"},{"1":"616.30","2":"20.81633","3":"22"},{"1":"659.90","2":"20.00000","3":"22"},{"1":"654.90","2":"19.68182","3":"22"},{"1":"634.90","2":"19.39018","3":"22"},{"1":"648.35","2":"20.92732","3":"22"},{"1":"666.85","2":"19.94437","3":"22"},{"1":"666.00","2":"20.79109","3":"22"},{"1":"643.20","2":"19.20354","3":"22"},{"1":"661.90","2":"19.02439","3":"22"},{"1":"684.35","2":"17.62058","3":"22"},{"1":"618.05","2":"20.23715","3":"22"},{"1":"639.50","2":"19.29374","3":"22"},{"1":"673.55","2":"18.82998","3":"22"},{"1":"644.55","2":"20.33949","3":"22"},{"1":"646.95","2":"19.22900","3":"22"},{"1":"666.70","2":"17.89130","3":"22"},{"1":"640.10","2":"19.51881","3":"22"},{"1":"653.70","2":"19.08451","3":"22"},{"1":"698.20","2":"19.93548","3":"22"},{"1":"672.05","2":"18.87326","3":"22"},{"1":"661.85","2":"20.14178","3":"22"},{"1":"645.55","2":"23.55637","3":"22"},{"1":"673.25","2":"21.46479","3":"22"},{"1":"616.30","2":"19.19101","3":"22"},{"1":"652.30","2":"20.13080","3":"22"},{"1":"626.90","2":"25.80000","3":"22"},{"1":"627.10","2":"18.77774","3":"22"},{"1":"664.70","2":"19.10982","3":"22"},{"1":"686.70","2":"19.70109","3":"22"},{"1":"636.50","2":"18.61594","3":"22"},{"1":"662.75","2":"20.99721","3":"22"},{"1":"634.70","2":"20.00000","3":"22"},{"1":"630.40","2":"20.98325","3":"22"},{"1":"612.50","2":"21.64262","3":"22"},{"1":"667.15","2":"20.02967","3":"22"},{"1":"677.95","2":"19.81140","3":"22"},{"1":"657.55","2":"18.00000","3":"22"},{"1":"649.85","2":"19.35811","3":"22"},{"1":"668.00","2":"20.17912","3":"22"},{"1":"667.20","2":"21.11986","3":"22"},{"1":"638.00","2":"23.38974","3":"22"},{"1":"659.35","2":"22.18182","3":"22"},{"1":"641.55","2":"19.94283","3":"22"},{"1":"634.00","2":"17.78826","3":"22"},{"1":"631.95","2":"14.70588","3":"22"},{"1":"657.40","2":"19.04077","3":"22"},{"1":"627.30","2":"20.89195","3":"22"},{"1":"681.90","2":"19.83851","3":"22"},{"1":"650.90","2":"19.52191","3":"22"},{"1":"638.20","2":"20.68622","3":"22"},{"1":"625.85","2":"18.18182","3":"22"},{"1":"669.35","2":"18.89224","3":"22"},{"1":"643.70","2":"24.88889","3":"22"},{"1":"666.10","2":"18.58064","3":"22"},{"1":"671.25","2":"18.04000","3":"22"},{"1":"645.10","2":"17.73399","3":"22"},{"1":"659.05","2":"21.45455","3":"22"},{"1":"623.45","2":"19.92343","3":"22"},{"1":"654.20","2":"20.33942","3":"22"},{"1":"651.15","2":"22.54608","3":"22"},{"1":"682.55","2":"21.10344","3":"22"},{"1":"655.70","2":"18.19743","3":"22"},{"1":"635.75","2":"20.10768","3":"22"},{"1":"639.75","2":"19.15984","3":"22"},{"1":"686.05","2":"19.54545","3":"22"},{"1":"640.75","2":"20.88889","3":"22"},{"1":"655.80","2":"18.39150","3":"22"},{"1":"706.75","2":"19.17990","3":"22"},{"1":"655.35","2":"19.39771","3":"22"},{"1":"628.55","2":"21.67827","3":"22"},{"1":"652.50","2":"19.28889","3":"22"},{"1":"647.05","2":"20.34927","3":"22"},{"1":"676.95","2":"20.96416","3":"22"},{"1":"666.55","2":"19.46039","3":"22"},{"1":"666.60","2":"19.28572","3":"22"},{"1":"636.10","2":"20.91979","3":"22"},{"1":"661.95","2":"20.90021","3":"22"},{"1":"667.60","2":"20.59575","3":"22"},{"1":"662.55","2":"19.37500","3":"22"},{"1":"671.60","2":"19.95122","3":"22"},{"1":"619.80","2":"18.84973","3":"22"},{"1":"660.95","2":"18.11787","3":"22"},{"1":"676.55","2":"19.18341","3":"22"},{"1":"634.20","2":"22.00000","3":"22"},{"1":"631.90","2":"21.58416","3":"22"},{"1":"651.20","2":"20.38889","3":"22"},{"1":"643.40","2":"16.29310","3":"22"},{"1":"669.30","2":"18.27778","3":"22"},{"1":"687.55","2":"19.37472","3":"22"},{"1":"646.55","2":"18.90909","3":"22"},{"1":"638.70","2":"16.40693","3":"22"},{"1":"633.05","2":"15.59140","3":"22"},{"1":"663.90","2":"18.70694","3":"22"},{"1":"657.15","2":"18.32985","3":"22"},{"1":"648.95","2":"17.90235","3":"22"},{"1":"641.45","2":"18.91157","3":"22"},{"1":"648.25","2":"20.32497","3":"22"},{"1":"642.75","2":"20.02457","3":"22"},{"1":"644.45","2":"24.00000","3":"22"},{"1":"672.45","2":"17.60784","3":"22"},{"1":"661.60","2":"19.34853","3":"22"},{"1":"654.10","2":"19.67846","3":"22"},{"1":"668.65","2":"18.72861","3":"22"},{"1":"635.60","2":"15.88235","3":"22"},{"1":"653.10","2":"20.05491","3":"22"},{"1":"681.30","2":"17.98825","3":"22"},{"1":"625.30","2":"16.96629","3":"22"},{"1":"671.75","2":"19.23937","3":"22"},{"1":"621.75","2":"19.19586","3":"22"},{"1":"662.40","2":"19.59906","3":"22"},{"1":"648.70","2":"20.54348","3":"22"},{"1":"658.80","2":"18.58848","3":"22"},{"1":"662.50","2":"15.60419","3":"22"},{"1":"649.15","2":"15.29304","3":"22"},{"1":"665.90","2":"17.65537","3":"22"},{"1":"694.25","2":"17.57976","3":"22"},{"1":"635.60","2":"22.33333","3":"22"},{"1":"631.40","2":"18.75000","3":"22"},{"1":"668.40","2":"18.10241","3":"22"},{"1":"656.40","2":"20.25641","3":"22"},{"1":"679.65","2":"18.80207","3":"22"},{"1":"664.00","2":"18.77230","3":"22"},{"1":"661.35","2":"20.40521","3":"22"},{"1":"668.80","2":"18.65079","3":"22"},{"1":"634.40","2":"20.70707","3":"22"},{"1":"643.70","2":"22.00000","3":"22"},{"1":"658.60","2":"17.69978","3":"22"},{"1":"623.20","2":"21.48329","3":"22"},{"1":"637.10","2":"16.70103","3":"22"},{"1":"643.40","2":"19.57567","3":"22"},{"1":"638.30","2":"17.25806","3":"22"},{"1":"633.15","2":"17.37526","3":"22"},{"1":"673.30","2":"17.34931","3":"22"},{"1":"640.85","2":"16.26229","3":"22"},{"1":"684.95","2":"17.70045","3":"22"},{"1":"643.05","2":"20.12881","3":"22"},{"1":"626.10","2":"18.26539","3":"22"},{"1":"645.75","2":"14.54214","3":"22"},{"1":"664.45","2":"19.15261","3":"22"},{"1":"670.70","2":"17.36574","3":"22"},{"1":"642.40","2":"15.13898","3":"22"},{"1":"655.40","2":"17.84266","3":"22"},{"1":"623.10","2":"15.40704","3":"22"},{"1":"668.60","2":"18.86534","3":"22"},{"1":"684.30","2":"16.47413","3":"22"},{"1":"676.60","2":"17.86263","3":"22"},{"1":"645.00","2":"21.88586","3":"22"},{"1":"653.40","2":"20.20000","3":"22"},{"1":"657.00","2":"19.03640","3":"22"},{"1":"668.60","2":"17.88991","3":"23"},{"1":"645.75","2":"21.52466","3":"23"},{"1":"695.30","2":"18.69723","3":"23"},{"1":"676.55","2":"17.35714","3":"23"},{"1":"646.90","2":"18.67133","3":"23"},{"1":"666.15","2":"21.40625","3":"23"},{"1":"628.75","2":"19.50000","3":"23"},{"1":"667.20","2":"20.89412","3":"23"},{"1":"676.15","2":"19.94737","3":"23"},{"1":"681.60","2":"20.80556","3":"23"},{"1":"669.95","2":"21.23809","3":"23"},{"1":"689.10","2":"21.00000","3":"23"},{"1":"671.25","2":"20.60000","3":"23"},{"1":"623.10","2":"20.00822","3":"23"},{"1":"671.60","2":"18.02778","3":"23"},{"1":"664.15","2":"20.25196","3":"23"},{"1":"636.70","2":"16.97787","3":"23"},{"1":"661.95","2":"16.50980","3":"23"},{"1":"655.55","2":"22.70402","3":"23"},{"1":"637.35","2":"19.91111","3":"23"},{"1":"656.55","2":"18.33333","3":"23"},{"1":"647.60","2":"22.61905","3":"23"},{"1":"661.85","2":"19.44828","3":"23"},{"1":"682.15","2":"25.05263","3":"23"},{"1":"612.50","2":"20.67544","3":"23"},{"1":"696.55","2":"18.68235","3":"23"},{"1":"694.80","2":"22.84553","3":"23"},{"1":"661.35","2":"19.26667","3":"23"},{"1":"617.35","2":"19.25000","3":"23"},{"1":"673.55","2":"20.54545","3":"23"},{"1":"638.55","2":"20.60697","3":"23"},{"1":"616.45","2":"21.07268","3":"23"},{"1":"698.20","2":"21.53581","3":"23"},{"1":"673.90","2":"19.90400","3":"23"},{"1":"678.05","2":"21.19407","3":"23"},{"1":"669.85","2":"21.86535","3":"23"},{"1":"652.10","2":"18.32965","3":"23"},{"1":"625.30","2":"16.22857","3":"23"},{"1":"666.15","2":"19.17857","3":"23"},{"1":"640.10","2":"20.27737","3":"23"},{"1":"644.95","2":"22.98614","3":"23"},{"1":"672.05","2":"20.44444","3":"23"},{"1":"638.35","2":"19.82085","3":"23"},{"1":"661.60","2":"23.20522","3":"23"},{"1":"662.70","2":"19.26697","3":"23"},{"1":"661.85","2":"23.30189","3":"23"},{"1":"680.05","2":"21.18829","3":"23"},{"1":"653.70","2":"20.87180","3":"23"},{"1":"634.70","2":"19.01749","3":"23"},{"1":"669.35","2":"21.91938","3":"23"},{"1":"637.95","2":"20.10124","3":"23"},{"1":"678.80","2":"21.47651","3":"23"},{"1":"620.50","2":"20.06579","3":"23"},{"1":"664.70","2":"20.37510","3":"23"},{"1":"652.35","2":"22.44648","3":"23"},{"1":"634.15","2":"22.89524","3":"23"},{"1":"652.85","2":"20.49797","3":"23"},{"1":"670.70","2":"20.00000","3":"23"},{"1":"665.90","2":"22.25658","3":"23"},{"1":"683.35","2":"21.56436","3":"23"},{"1":"627.10","2":"19.47737","3":"23"},{"1":"666.70","2":"17.67002","3":"23"},{"1":"657.00","2":"21.94756","3":"23"},{"1":"626.90","2":"21.78339","3":"23"},{"1":"630.55","2":"19.14000","3":"23"},{"1":"640.50","2":"18.11050","3":"23"},{"1":"619.80","2":"20.68242","3":"23"},{"1":"633.65","2":"22.62361","3":"23"},{"1":"676.60","2":"21.78650","3":"23"},{"1":"657.00","2":"18.58293","3":"23"},{"1":"635.20","2":"21.54545","3":"23"},{"1":"666.65","2":"21.15289","3":"23"},{"1":"648.70","2":"16.63333","3":"23"},{"1":"660.05","2":"21.14438","3":"23"},{"1":"664.95","2":"19.78182","3":"23"},{"1":"666.55","2":"18.98373","3":"23"},{"1":"684.30","2":"17.66767","3":"23"},{"1":"664.30","2":"17.75499","3":"23"},{"1":"656.40","2":"15.27273","3":"23"},{"1":"634.40","2":"14.00000","3":"23"},{"1":"649.70","2":"20.59613","3":"23"},{"1":"666.65","2":"16.31169","3":"23"},{"1":"663.45","2":"21.12796","3":"23"},{"1":"636.10","2":"17.48801","3":"23"},{"1":"634.05","2":"17.88679","3":"23"},{"1":"644.40","2":"19.30676","3":"23"},{"1":"652.10","2":"20.89231","3":"23"},{"1":"628.65","2":"21.28684","3":"23"},{"1":"650.90","2":"20.19560","3":"23"},{"1":"665.95","2":"24.95000","3":"23"},{"1":"635.60","2":"18.13043","3":"23"},{"1":"663.85","2":"20.00000","3":"23"},{"1":"624.55","2":"18.72951","3":"23"},{"1":"642.40","2":"18.25000","3":"23"},{"1":"650.90","2":"18.99257","3":"23"},{"1":"666.10","2":"19.88764","3":"23"},{"1":"659.35","2":"19.37895","3":"23"},{"1":"616.30","2":"20.46259","3":"23"},{"1":"672.55","2":"22.29157","3":"23"},{"1":"700.30","2":"20.70474","3":"23"},{"1":"672.45","2":"19.06005","3":"23"},{"1":"684.95","2":"20.23247","3":"23"},{"1":"649.30","2":"19.69012","3":"23"},{"1":"645.10","2":"20.36254","3":"23"},{"1":"666.85","2":"19.75422","3":"23"},{"1":"657.50","2":"19.37977","3":"23"},{"1":"672.20","2":"22.92351","3":"23"},{"1":"662.90","2":"19.37340","3":"23"},{"1":"618.05","2":"19.15516","3":"23"},{"1":"627.25","2":"21.30000","3":"23"},{"1":"673.25","2":"18.30357","3":"23"},{"1":"630.55","2":"21.07926","3":"23"},{"1":"693.95","2":"18.79121","3":"23"},{"1":"638.30","2":"19.62662","3":"23"},{"1":"646.70","2":"19.59016","3":"23"},{"1":"654.85","2":"20.87187","3":"23"},{"1":"682.45","2":"21.11500","3":"23"},{"1":"695.20","2":"20.08452","3":"23"},{"1":"646.55","2":"19.91049","3":"23"},{"1":"653.55","2":"17.81285","3":"23"},{"1":"671.60","2":"18.13333","3":"23"},{"1":"669.10","2":"19.22221","3":"23"},{"1":"657.80","2":"18.66072","3":"23"},{"1":"634.55","2":"19.60000","3":"23"},{"1":"638.00","2":"19.28384","3":"23"},{"1":"676.95","2":"22.81818","3":"23"},{"1":"643.40","2":"18.80922","3":"23"},{"1":"639.75","2":"21.37363","3":"23"},{"1":"655.80","2":"20.02041","3":"23"},{"1":"650.60","2":"21.49862","3":"23"},{"1":"653.50","2":"15.42857","3":"23"},{"1":"657.15","2":"22.40000","3":"23"},{"1":"649.85","2":"20.12709","3":"23"},{"1":"679.80","2":"19.03798","3":"23"},{"1":"640.75","2":"17.34216","3":"23"},{"1":"656.50","2":"17.01863","3":"23"},{"1":"643.40","2":"20.80000","3":"23"},{"1":"634.20","2":"21.15385","3":"23"},{"1":"634.00","2":"18.45833","3":"23"},{"1":"634.95","2":"19.14082","3":"23"},{"1":"631.85","2":"19.40766","3":"23"},{"1":"666.45","2":"19.56896","3":"23"},{"1":"679.75","2":"21.50120","3":"23"},{"1":"655.35","2":"17.52941","3":"23"},{"1":"618.30","2":"16.43017","3":"23"},{"1":"662.40","2":"19.79654","3":"23"},{"1":"646.95","2":"17.18613","3":"23"},{"1":"645.60","2":"17.61589","3":"23"},{"1":"674.25","2":"20.12537","3":"23"},{"1":"660.20","2":"22.16667","3":"23"},{"1":"615.75","2":"19.96154","3":"23"},{"1":"631.05","2":"19.03945","3":"23"},{"1":"682.65","2":"15.22436","3":"23"},{"1":"621.40","2":"21.14475","3":"23"},{"1":"612.65","2":"19.64390","3":"23"},{"1":"673.30","2":"21.04869","3":"23"},{"1":"655.20","2":"20.17544","3":"23"},{"1":"651.90","2":"21.39130","3":"23"},{"1":"672.05","2":"20.00833","3":"23"},{"1":"648.35","2":"20.29137","3":"23"},{"1":"642.75","2":"17.66667","3":"23"},{"1":"631.40","2":"18.22055","3":"23"},{"1":"653.10","2":"20.27100","3":"23"},{"1":"652.50","2":"20.19895","3":"23"},{"1":"640.85","2":"21.38424","3":"23"},{"1":"659.15","2":"20.97368","3":"23"},{"1":"635.60","2":"20.00000","3":"23"},{"1":"642.20","2":"17.15328","3":"23"},{"1":"657.90","2":"22.34977","3":"23"},{"1":"632.95","2":"22.17007","3":"23"},{"1":"691.35","2":"18.18182","3":"23"},{"1":"675.40","2":"18.95714","3":"23"},{"1":"650.55","2":"19.74533","3":"23"},{"1":"667.60","2":"16.42623","3":"23"},{"1":"632.85","2":"16.62540","3":"23"},{"1":"681.90","2":"16.38177","3":"23"},{"1":"624.95","2":"20.07416","3":"23"},{"1":"671.65","2":"17.99544","3":"23"},{"1":"662.55","2":"19.39130","3":"23"},{"1":"644.45","2":"16.42857","3":"23"},{"1":"644.20","2":"16.72949","3":"23"},{"1":"643.25","2":"24.41345","3":"23"},{"1":"662.40","2":"18.26415","3":"23"},{"1":"671.90","2":"18.95504","3":"23"},{"1":"676.85","2":"21.03896","3":"23"},{"1":"671.75","2":"20.74074","3":"23"},{"1":"628.55","2":"18.10000","3":"23"},{"1":"677.25","2":"19.84615","3":"23"},{"1":"670.00","2":"21.60000","3":"23"},{"1":"645.25","2":"22.44242","3":"23"},{"1":"668.65","2":"23.01438","3":"23"},{"1":"651.20","2":"17.74892","3":"23"},{"1":"647.05","2":"18.28664","3":"23"},{"1":"706.75","2":"19.26544","3":"23"},{"1":"679.65","2":"22.66667","3":"23"},{"1":"634.10","2":"19.29412","3":"23"},{"1":"659.05","2":"17.36364","3":"23"},{"1":"672.70","2":"19.82143","3":"23"},{"1":"655.30","2":"20.43378","3":"23"},{"1":"661.90","2":"21.03721","3":"23"},{"1":"650.65","2":"19.92462","3":"23"},{"1":"644.50","2":"19.00986","3":"23"},{"1":"682.55","2":"23.82222","3":"23"},{"1":"669.30","2":"19.36909","3":"23"},{"1":"668.80","2":"19.82857","3":"23"},{"1":"637.65","2":"15.25885","3":"23"},{"1":"647.60","2":"17.16129","3":"23"},{"1":"681.30","2":"21.81333","3":"23"},{"1":"669.80","2":"19.07471","3":"23"},{"1":"635.45","2":"25.78512","3":"23"},{"1":"669.30","2":"18.21261","3":"23"},{"1":"632.25","2":"18.16606","3":"23"},{"1":"641.55","2":"16.97297","3":"23"},{"1":"656.80","2":"21.50087","3":"23"},{"1":"646.50","2":"20.60000","3":"23"},{"1":"658.80","2":"16.99029","3":"23"},{"1":"634.10","2":"20.77954","3":"23"},{"1":"687.55","2":"15.51247","3":"23"},{"1":"621.75","2":"19.88506","3":"23"},{"1":"698.25","2":"21.39882","3":"23"},{"1":"679.40","2":"20.49751","3":"23"},{"1":"652.40","2":"19.36376","3":"23"},{"1":"647.70","2":"17.65957","3":"23"},{"1":"653.95","2":"21.01796","3":"23"},{"1":"622.05","2":"19.05565","3":"23"},{"1":"648.00","2":"22.53846","3":"23"},{"1":"664.00","2":"21.10787","3":"23"},{"1":"662.55","2":"20.05135","3":"23"},{"1":"659.90","2":"14.20176","3":"23"},{"1":"663.90","2":"18.47687","3":"23"},{"1":"691.05","2":"18.63542","3":"23"},{"1":"654.20","2":"20.94595","3":"23"},{"1":"653.40","2":"21.08548","3":"23"},{"1":"644.70","2":"18.69288","3":"23"},{"1":"653.80","2":"20.86808","3":"23"},{"1":"609.00","2":"19.82558","3":"23"},{"1":"626.10","2":"19.75000","3":"23"},{"1":"655.35","2":"19.50000","3":"23"},{"1":"630.35","2":"18.39080","3":"23"},{"1":"639.50","2":"18.78676","3":"23"},{"1":"639.80","2":"19.77018","3":"23"},{"1":"660.75","2":"19.33333","3":"23"},{"1":"655.70","2":"21.46392","3":"23"},{"1":"643.60","2":"23.08492","3":"23"},{"1":"664.95","2":"21.06299","3":"23"},{"1":"624.15","2":"18.68687","3":"23"},{"1":"646.40","2":"20.77024","3":"23"},{"1":"652.30","2":"19.30556","3":"23"},{"1":"622.60","2":"20.13280","3":"23"},{"1":"665.20","2":"20.66964","3":"23"},{"1":"638.30","2":"22.28155","3":"23"},{"1":"643.50","2":"20.60027","3":"23"},{"1":"643.20","2":"20.82734","3":"23"},{"1":"639.85","2":"19.22492","3":"23"},{"1":"655.85","2":"17.65477","3":"23"},{"1":"644.45","2":"17.00000","3":"23"},{"1":"660.10","2":"16.49773","3":"23"},{"1":"686.05","2":"19.78261","3":"23"},{"1":"654.30","2":"22.30216","3":"23"},{"1":"647.30","2":"17.73077","3":"23"},{"1":"632.45","2":"20.44836","3":"23"},{"1":"633.90","2":"20.37169","3":"23"},{"1":"657.65","2":"20.16479","3":"23"},{"1":"656.70","2":"21.61538","3":"23"},{"1":"668.10","2":"20.56143","3":"23"},{"1":"665.65","2":"19.95551","3":"23"},{"1":"605.55","2":"21.18387","3":"23"},{"1":"645.75","2":"18.81042","3":"23"},{"1":"654.85","2":"20.57838","3":"23"},{"1":"630.40","2":"18.32461","3":"23"},{"1":"659.80","2":"18.82063","3":"23"},{"1":"635.95","2":"20.81633","3":"23"},{"1":"645.00","2":"20.00000","3":"23"},{"1":"665.10","2":"19.68182","3":"23"},{"1":"641.45","2":"19.39018","3":"23"},{"1":"671.95","2":"20.92732","3":"23"},{"1":"690.80","2":"19.94437","3":"23"},{"1":"616.30","2":"20.79109","3":"23"},{"1":"645.55","2":"19.20354","3":"23"},{"1":"639.25","2":"19.02439","3":"23"},{"1":"675.70","2":"17.62058","3":"23"},{"1":"661.90","2":"20.23715","3":"23"},{"1":"623.20","2":"19.29374","3":"23"},{"1":"671.90","2":"18.82998","3":"23"},{"1":"664.15","2":"20.33949","3":"23"},{"1":"661.20","2":"19.22900","3":"23"},{"1":"659.40","2":"17.89130","3":"23"},{"1":"636.50","2":"19.51881","3":"23"},{"1":"657.75","2":"19.08451","3":"23"},{"1":"655.05","2":"19.93548","3":"23"},{"1":"651.35","2":"18.87326","3":"23"},{"1":"655.75","2":"20.14178","3":"23"},{"1":"654.60","2":"23.55637","3":"23"},{"1":"653.85","2":"21.46479","3":"23"},{"1":"683.40","2":"19.19101","3":"23"},{"1":"698.45","2":"20.13080","3":"23"},{"1":"639.30","2":"25.80000","3":"23"},{"1":"704.30","2":"18.77774","3":"23"},{"1":"636.60","2":"19.10982","3":"23"},{"1":"632.00","2":"19.70109","3":"23"},{"1":"646.35","2":"18.61594","3":"23"},{"1":"694.25","2":"20.99721","3":"23"},{"1":"650.45","2":"20.00000","3":"23"},{"1":"639.90","2":"20.98325","3":"23"},{"1":"663.85","2":"21.64262","3":"23"},{"1":"631.95","2":"20.02967","3":"23"},{"1":"661.45","2":"19.81140","3":"23"},{"1":"657.40","2":"18.00000","3":"23"},{"1":"654.10","2":"19.35811","3":"23"},{"1":"652.30","2":"20.17912","3":"23"},{"1":"638.70","2":"21.11986","3":"23"},{"1":"664.75","2":"23.38974","3":"23"},{"1":"653.55","2":"22.18182","3":"23"},{"1":"652.40","2":"19.94283","3":"23"},{"1":"649.15","2":"17.78826","3":"23"},{"1":"636.90","2":"14.70588","3":"23"},{"1":"662.65","2":"19.04077","3":"23"},{"1":"648.95","2":"20.89195","3":"23"},{"1":"629.80","2":"19.83851","3":"23"},{"1":"632.20","2":"19.52191","3":"23"},{"1":"628.40","2":"20.68622","3":"23"},{"1":"666.60","2":"18.18182","3":"23"},{"1":"638.20","2":"18.89224","3":"23"},{"1":"643.50","2":"24.88889","3":"23"},{"1":"684.35","2":"18.58064","3":"23"},{"1":"672.30","2":"18.04000","3":"23"},{"1":"666.05","2":"17.73399","3":"23"},{"1":"666.00","2":"21.45455","3":"23"},{"1":"640.90","2":"19.92343","3":"23"},{"1":"654.90","2":"20.33942","3":"23"},{"1":"616.30","2":"22.54608","3":"23"},{"1":"639.35","2":"21.10344","3":"23"},{"1":"633.15","2":"18.19743","3":"23"},{"1":"641.10","2":"20.10768","3":"23"},{"1":"672.35","2":"19.15984","3":"23"},{"1":"643.05","2":"19.54545","3":"23"},{"1":"673.55","2":"20.88889","3":"23"},{"1":"645.55","2":"18.39150","3":"23"},{"1":"691.90","2":"19.17990","3":"23"},{"1":"667.15","2":"19.39771","3":"23"},{"1":"677.95","2":"21.67827","3":"23"},{"1":"637.95","2":"19.28889","3":"23"},{"1":"664.00","2":"20.34927","3":"23"},{"1":"625.85","2":"20.96416","3":"23"},{"1":"664.40","2":"19.46039","3":"23"},{"1":"664.45","2":"19.28572","3":"23"},{"1":"668.00","2":"20.91979","3":"23"},{"1":"658.60","2":"20.90021","3":"23"},{"1":"606.75","2":"20.59575","3":"23"},{"1":"656.65","2":"19.37500","3":"23"},{"1":"667.45","2":"19.95122","3":"23"},{"1":"631.90","2":"18.84973","3":"23"},{"1":"662.50","2":"18.11787","3":"23"},{"1":"646.00","2":"19.18341","3":"23"},{"1":"644.20","2":"22.00000","3":"23"},{"1":"637.10","2":"21.58416","3":"23"},{"1":"667.45","2":"20.38889","3":"23"},{"1":"647.25","2":"16.29310","3":"23"},{"1":"656.80","2":"18.27778","3":"23"},{"1":"623.60","2":"19.37472","3":"23"},{"1":"649.50","2":"18.90909","3":"23"},{"1":"641.45","2":"16.40693","3":"23"},{"1":"699.10","2":"15.59140","3":"23"},{"1":"637.00","2":"18.70694","3":"23"},{"1":"679.50","2":"18.32985","3":"23"},{"1":"655.05","2":"17.90235","3":"23"},{"1":"644.55","2":"18.91157","3":"23"},{"1":"636.95","2":"20.32497","3":"23"},{"1":"684.80","2":"20.02457","3":"23"},{"1":"626.80","2":"24.00000","3":"23"},{"1":"651.40","2":"17.60784","3":"23"},{"1":"651.85","2":"19.34853","3":"23"},{"1":"662.45","2":"19.67846","3":"23"},{"1":"678.40","2":"18.72861","3":"23"},{"1":"648.20","2":"15.88235","3":"23"},{"1":"643.70","2":"20.05491","3":"23"},{"1":"659.40","2":"17.98825","3":"23"},{"1":"668.95","2":"16.96629","3":"23"},{"1":"665.35","2":"19.23937","3":"23"},{"1":"633.05","2":"19.19586","3":"23"},{"1":"643.70","2":"19.59906","3":"23"},{"1":"673.05","2":"20.54348","3":"23"},{"1":"660.30","2":"18.58848","3":"23"},{"1":"663.50","2":"15.60419","3":"23"},{"1":"655.05","2":"15.29304","3":"23"},{"1":"628.25","2":"17.65537","3":"23"},{"1":"651.15","2":"17.57976","3":"23"},{"1":"658.35","2":"22.33333","3":"23"},{"1":"642.20","2":"18.75000","3":"23"},{"1":"651.45","2":"18.10241","3":"23"},{"1":"669.35","2":"20.25641","3":"23"},{"1":"635.05","2":"18.80207","3":"23"},{"1":"661.60","2":"18.77230","3":"23"},{"1":"634.90","2":"20.40521","3":"23"},{"1":"671.70","2":"18.65079","3":"23"},{"1":"657.55","2":"20.70707","3":"23"},{"1":"641.80","2":"22.00000","3":"23"},{"1":"671.30","2":"17.69978","3":"23"},{"1":"663.35","2":"21.48329","3":"23"},{"1":"620.30","2":"16.70103","3":"23"},{"1":"652.00","2":"19.57567","3":"23"},{"1":"666.85","2":"17.25806","3":"23"},{"1":"651.80","2":"17.37526","3":"23"},{"1":"661.85","2":"17.34931","3":"23"},{"1":"655.40","2":"16.26229","3":"23"},{"1":"668.40","2":"17.70045","3":"23"},{"1":"635.75","2":"20.12881","3":"23"},{"1":"660.95","2":"18.26539","3":"23"},{"1":"654.20","2":"14.54214","3":"23"},{"1":"640.15","2":"19.15261","3":"23"},{"1":"662.75","2":"17.36574","3":"23"},{"1":"658.00","2":"15.13898","3":"23"},{"1":"686.70","2":"17.84266","3":"23"},{"1":"623.45","2":"15.40704","3":"23"},{"1":"627.30","2":"18.86534","3":"23"},{"1":"648.25","2":"16.47413","3":"23"},{"1":"681.30","2":"17.86263","3":"23"},{"1":"680.45","2":"21.88586","3":"23"},{"1":"668.90","2":"20.20000","3":"23"},{"1":"646.20","2":"19.03640","3":"23"},{"1":"639.80","2":"17.88991","3":"24"},{"1":"663.35","2":"21.52466","3":"24"},{"1":"653.55","2":"18.69723","3":"24"},{"1":"644.45","2":"17.35714","3":"24"},{"1":"641.45","2":"18.67133","3":"24"},{"1":"628.75","2":"21.40625","3":"24"},{"1":"655.05","2":"19.50000","3":"24"},{"1":"650.90","2":"20.89412","3":"24"},{"1":"627.10","2":"19.94737","3":"24"},{"1":"661.35","2":"20.80556","3":"24"},{"1":"638.70","2":"21.23809","3":"24"},{"1":"680.45","2":"21.00000","3":"24"},{"1":"647.25","2":"20.60000","3":"24"},{"1":"645.75","2":"20.00822","3":"24"},{"1":"639.85","2":"18.02778","3":"24"},{"1":"665.35","2":"20.25196","3":"24"},{"1":"693.95","2":"16.97787","3":"24"},{"1":"616.30","2":"16.50980","3":"24"},{"1":"634.20","2":"22.70402","3":"24"},{"1":"633.05","2":"19.91111","3":"24"},{"1":"672.70","2":"18.33333","3":"24"},{"1":"656.80","2":"22.61905","3":"24"},{"1":"649.15","2":"19.44828","3":"24"},{"1":"664.45","2":"25.05263","3":"24"},{"1":"655.05","2":"20.67544","3":"24"},{"1":"656.40","2":"18.68235","3":"24"},{"1":"678.80","2":"22.84553","3":"24"},{"1":"676.60","2":"19.26667","3":"24"},{"1":"646.70","2":"19.25000","3":"24"},{"1":"661.60","2":"20.54545","3":"24"},{"1":"651.80","2":"20.60697","3":"24"},{"1":"657.75","2":"21.07268","3":"24"},{"1":"698.45","2":"21.53581","3":"24"},{"1":"649.50","2":"19.90400","3":"24"},{"1":"657.50","2":"21.19407","3":"24"},{"1":"639.35","2":"21.86535","3":"24"},{"1":"624.55","2":"18.32965","3":"24"},{"1":"666.85","2":"16.22857","3":"24"},{"1":"652.40","2":"19.17857","3":"24"},{"1":"648.20","2":"20.27737","3":"24"},{"1":"640.10","2":"22.98614","3":"24"},{"1":"664.95","2":"20.44444","3":"24"},{"1":"644.55","2":"19.82085","3":"24"},{"1":"645.55","2":"23.20522","3":"24"},{"1":"638.20","2":"19.26697","3":"24"},{"1":"654.60","2":"23.30189","3":"24"},{"1":"668.10","2":"21.18829","3":"24"},{"1":"700.30","2":"20.87180","3":"24"},{"1":"655.35","2":"19.01749","3":"24"},{"1":"695.20","2":"21.91938","3":"24"},{"1":"618.05","2":"20.10124","3":"24"},{"1":"644.20","2":"21.47651","3":"24"},{"1":"666.60","2":"20.06579","3":"24"},{"1":"636.70","2":"20.37510","3":"24"},{"1":"646.35","2":"22.44648","3":"24"},{"1":"655.20","2":"22.89524","3":"24"},{"1":"671.95","2":"20.49797","3":"24"},{"1":"670.00","2":"20.00000","3":"24"},{"1":"643.50","2":"22.25658","3":"24"},{"1":"666.00","2":"21.56436","3":"24"},{"1":"657.65","2":"19.47737","3":"24"},{"1":"656.55","2":"17.67002","3":"24"},{"1":"632.00","2":"21.94756","3":"24"},{"1":"676.95","2":"21.78339","3":"24"},{"1":"661.85","2":"19.14000","3":"24"},{"1":"664.75","2":"18.11050","3":"24"},{"1":"671.70","2":"20.68242","3":"24"},{"1":"639.90","2":"22.62361","3":"24"},{"1":"641.80","2":"21.78650","3":"24"},{"1":"649.85","2":"18.58293","3":"24"},{"1":"652.40","2":"21.54545","3":"24"},{"1":"612.65","2":"21.15289","3":"24"},{"1":"650.90","2":"16.63333","3":"24"},{"1":"662.50","2":"21.14438","3":"24"},{"1":"651.90","2":"19.78182","3":"24"},{"1":"698.20","2":"18.98373","3":"24"},{"1":"637.65","2":"17.66767","3":"24"},{"1":"681.60","2":"17.75499","3":"24"},{"1":"645.00","2":"15.27273","3":"24"},{"1":"650.65","2":"14.00000","3":"24"},{"1":"645.25","2":"20.59613","3":"24"},{"1":"652.30","2":"16.31169","3":"24"},{"1":"663.85","2":"21.12796","3":"24"},{"1":"636.50","2":"17.48801","3":"24"},{"1":"660.30","2":"17.88679","3":"24"},{"1":"669.30","2":"19.30676","3":"24"},{"1":"667.20","2":"20.89231","3":"24"},{"1":"691.90","2":"21.28684","3":"24"},{"1":"643.70","2":"20.19560","3":"24"},{"1":"681.90","2":"24.95000","3":"24"},{"1":"620.50","2":"18.13043","3":"24"},{"1":"651.20","2":"20.00000","3":"24"},{"1":"668.90","2":"18.72951","3":"24"},{"1":"665.65","2":"18.25000","3":"24"},{"1":"648.70","2":"18.99257","3":"24"},{"1":"669.30","2":"19.88764","3":"24"},{"1":"668.40","2":"19.37895","3":"24"},{"1":"640.15","2":"20.46259","3":"24"},{"1":"664.15","2":"22.29157","3":"24"},{"1":"681.30","2":"20.70474","3":"24"},{"1":"663.50","2":"19.06005","3":"24"},{"1":"662.40","2":"20.23247","3":"24"},{"1":"658.80","2":"19.69012","3":"24"},{"1":"671.60","2":"20.36254","3":"24"},{"1":"684.35","2":"19.75422","3":"24"},{"1":"704.30","2":"19.37977","3":"24"},{"1":"640.75","2":"22.92351","3":"24"},{"1":"653.95","2":"19.37340","3":"24"},{"1":"633.90","2":"19.15516","3":"24"},{"1":"671.25","2":"21.30000","3":"24"},{"1":"617.35","2":"18.30357","3":"24"},{"1":"673.55","2":"21.07926","3":"24"},{"1":"625.30","2":"18.79121","3":"24"},{"1":"630.35","2":"19.62662","3":"24"},{"1":"674.25","2":"19.59016","3":"24"},{"1":"655.75","2":"20.87187","3":"24"},{"1":"681.30","2":"21.11500","3":"24"},{"1":"643.20","2":"20.08452","3":"24"},{"1":"644.95","2":"19.91049","3":"24"},{"1":"653.50","2":"17.81285","3":"24"},{"1":"646.55","2":"18.13333","3":"24"},{"1":"662.55","2":"19.22221","3":"24"},{"1":"665.95","2":"18.66072","3":"24"},{"1":"656.65","2":"19.60000","3":"24"},{"1":"631.05","2":"19.28384","3":"24"},{"1":"652.50","2":"22.81818","3":"24"},{"1":"666.55","2":"18.80922","3":"24"},{"1":"644.45","2":"21.37363","3":"24"},{"1":"666.15","2":"20.02041","3":"24"},{"1":"644.40","2":"21.49862","3":"24"},{"1":"632.20","2":"15.42857","3":"24"},{"1":"659.35","2":"22.40000","3":"24"},{"1":"619.80","2":"20.12709","3":"24"},{"1":"634.95","2":"19.03798","3":"24"},{"1":"631.85","2":"17.34216","3":"24"},{"1":"684.80","2":"17.01863","3":"24"},{"1":"699.10","2":"20.80000","3":"24"},{"1":"645.75","2":"21.15385","3":"24"},{"1":"686.05","2":"18.45833","3":"24"},{"1":"635.95","2":"19.14082","3":"24"},{"1":"662.55","2":"19.40766","3":"24"},{"1":"671.90","2":"19.56896","3":"24"},{"1":"631.95","2":"21.50120","3":"24"},{"1":"679.65","2":"17.52941","3":"24"},{"1":"643.60","2":"16.43017","3":"24"},{"1":"672.30","2":"19.79654","3":"24"},{"1":"638.30","2":"17.18613","3":"24"},{"1":"616.45","2":"17.61589","3":"24"},{"1":"663.45","2":"20.12537","3":"24"},{"1":"665.90","2":"22.16667","3":"24"},{"1":"644.50","2":"19.96154","3":"24"},{"1":"645.10","2":"19.03945","3":"24"},{"1":"653.85","2":"15.22436","3":"24"},{"1":"662.90","2":"21.14475","3":"24"},{"1":"679.40","2":"19.64390","3":"24"},{"1":"630.40","2":"21.04869","3":"24"},{"1":"609.00","2":"20.17544","3":"24"},{"1":"651.45","2":"21.39130","3":"24"},{"1":"658.00","2":"20.00833","3":"24"},{"1":"679.50","2":"20.29137","3":"24"},{"1":"642.20","2":"17.66667","3":"24"},{"1":"672.45","2":"18.22055","3":"24"},{"1":"651.40","2":"20.27100","3":"24"},{"1":"662.75","2":"20.19895","3":"24"},{"1":"672.20","2":"21.38424","3":"24"},{"1":"673.55","2":"20.97368","3":"24"},{"1":"675.70","2":"20.00000","3":"24"},{"1":"634.10","2":"17.15328","3":"24"},{"1":"620.30","2":"22.34977","3":"24"},{"1":"643.05","2":"22.17007","3":"24"},{"1":"667.60","2":"18.18182","3":"24"},{"1":"644.70","2":"18.95714","3":"24"},{"1":"661.45","2":"19.74533","3":"24"},{"1":"658.35","2":"16.42623","3":"24"},{"1":"668.80","2":"16.62540","3":"24"},{"1":"637.95","2":"16.38177","3":"24"},{"1":"672.35","2":"20.07416","3":"24"},{"1":"660.10","2":"17.99544","3":"24"},{"1":"664.00","2":"19.39130","3":"24"},{"1":"657.55","2":"16.42857","3":"24"},{"1":"664.00","2":"16.72949","3":"24"},{"1":"623.60","2":"24.41345","3":"24"},{"1":"628.65","2":"18.26415","3":"24"},{"1":"662.45","2":"18.95504","3":"24"},{"1":"628.55","2":"21.03896","3":"24"},{"1":"679.75","2":"20.74074","3":"24"},{"1":"682.45","2":"18.10000","3":"24"},{"1":"654.20","2":"19.84615","3":"24"},{"1":"655.35","2":"21.60000","3":"24"},{"1":"652.00","2":"22.44242","3":"24"},{"1":"642.20","2":"23.01438","3":"24"},{"1":"698.25","2":"17.74892","3":"24"},{"1":"684.30","2":"18.28664","3":"24"},{"1":"679.80","2":"19.26544","3":"24"},{"1":"690.80","2":"22.66667","3":"24"},{"1":"666.05","2":"19.29412","3":"24"},{"1":"666.10","2":"17.36364","3":"24"},{"1":"660.75","2":"19.82143","3":"24"},{"1":"666.65","2":"20.43378","3":"24"},{"1":"642.75","2":"21.03721","3":"24"},{"1":"661.90","2":"19.92462","3":"24"},{"1":"647.30","2":"19.00986","3":"24"},{"1":"667.45","2":"23.82222","3":"24"},{"1":"625.85","2":"19.36909","3":"24"},{"1":"645.55","2":"19.82857","3":"24"},{"1":"668.60","2":"15.25885","3":"24"},{"1":"624.15","2":"17.16129","3":"24"},{"1":"653.10","2":"21.81333","3":"24"},{"1":"694.80","2":"19.07471","3":"24"},{"1":"655.80","2":"25.78512","3":"24"},{"1":"626.90","2":"18.21261","3":"24"},{"1":"632.45","2":"18.16606","3":"24"},{"1":"666.70","2":"16.97297","3":"24"},{"1":"657.90","2":"21.50087","3":"24"},{"1":"656.70","2":"20.60000","3":"24"},{"1":"658.60","2":"16.99029","3":"24"},{"1":"660.95","2":"20.77954","3":"24"},{"1":"643.50","2":"15.51247","3":"24"},{"1":"648.00","2":"19.88506","3":"24"},{"1":"671.60","2":"21.39882","3":"24"},{"1":"651.15","2":"20.49751","3":"24"},{"1":"682.65","2":"19.36376","3":"24"},{"1":"636.10","2":"17.65957","3":"24"},{"1":"662.40","2":"21.01796","3":"24"},{"1":"653.80","2":"19.05565","3":"24"},{"1":"657.15","2":"22.53846","3":"24"},{"1":"684.95","2":"21.10787","3":"24"},{"1":"660.20","2":"20.05135","3":"24"},{"1":"634.15","2":"14.20176","3":"24"},{"1":"677.95","2":"18.47687","3":"24"},{"1":"639.50","2":"18.63542","3":"24"},{"1":"654.10","2":"20.94595","3":"24"},{"1":"612.50","2":"21.08548","3":"24"},{"1":"687.55","2":"18.69288","3":"24"},{"1":"661.85","2":"20.86808","3":"24"},{"1":"659.90","2":"19.82558","3":"24"},{"1":"643.25","2":"19.75000","3":"24"},{"1":"633.15","2":"19.50000","3":"24"},{"1":"669.10","2":"18.39080","3":"24"},{"1":"636.90","2":"18.78676","3":"24"},{"1":"676.55","2":"19.77018","3":"24"},{"1":"630.55","2":"19.33333","3":"24"},{"1":"673.25","2":"21.46392","3":"24"},{"1":"652.10","2":"23.08492","3":"24"},{"1":"634.40","2":"21.06299","3":"24"},{"1":"637.35","2":"18.68687","3":"24"},{"1":"668.00","2":"20.77024","3":"24"},{"1":"651.85","2":"19.30556","3":"24"},{"1":"639.30","2":"20.13280","3":"24"},{"1":"669.95","2":"20.66964","3":"24"},{"1":"652.30","2":"22.28155","3":"24"},{"1":"659.40","2":"20.60027","3":"24"},{"1":"664.30","2":"20.82734","3":"24"},{"1":"654.90","2":"19.22492","3":"24"},{"1":"683.35","2":"17.65477","3":"24"},{"1":"626.10","2":"17.00000","3":"24"},{"1":"635.60","2":"16.49773","3":"24"},{"1":"672.55","2":"19.78261","3":"24"},{"1":"663.90","2":"22.30216","3":"24"},{"1":"631.90","2":"17.73077","3":"24"},{"1":"633.65","2":"20.44836","3":"24"},{"1":"652.10","2":"20.37169","3":"24"},{"1":"661.60","2":"20.16479","3":"24"},{"1":"678.05","2":"21.61538","3":"24"},{"1":"664.40","2":"20.56143","3":"24"},{"1":"646.90","2":"19.95551","3":"24"},{"1":"624.95","2":"21.18387","3":"24"},{"1":"637.95","2":"18.81042","3":"24"},{"1":"636.60","2":"20.57838","3":"24"},{"1":"683.40","2":"18.32461","3":"24"},{"1":"654.85","2":"18.82063","3":"24"},{"1":"632.85","2":"20.81633","3":"24"},{"1":"634.10","2":"20.00000","3":"24"},{"1":"659.80","2":"19.68182","3":"24"},{"1":"616.30","2":"19.39018","3":"24"},{"1":"665.20","2":"20.92732","3":"24"},{"1":"648.25","2":"19.94437","3":"24"},{"1":"646.00","2":"20.79109","3":"24"},{"1":"664.95","2":"19.20354","3":"24"},{"1":"634.70","2":"19.02439","3":"24"},{"1":"628.40","2":"17.62058","3":"24"},{"1":"638.30","2":"20.23715","3":"24"},{"1":"650.45","2":"19.29374","3":"24"},{"1":"655.85","2":"18.82998","3":"24"},{"1":"646.40","2":"20.33949","3":"24"},{"1":"671.90","2":"19.22900","3":"24"},{"1":"661.90","2":"17.89130","3":"24"},{"1":"644.20","2":"19.51881","3":"24"},{"1":"666.85","2":"19.08451","3":"24"},{"1":"646.20","2":"19.93548","3":"24"},{"1":"623.10","2":"18.87326","3":"24"},{"1":"680.05","2":"20.14178","3":"24"},{"1":"678.40","2":"23.55637","3":"24"},{"1":"634.00","2":"21.46479","3":"24"},{"1":"646.50","2":"19.19101","3":"24"},{"1":"606.75","2":"20.13080","3":"24"},{"1":"676.85","2":"25.80000","3":"24"},{"1":"647.60","2":"18.77774","3":"24"},{"1":"669.80","2":"19.10982","3":"24"},{"1":"657.00","2":"19.70109","3":"24"},{"1":"621.75","2":"18.61594","3":"24"},{"1":"634.90","2":"20.99721","3":"24"},{"1":"635.45","2":"20.00000","3":"24"},{"1":"666.45","2":"20.98325","3":"24"},{"1":"659.40","2":"21.64262","3":"24"},{"1":"668.95","2":"20.02967","3":"24"},{"1":"635.20","2":"19.81140","3":"24"},{"1":"638.00","2":"18.00000","3":"24"},{"1":"695.30","2":"19.35811","3":"24"},{"1":"643.40","2":"20.17912","3":"24"},{"1":"638.55","2":"21.11986","3":"24"},{"1":"621.40","2":"23.38974","3":"24"},{"1":"635.05","2":"22.18182","3":"24"},{"1":"649.30","2":"19.94283","3":"24"},{"1":"636.95","2":"17.78826","3":"24"},{"1":"635.75","2":"14.70588","3":"24"},{"1":"673.30","2":"19.04077","3":"24"},{"1":"691.35","2":"20.89195","3":"24"},{"1":"652.85","2":"19.83851","3":"24"},{"1":"647.70","2":"19.52191","3":"24"},{"1":"672.05","2":"20.68622","3":"24"},{"1":"654.30","2":"18.18182","3":"24"},{"1":"652.35","2":"18.89224","3":"24"},{"1":"670.70","2":"24.88889","3":"24"},{"1":"645.60","2":"18.58064","3":"24"},{"1":"691.05","2":"18.04000","3":"24"},{"1":"657.80","2":"17.73399","3":"24"},{"1":"661.20","2":"21.45455","3":"24"},{"1":"669.35","2":"19.92343","3":"24"},{"1":"664.70","2":"20.33942","3":"24"},{"1":"668.65","2":"22.54608","3":"24"},{"1":"659.05","2":"21.10344","3":"24"},{"1":"622.05","2":"18.19743","3":"24"},{"1":"663.85","2":"20.10768","3":"24"},{"1":"616.30","2":"19.15984","3":"24"},{"1":"632.25","2":"19.54545","3":"24"},{"1":"655.55","2":"20.88889","3":"24"},{"1":"673.05","2":"18.39150","3":"24"},{"1":"671.75","2":"19.17990","3":"24"},{"1":"640.50","2":"19.39771","3":"24"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] ] --- # The *infer* Pipeline: Calculate I .center[  ] --- # The *infer* Pipeline: Calculate I .left-code[ ### Specify ### Hypothesize ### Generate ### Calculate `%>% calculate(stat = "")` ] .right-plot[ .smallest[ - We `calculate` sample statistics for each of the 1,000 `replicate` samples - In our case, calculate the slope, `\((\hat{\beta_1})\)` for each `replicate` ```r CASchool %>% specify(testscr ~ str) %>% hypothesize(null = "independence") %>% generate(reps = 1000, type = "permute") %>% * calculate(stat = "slope") ``` - Other `stat`s for calculation: `"mean"`, `"median"`, `"prop"`, `"diff in means"`, `"diff in props"`, etc. (see [package information](https://infer.netlify.com/)) ] ] --- # The *infer* Pipeline: Calculate II .left-code[ ### Specify ### Hypothesize ### Generate ### Calculate `%>% calculate(stat = "")` ] .right-plot[ .smallest[ <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["replicate"],"name":[1],"type":["int"],"align":["right"]},{"label":["stat"],"name":[2],"type":["dbl"],"align":["right"]}],"data":[{"1":"1","2":"0.384783281"},{"1":"2","2":"0.241700895"},{"1":"3","2":"0.268799843"},{"1":"4","2":"-0.189039951"},{"1":"5","2":"1.215030315"},{"1":"6","2":"0.511783627"},{"1":"7","2":"-0.457378304"},{"1":"8","2":"1.008206723"},{"1":"9","2":"0.092043084"},{"1":"10","2":"0.233837801"},{"1":"11","2":"-0.445676971"},{"1":"12","2":"-0.036813328"},{"1":"13","2":"-0.043118898"},{"1":"14","2":"-0.177525696"},{"1":"15","2":"-0.406351992"},{"1":"16","2":"-0.311728867"},{"1":"17","2":"0.376747649"},{"1":"18","2":"-0.201299988"},{"1":"19","2":"0.236642852"},{"1":"20","2":"-0.742301389"},{"1":"21","2":"0.773978816"},{"1":"22","2":"0.283806177"},{"1":"23","2":"-0.446008426"},{"1":"24","2":"-0.301509287"},{"1":"25","2":"-0.250904829"},{"1":"26","2":"0.155393905"},{"1":"27","2":"-0.649649642"},{"1":"28","2":"0.277685932"},{"1":"29","2":"0.450507687"},{"1":"30","2":"0.161334900"},{"1":"31","2":"0.093849947"},{"1":"32","2":"-0.427373560"},{"1":"33","2":"0.650190354"},{"1":"34","2":"0.544727649"},{"1":"35","2":"-0.155540584"},{"1":"36","2":"0.101344971"},{"1":"37","2":"0.422808352"},{"1":"38","2":"-0.183895207"},{"1":"39","2":"0.341437535"},{"1":"40","2":"-0.623312627"},{"1":"41","2":"0.251732645"},{"1":"42","2":"0.181194538"},{"1":"43","2":"-0.124123737"},{"1":"44","2":"0.967033768"},{"1":"45","2":"-0.190687589"},{"1":"46","2":"0.134063369"},{"1":"47","2":"-0.217879354"},{"1":"48","2":"-0.004629455"},{"1":"49","2":"0.193039075"},{"1":"50","2":"-0.168591669"},{"1":"51","2":"-0.103141833"},{"1":"52","2":"0.124380374"},{"1":"53","2":"0.221131052"},{"1":"54","2":"0.117136592"},{"1":"55","2":"0.892638832"},{"1":"56","2":"-0.040416949"},{"1":"57","2":"0.819786957"},{"1":"58","2":"0.205001272"},{"1":"59","2":"0.151970308"},{"1":"60","2":"0.134828268"},{"1":"61","2":"-0.321222382"},{"1":"62","2":"-0.613999902"},{"1":"63","2":"-0.008797143"},{"1":"64","2":"0.754121948"},{"1":"65","2":"-0.093272902"},{"1":"66","2":"0.160970525"},{"1":"67","2":"0.018874146"},{"1":"68","2":"0.030703061"},{"1":"69","2":"0.872196248"},{"1":"70","2":"-0.271602375"},{"1":"71","2":"0.146690136"},{"1":"72","2":"-0.103976880"},{"1":"73","2":"0.327961214"},{"1":"74","2":"-0.276573383"},{"1":"75","2":"-0.709912058"},{"1":"76","2":"-0.612175461"},{"1":"77","2":"-0.345773372"},{"1":"78","2":"0.009255478"},{"1":"79","2":"-0.415895902"},{"1":"80","2":"0.144022046"},{"1":"81","2":"0.357361106"},{"1":"82","2":"-0.182868419"},{"1":"83","2":"0.137138091"},{"1":"84","2":"0.588399050"},{"1":"85","2":"-0.794232157"},{"1":"86","2":"-0.461471728"},{"1":"87","2":"-0.178585772"},{"1":"88","2":"0.247112387"},{"1":"89","2":"0.107803822"},{"1":"90","2":"-0.266329362"},{"1":"91","2":"0.084674046"},{"1":"92","2":"0.539791560"},{"1":"93","2":"-0.127008427"},{"1":"94","2":"-0.566030015"},{"1":"95","2":"-0.050963910"},{"1":"96","2":"0.985269956"},{"1":"97","2":"0.032733209"},{"1":"98","2":"-0.493801785"},{"1":"99","2":"-0.564488516"},{"1":"100","2":"0.605097549"},{"1":"101","2":"1.158663277"},{"1":"102","2":"0.111880621"},{"1":"103","2":"-1.013596175"},{"1":"104","2":"0.055783471"},{"1":"105","2":"0.085244755"},{"1":"106","2":"-0.689072722"},{"1":"107","2":"0.116025236"},{"1":"108","2":"0.374462000"},{"1":"109","2":"0.555295292"},{"1":"110","2":"-0.350166440"},{"1":"111","2":"-0.057428613"},{"1":"112","2":"0.215446568"},{"1":"113","2":"0.127589236"},{"1":"114","2":"0.409151797"},{"1":"115","2":"-0.106791445"},{"1":"116","2":"0.738261903"},{"1":"117","2":"0.274467112"},{"1":"118","2":"0.058143261"},{"1":"119","2":"0.165162335"},{"1":"120","2":"-0.442223744"},{"1":"121","2":"-0.296436738"},{"1":"122","2":"-0.465844816"},{"1":"123","2":"0.940190628"},{"1":"124","2":"-0.268824485"},{"1":"125","2":"-0.341673562"},{"1":"126","2":"0.383824805"},{"1":"127","2":"0.022294381"},{"1":"128","2":"0.723093117"},{"1":"129","2":"-0.515045247"},{"1":"130","2":"-0.811833469"},{"1":"131","2":"0.361797399"},{"1":"132","2":"0.742750495"},{"1":"133","2":"-0.974416696"},{"1":"134","2":"-1.005882693"},{"1":"135","2":"-0.049904851"},{"1":"136","2":"0.718178393"},{"1":"137","2":"0.353763483"},{"1":"138","2":"-1.093791135"},{"1":"139","2":"0.177423371"},{"1":"140","2":"-0.516159873"},{"1":"141","2":"-0.500447550"},{"1":"142","2":"-0.363266393"},{"1":"143","2":"0.771094692"},{"1":"144","2":"-0.267763553"},{"1":"145","2":"0.467830784"},{"1":"146","2":"0.215496640"},{"1":"147","2":"0.932455129"},{"1":"148","2":"-0.580508451"},{"1":"149","2":"-1.542863084"},{"1":"150","2":"-1.138057707"},{"1":"151","2":"1.515074005"},{"1":"152","2":"0.827998212"},{"1":"153","2":"-0.946618798"},{"1":"154","2":"1.434033286"},{"1":"155","2":"-0.231243568"},{"1":"156","2":"0.310256878"},{"1":"157","2":"-0.060357036"},{"1":"158","2":"0.047984462"},{"1":"159","2":"0.170803807"},{"1":"160","2":"-0.346651465"},{"1":"161","2":"0.074042942"},{"1":"162","2":"0.292340737"},{"1":"163","2":"-0.013259074"},{"1":"164","2":"1.092762081"},{"1":"165","2":"-0.487574644"},{"1":"166","2":"-0.189834536"},{"1":"167","2":"0.647436122"},{"1":"168","2":"-0.381581015"},{"1":"169","2":"-0.135456386"},{"1":"170","2":"0.367657290"},{"1":"171","2":"-0.114425532"},{"1":"172","2":"0.114871690"},{"1":"173","2":"0.087953109"},{"1":"174","2":"-0.172484057"},{"1":"175","2":"0.244612878"},{"1":"176","2":"0.513783318"},{"1":"177","2":"-0.541663976"},{"1":"178","2":"-1.032394179"},{"1":"179","2":"-0.506948372"},{"1":"180","2":"0.375834776"},{"1":"181","2":"-0.146353305"},{"1":"182","2":"-0.523263639"},{"1":"183","2":"-0.437623485"},{"1":"184","2":"0.912232333"},{"1":"185","2":"-0.589911526"},{"1":"186","2":"0.190165615"},{"1":"187","2":"0.077886078"},{"1":"188","2":"-0.271806208"},{"1":"189","2":"0.010233108"},{"1":"190","2":"0.418141113"},{"1":"191","2":"-0.325992479"},{"1":"192","2":"0.709060722"},{"1":"193","2":"0.534623918"},{"1":"194","2":"0.078128109"},{"1":"195","2":"-0.637697736"},{"1":"196","2":"-0.701659931"},{"1":"197","2":"0.323236916"},{"1":"198","2":"-0.049196055"},{"1":"199","2":"0.500416363"},{"1":"200","2":"0.451814231"},{"1":"201","2":"-0.466313473"},{"1":"202","2":"-0.117749517"},{"1":"203","2":"0.455481076"},{"1":"204","2":"-0.466297636"},{"1":"205","2":"0.843522171"},{"1":"206","2":"0.145318409"},{"1":"207","2":"-0.127785904"},{"1":"208","2":"0.769661818"},{"1":"209","2":"-0.139451996"},{"1":"210","2":"-0.345445767"},{"1":"211","2":"-0.079822069"},{"1":"212","2":"-0.266155972"},{"1":"213","2":"0.321488736"},{"1":"214","2":"-0.140491604"},{"1":"215","2":"0.162167817"},{"1":"216","2":"0.454341288"},{"1":"217","2":"1.345292985"},{"1":"218","2":"-0.047458319"},{"1":"219","2":"-0.824461139"},{"1":"220","2":"0.184351540"},{"1":"221","2":"-0.441437874"},{"1":"222","2":"-0.389260488"},{"1":"223","2":"0.839969387"},{"1":"224","2":"-0.553853562"},{"1":"225","2":"-0.495956211"},{"1":"226","2":"-0.504665618"},{"1":"227","2":"0.400194567"},{"1":"228","2":"-0.174515554"},{"1":"229","2":"0.257287450"},{"1":"230","2":"0.531195131"},{"1":"231","2":"0.326883855"},{"1":"232","2":"-0.092122095"},{"1":"233","2":"0.064613365"},{"1":"234","2":"0.282009046"},{"1":"235","2":"-0.419463259"},{"1":"236","2":"0.284828573"},{"1":"237","2":"0.140074899"},{"1":"238","2":"0.448234036"},{"1":"239","2":"-0.154727886"},{"1":"240","2":"0.451633731"},{"1":"241","2":"0.280397223"},{"1":"242","2":"-0.390214568"},{"1":"243","2":"-0.939459542"},{"1":"244","2":"0.548623974"},{"1":"245","2":"-0.640197137"},{"1":"246","2":"1.326982348"},{"1":"247","2":"0.525704986"},{"1":"248","2":"0.696793162"},{"1":"249","2":"0.858641917"},{"1":"250","2":"0.013868941"},{"1":"251","2":"-0.958033028"},{"1":"252","2":"-0.738986072"},{"1":"253","2":"-1.029774494"},{"1":"254","2":"-0.012679553"},{"1":"255","2":"0.741874585"},{"1":"256","2":"-0.691893360"},{"1":"257","2":"-0.538200303"},{"1":"258","2":"-0.094533103"},{"1":"259","2":"-0.387957009"},{"1":"260","2":"0.150446273"},{"1":"261","2":"-0.178290332"},{"1":"262","2":"-0.068633705"},{"1":"263","2":"-0.304457124"},{"1":"264","2":"-0.450191399"},{"1":"265","2":"-0.518622480"},{"1":"266","2":"0.660751972"},{"1":"267","2":"0.614295507"},{"1":"268","2":"0.006756848"},{"1":"269","2":"0.070004742"},{"1":"270","2":"-0.543069604"},{"1":"271","2":"0.181761428"},{"1":"272","2":"0.037976260"},{"1":"273","2":"-0.670125486"},{"1":"274","2":"0.179539930"},{"1":"275","2":"-0.108014853"},{"1":"276","2":"0.462090110"},{"1":"277","2":"-1.394443787"},{"1":"278","2":"0.154330259"},{"1":"279","2":"-0.285384337"},{"1":"280","2":"0.084168355"},{"1":"281","2":"-0.299996827"},{"1":"282","2":"-0.580578487"},{"1":"283","2":"0.165750272"},{"1":"284","2":"-0.106883549"},{"1":"285","2":"0.421248908"},{"1":"286","2":"-0.617698229"},{"1":"287","2":"-0.069431893"},{"1":"288","2":"-0.322109913"},{"1":"289","2":"-0.321276682"},{"1":"290","2":"-0.906676895"},{"1":"291","2":"0.643863072"},{"1":"292","2":"-0.216617106"},{"1":"293","2":"0.765933770"},{"1":"294","2":"-0.404250909"},{"1":"295","2":"0.530721203"},{"1":"296","2":"-0.809997458"},{"1":"297","2":"0.146559983"},{"1":"298","2":"-0.213540649"},{"1":"299","2":"-0.722830086"},{"1":"300","2":"-0.242878453"},{"1":"301","2":"-0.656718170"},{"1":"302","2":"-0.609696404"},{"1":"303","2":"-0.687729683"},{"1":"304","2":"-0.522236174"},{"1":"305","2":"0.397509345"},{"1":"306","2":"-0.039664721"},{"1":"307","2":"0.096483262"},{"1":"308","2":"0.628294995"},{"1":"309","2":"0.032773066"},{"1":"310","2":"0.267288133"},{"1":"311","2":"1.113652150"},{"1":"312","2":"0.709246813"},{"1":"313","2":"-0.442665408"},{"1":"314","2":"0.743312158"},{"1":"315","2":"0.395061150"},{"1":"316","2":"-0.016789103"},{"1":"317","2":"-0.368734683"},{"1":"318","2":"-0.503695487"},{"1":"319","2":"0.780282684"},{"1":"320","2":"0.041608833"},{"1":"321","2":"-0.439993905"},{"1":"322","2":"-0.669808317"},{"1":"323","2":"-0.783970957"},{"1":"324","2":"-0.137154895"},{"1":"325","2":"-0.215248116"},{"1":"326","2":"-0.326008171"},{"1":"327","2":"0.256482981"},{"1":"328","2":"-0.838986015"},{"1":"329","2":"-0.171062097"},{"1":"330","2":"0.389152684"},{"1":"331","2":"0.551004334"},{"1":"332","2":"0.178554217"},{"1":"333","2":"-0.654779294"},{"1":"334","2":"0.456037671"},{"1":"335","2":"0.133692707"},{"1":"336","2":"0.357032259"},{"1":"337","2":"0.254848570"},{"1":"338","2":"0.128421916"},{"1":"339","2":"0.297332422"},{"1":"340","2":"-0.609553451"},{"1":"341","2":"0.477694702"},{"1":"342","2":"0.531496759"},{"1":"343","2":"-0.159482874"},{"1":"344","2":"-0.532020593"},{"1":"345","2":"1.292737270"},{"1":"346","2":"-0.927207642"},{"1":"347","2":"-1.345167877"},{"1":"348","2":"-0.355834455"},{"1":"349","2":"0.513911840"},{"1":"350","2":"-0.648253721"},{"1":"351","2":"-0.105561625"},{"1":"352","2":"-0.263280518"},{"1":"353","2":"-0.466511828"},{"1":"354","2":"-0.177470625"},{"1":"355","2":"-0.111238516"},{"1":"356","2":"-0.095027444"},{"1":"357","2":"-0.720628081"},{"1":"358","2":"-0.708581213"},{"1":"359","2":"-0.053765527"},{"1":"360","2":"0.768134437"},{"1":"361","2":"-0.348808655"},{"1":"362","2":"0.406713925"},{"1":"363","2":"-0.476088160"},{"1":"364","2":"-0.247155470"},{"1":"365","2":"-0.207932895"},{"1":"366","2":"0.557667344"},{"1":"367","2":"-0.963266258"},{"1":"368","2":"-0.411052515"},{"1":"369","2":"-0.834509407"},{"1":"370","2":"0.109209559"},{"1":"371","2":"-0.257853210"},{"1":"372","2":"-1.044283522"},{"1":"373","2":"0.080542166"},{"1":"374","2":"0.358106665"},{"1":"375","2":"-0.234374140"},{"1":"376","2":"0.612093045"},{"1":"377","2":"0.555558793"},{"1":"378","2":"0.163709686"},{"1":"379","2":"1.367249373"},{"1":"380","2":"-0.374362906"},{"1":"381","2":"0.005782268"},{"1":"382","2":"-0.323735885"},{"1":"383","2":"-0.287131921"},{"1":"384","2":"0.566027985"},{"1":"385","2":"-0.861746597"},{"1":"386","2":"0.953560735"},{"1":"387","2":"1.107853955"},{"1":"388","2":"0.695068731"},{"1":"389","2":"0.168042952"},{"1":"390","2":"0.335300213"},{"1":"391","2":"0.080872197"},{"1":"392","2":"0.164365328"},{"1":"393","2":"0.373794430"},{"1":"394","2":"-0.327966505"},{"1":"395","2":"-0.140736543"},{"1":"396","2":"0.302787781"},{"1":"397","2":"0.863149362"},{"1":"398","2":"-0.038769296"},{"1":"399","2":"0.661849685"},{"1":"400","2":"0.034183931"},{"1":"401","2":"0.212317035"},{"1":"402","2":"-0.033855405"},{"1":"403","2":"0.250894845"},{"1":"404","2":"-0.280594816"},{"1":"405","2":"-0.092689028"},{"1":"406","2":"0.386305198"},{"1":"407","2":"-0.863742790"},{"1":"408","2":"-1.024612150"},{"1":"409","2":"0.940570725"},{"1":"410","2":"0.891284682"},{"1":"411","2":"-0.274163415"},{"1":"412","2":"-0.956715970"},{"1":"413","2":"-0.154214172"},{"1":"414","2":"0.442316057"},{"1":"415","2":"0.686017459"},{"1":"416","2":"0.744311549"},{"1":"417","2":"-0.057776987"},{"1":"418","2":"-0.726031949"},{"1":"419","2":"-0.549983754"},{"1":"420","2":"0.657189643"},{"1":"421","2":"-0.419228393"},{"1":"422","2":"0.246357726"},{"1":"423","2":"-0.048257104"},{"1":"424","2":"-0.294865276"},{"1":"425","2":"0.722857537"},{"1":"426","2":"0.878027307"},{"1":"427","2":"-0.081443206"},{"1":"428","2":"0.525630683"},{"1":"429","2":"-0.187010808"},{"1":"430","2":"0.600955052"},{"1":"431","2":"0.226404296"},{"1":"432","2":"0.467826156"},{"1":"433","2":"-0.259687428"},{"1":"434","2":"0.022420433"},{"1":"435","2":"-0.656136339"},{"1":"436","2":"-0.214424339"},{"1":"437","2":"-0.146469363"},{"1":"438","2":"0.020299830"},{"1":"439","2":"0.242355971"},{"1":"440","2":"0.666680098"},{"1":"441","2":"0.003656225"},{"1":"442","2":"-0.502710628"},{"1":"443","2":"0.265717339"},{"1":"444","2":"0.200212561"},{"1":"445","2":"-0.451076301"},{"1":"446","2":"0.107194343"},{"1":"447","2":"0.559516228"},{"1":"448","2":"1.297326645"},{"1":"449","2":"0.371931683"},{"1":"450","2":"-1.420791113"},{"1":"451","2":"-0.619012281"},{"1":"452","2":"-0.140396889"},{"1":"453","2":"-0.444358025"},{"1":"454","2":"0.205260787"},{"1":"455","2":"-0.495145441"},{"1":"456","2":"0.161150715"},{"1":"457","2":"0.444352045"},{"1":"458","2":"-0.218760457"},{"1":"459","2":"-0.543016368"},{"1":"460","2":"0.346774807"},{"1":"461","2":"-0.307538789"},{"1":"462","2":"-0.129555235"},{"1":"463","2":"0.319658527"},{"1":"464","2":"-0.633282247"},{"1":"465","2":"0.358656602"},{"1":"466","2":"0.362487431"},{"1":"467","2":"-0.866341882"},{"1":"468","2":"0.214356949"},{"1":"469","2":"-0.405726231"},{"1":"470","2":"-0.644303697"},{"1":"471","2":"0.386367492"},{"1":"472","2":"0.033513429"},{"1":"473","2":"-0.688195177"},{"1":"474","2":"0.302358173"},{"1":"475","2":"0.272413463"},{"1":"476","2":"-0.529602267"},{"1":"477","2":"0.733891062"},{"1":"478","2":"-0.294163691"},{"1":"479","2":"0.150363793"},{"1":"480","2":"-0.837140581"},{"1":"481","2":"-0.238761485"},{"1":"482","2":"-0.589805161"},{"1":"483","2":"-0.810163378"},{"1":"484","2":"-0.619846624"},{"1":"485","2":"-0.217378163"},{"1":"486","2":"-0.320612243"},{"1":"487","2":"-0.273771081"},{"1":"488","2":"0.825098052"},{"1":"489","2":"0.345310068"},{"1":"490","2":"0.873734869"},{"1":"491","2":"0.103017297"},{"1":"492","2":"-0.260816377"},{"1":"493","2":"-0.299346754"},{"1":"494","2":"0.302302208"},{"1":"495","2":"-1.086764537"},{"1":"496","2":"-0.305511293"},{"1":"497","2":"-0.282852667"},{"1":"498","2":"-0.934307695"},{"1":"499","2":"0.631162672"},{"1":"500","2":"-0.074206878"},{"1":"501","2":"0.195768995"},{"1":"502","2":"-0.421480500"},{"1":"503","2":"-0.136892941"},{"1":"504","2":"-1.009331386"},{"1":"505","2":"0.399526007"},{"1":"506","2":"0.754767003"},{"1":"507","2":"0.487931315"},{"1":"508","2":"0.396142960"},{"1":"509","2":"0.599957247"},{"1":"510","2":"0.126644972"},{"1":"511","2":"-0.202882955"},{"1":"512","2":"-0.369928459"},{"1":"513","2":"0.753018852"},{"1":"514","2":"1.067920756"},{"1":"515","2":"-0.322992692"},{"1":"516","2":"1.021169180"},{"1":"517","2":"-0.313989560"},{"1":"518","2":"-0.274286629"},{"1":"519","2":"-0.953021674"},{"1":"520","2":"-0.140997290"},{"1":"521","2":"0.333062506"},{"1":"522","2":"0.151699637"},{"1":"523","2":"0.458256115"},{"1":"524","2":"0.744691076"},{"1":"525","2":"-0.586834310"},{"1":"526","2":"-1.097598845"},{"1":"527","2":"-0.769167937"},{"1":"528","2":"-0.509747841"},{"1":"529","2":"0.004045690"},{"1":"530","2":"0.433581241"},{"1":"531","2":"0.405411820"},{"1":"532","2":"-0.338023782"},{"1":"533","2":"0.130707881"},{"1":"534","2":"-0.004370268"},{"1":"535","2":"-0.112289742"},{"1":"536","2":"-0.507562392"},{"1":"537","2":"0.072765615"},{"1":"538","2":"-0.347437441"},{"1":"539","2":"-0.419343388"},{"1":"540","2":"-0.322308101"},{"1":"541","2":"-0.170418705"},{"1":"542","2":"-0.328832639"},{"1":"543","2":"0.071284287"},{"1":"544","2":"0.650544304"},{"1":"545","2":"-0.117987192"},{"1":"546","2":"1.179244992"},{"1":"547","2":"0.081611742"},{"1":"548","2":"-0.387813462"},{"1":"549","2":"-0.012385994"},{"1":"550","2":"-0.462926047"},{"1":"551","2":"0.349155458"},{"1":"552","2":"0.512204689"},{"1":"553","2":"-0.034607938"},{"1":"554","2":"-0.979060560"},{"1":"555","2":"-0.312203226"},{"1":"556","2":"-0.233687894"},{"1":"557","2":"-0.044753791"},{"1":"558","2":"0.768757577"},{"1":"559","2":"0.672804711"},{"1":"560","2":"0.342409201"},{"1":"561","2":"-0.082578029"},{"1":"562","2":"0.060763880"},{"1":"563","2":"0.361461452"},{"1":"564","2":"0.008025913"},{"1":"565","2":"-0.278122501"},{"1":"566","2":"0.862956423"},{"1":"567","2":"-0.033773855"},{"1":"568","2":"-0.120734869"},{"1":"569","2":"0.227769528"},{"1":"570","2":"0.045893456"},{"1":"571","2":"-0.147372190"},{"1":"572","2":"0.329491933"},{"1":"573","2":"-0.055884817"},{"1":"574","2":"-0.787415708"},{"1":"575","2":"0.398953336"},{"1":"576","2":"-0.709699428"},{"1":"577","2":"-0.315014143"},{"1":"578","2":"0.126468815"},{"1":"579","2":"-0.022193885"},{"1":"580","2":"0.504389006"},{"1":"581","2":"0.536033134"},{"1":"582","2":"-0.466639088"},{"1":"583","2":"-0.299861151"},{"1":"584","2":"-0.834101283"},{"1":"585","2":"0.706200704"},{"1":"586","2":"0.452427016"},{"1":"587","2":"0.667507978"},{"1":"588","2":"-0.229383506"},{"1":"589","2":"0.116028201"},{"1":"590","2":"-0.070312370"},{"1":"591","2":"0.393783245"},{"1":"592","2":"0.026009698"},{"1":"593","2":"0.156005272"},{"1":"594","2":"0.319576653"},{"1":"595","2":"-0.061463999"},{"1":"596","2":"0.911079234"},{"1":"597","2":"0.318043384"},{"1":"598","2":"0.402852314"},{"1":"599","2":"-0.397657391"},{"1":"600","2":"-0.150325340"},{"1":"601","2":"-0.440204726"},{"1":"602","2":"0.491778967"},{"1":"603","2":"0.418827492"},{"1":"604","2":"-0.807693330"},{"1":"605","2":"0.303109652"},{"1":"606","2":"0.437337111"},{"1":"607","2":"-0.174324414"},{"1":"608","2":"0.097828501"},{"1":"609","2":"-0.676622908"},{"1":"610","2":"0.152309802"},{"1":"611","2":"-0.311067483"},{"1":"612","2":"-0.213655381"},{"1":"613","2":"-0.026828099"},{"1":"614","2":"-0.406615192"},{"1":"615","2":"0.850914110"},{"1":"616","2":"-0.052961230"},{"1":"617","2":"0.425246139"},{"1":"618","2":"-1.068085798"},{"1":"619","2":"-0.186469878"},{"1":"620","2":"-0.613539676"},{"1":"621","2":"0.472193236"},{"1":"622","2":"0.839267527"},{"1":"623","2":"-0.951613332"},{"1":"624","2":"0.996436330"},{"1":"625","2":"0.078191175"},{"1":"626","2":"0.550498660"},{"1":"627","2":"-0.192557582"},{"1":"628","2":"-0.165671874"},{"1":"629","2":"-0.268626815"},{"1":"630","2":"-0.596783949"},{"1":"631","2":"0.491446604"},{"1":"632","2":"-0.593543184"},{"1":"633","2":"-1.005063108"},{"1":"634","2":"-0.657378941"},{"1":"635","2":"0.037083686"},{"1":"636","2":"0.136445539"},{"1":"637","2":"-0.417857809"},{"1":"638","2":"0.207994791"},{"1":"639","2":"-0.474869969"},{"1":"640","2":"-1.177796979"},{"1":"641","2":"-0.585657936"},{"1":"642","2":"-1.000782845"},{"1":"643","2":"0.102189349"},{"1":"644","2":"0.337059290"},{"1":"645","2":"-0.087969355"},{"1":"646","2":"-0.573166493"},{"1":"647","2":"0.389817706"},{"1":"648","2":"0.781200726"},{"1":"649","2":"-0.132578435"},{"1":"650","2":"0.418229239"},{"1":"651","2":"0.328300902"},{"1":"652","2":"-0.017070103"},{"1":"653","2":"0.930114143"},{"1":"654","2":"1.290330474"},{"1":"655","2":"1.128467030"},{"1":"656","2":"-0.774667056"},{"1":"657","2":"0.458408656"},{"1":"658","2":"-0.444991415"},{"1":"659","2":"0.142835689"},{"1":"660","2":"-0.158697472"},{"1":"661","2":"-0.871227948"},{"1":"662","2":"-0.076296461"},{"1":"663","2":"0.123900430"},{"1":"664","2":"0.010820338"},{"1":"665","2":"0.426133938"},{"1":"666","2":"-0.415784405"},{"1":"667","2":"-0.060669814"},{"1":"668","2":"0.011292388"},{"1":"669","2":"-0.419750142"},{"1":"670","2":"0.379208553"},{"1":"671","2":"-0.443830564"},{"1":"672","2":"0.614854426"},{"1":"673","2":"-0.659886632"},{"1":"674","2":"-0.758181777"},{"1":"675","2":"-0.755836944"},{"1":"676","2":"-1.226742261"},{"1":"677","2":"-0.182653356"},{"1":"678","2":"-0.568889686"},{"1":"679","2":"-0.938811554"},{"1":"680","2":"-0.933366153"},{"1":"681","2":"-0.398588431"},{"1":"682","2":"0.383146087"},{"1":"683","2":"-1.381153604"},{"1":"684","2":"0.538949884"},{"1":"685","2":"-0.774088539"},{"1":"686","2":"-0.570857172"},{"1":"687","2":"-0.374231458"},{"1":"688","2":"-0.199611870"},{"1":"689","2":"0.263824586"},{"1":"690","2":"-0.393575265"},{"1":"691","2":"0.050447483"},{"1":"692","2":"0.539710143"},{"1":"693","2":"-0.337531119"},{"1":"694","2":"-0.135724207"},{"1":"695","2":"1.098073331"},{"1":"696","2":"0.348330529"},{"1":"697","2":"0.729599978"},{"1":"698","2":"-0.606679158"},{"1":"699","2":"-0.456479514"},{"1":"700","2":"-0.466648304"},{"1":"701","2":"0.082042171"},{"1":"702","2":"-0.520363618"},{"1":"703","2":"0.373744681"},{"1":"704","2":"-0.149980989"},{"1":"705","2":"-0.306022256"},{"1":"706","2":"-0.607344387"},{"1":"707","2":"-0.793082097"},{"1":"708","2":"-0.027971565"},{"1":"709","2":"-0.105164172"},{"1":"710","2":"1.461637118"},{"1":"711","2":"-0.484662876"},{"1":"712","2":"-0.645819894"},{"1":"713","2":"-0.730086927"},{"1":"714","2":"0.150198145"},{"1":"715","2":"0.069811309"},{"1":"716","2":"0.034853357"},{"1":"717","2":"0.140667255"},{"1":"718","2":"-0.470300547"},{"1":"719","2":"0.809065958"},{"1":"720","2":"-0.586613987"},{"1":"721","2":"0.122593898"},{"1":"722","2":"-0.013259455"},{"1":"723","2":"-0.065876871"},{"1":"724","2":"0.166245013"},{"1":"725","2":"0.702434868"},{"1":"726","2":"0.364887529"},{"1":"727","2":"-0.366288183"},{"1":"728","2":"0.376309429"},{"1":"729","2":"0.583515088"},{"1":"730","2":"0.046589530"},{"1":"731","2":"0.869632541"},{"1":"732","2":"0.199692344"},{"1":"733","2":"0.616495357"},{"1":"734","2":"0.181932081"},{"1":"735","2":"-0.112843336"},{"1":"736","2":"0.991113144"},{"1":"737","2":"-0.041540017"},{"1":"738","2":"-0.251468922"},{"1":"739","2":"0.065506681"},{"1":"740","2":"-0.473748022"},{"1":"741","2":"0.702127043"},{"1":"742","2":"-0.538663966"},{"1":"743","2":"0.269264095"},{"1":"744","2":"0.789738536"},{"1":"745","2":"0.361289065"},{"1":"746","2":"0.384796910"},{"1":"747","2":"0.143489368"},{"1":"748","2":"0.619538618"},{"1":"749","2":"0.296182971"},{"1":"750","2":"0.069027012"},{"1":"751","2":"0.691891746"},{"1":"752","2":"-0.139516893"},{"1":"753","2":"0.334803522"},{"1":"754","2":"0.173413799"},{"1":"755","2":"0.585025755"},{"1":"756","2":"0.312183439"},{"1":"757","2":"0.647655930"},{"1":"758","2":"0.508753857"},{"1":"759","2":"-0.584338567"},{"1":"760","2":"0.764005001"},{"1":"761","2":"-0.120799430"},{"1":"762","2":"-0.120410832"},{"1":"763","2":"-0.352248618"},{"1":"764","2":"0.289429505"},{"1":"765","2":"-0.297584170"},{"1":"766","2":"-0.777011079"},{"1":"767","2":"-0.032617173"},{"1":"768","2":"-0.424977296"},{"1":"769","2":"-0.506216649"},{"1":"770","2":"0.151951758"},{"1":"771","2":"-1.097682811"},{"1":"772","2":"-0.502005295"},{"1":"773","2":"0.408373748"},{"1":"774","2":"0.174283169"},{"1":"775","2":"-0.357446385"},{"1":"776","2":"-0.279777843"},{"1":"777","2":"-0.478527562"},{"1":"778","2":"-0.729158898"},{"1":"779","2":"0.637112058"},{"1":"780","2":"-0.112293413"},{"1":"781","2":"-0.070220860"},{"1":"782","2":"0.407416211"},{"1":"783","2":"-0.002881228"},{"1":"784","2":"-0.056183899"},{"1":"785","2":"-0.312043791"},{"1":"786","2":"-0.228790992"},{"1":"787","2":"-0.165929988"},{"1":"788","2":"-0.353336846"},{"1":"789","2":"0.304699204"},{"1":"790","2":"-0.362306016"},{"1":"791","2":"-0.051849060"},{"1":"792","2":"-0.047957252"},{"1":"793","2":"0.490019006"},{"1":"794","2":"0.781568586"},{"1":"795","2":"-0.408135192"},{"1":"796","2":"1.458086863"},{"1":"797","2":"0.841364928"},{"1":"798","2":"0.292415118"},{"1":"799","2":"-0.619146032"},{"1":"800","2":"0.642447077"},{"1":"801","2":"0.101299795"},{"1":"802","2":"-1.256690870"},{"1":"803","2":"-0.332366932"},{"1":"804","2":"-0.188850980"},{"1":"805","2":"-0.737220033"},{"1":"806","2":"-0.144158131"},{"1":"807","2":"-0.008095132"},{"1":"808","2":"-0.063249991"},{"1":"809","2":"0.808895908"},{"1":"810","2":"-0.048566442"},{"1":"811","2":"-0.158964599"},{"1":"812","2":"-0.520431400"},{"1":"813","2":"0.481771045"},{"1":"814","2":"0.346026910"},{"1":"815","2":"-0.076063600"},{"1":"816","2":"-0.101980512"},{"1":"817","2":"0.078577813"},{"1":"818","2":"-0.149961473"},{"1":"819","2":"0.421958287"},{"1":"820","2":"-0.547580608"},{"1":"821","2":"-0.018883204"},{"1":"822","2":"1.232858475"},{"1":"823","2":"0.574304283"},{"1":"824","2":"0.059057374"},{"1":"825","2":"0.497487366"},{"1":"826","2":"-1.092174796"},{"1":"827","2":"0.418161740"},{"1":"828","2":"-0.085055833"},{"1":"829","2":"0.592299198"},{"1":"830","2":"-0.998836834"},{"1":"831","2":"-0.566930217"},{"1":"832","2":"-0.237226969"},{"1":"833","2":"0.422449299"},{"1":"834","2":"-0.515656325"},{"1":"835","2":"-0.380845237"},{"1":"836","2":"0.324779114"},{"1":"837","2":"-0.132888384"},{"1":"838","2":"-0.677077484"},{"1":"839","2":"0.108494114"},{"1":"840","2":"0.383975674"},{"1":"841","2":"-0.374830480"},{"1":"842","2":"-0.057190873"},{"1":"843","2":"-0.198594583"},{"1":"844","2":"-0.353450404"},{"1":"845","2":"-0.409904739"},{"1":"846","2":"0.534769835"},{"1":"847","2":"-0.195609476"},{"1":"848","2":"0.070824073"},{"1":"849","2":"0.456170129"},{"1":"850","2":"0.826570160"},{"1":"851","2":"0.046111801"},{"1":"852","2":"-0.774474013"},{"1":"853","2":"-0.598771111"},{"1":"854","2":"-0.417561245"},{"1":"855","2":"-0.046458850"},{"1":"856","2":"0.922080107"},{"1":"857","2":"0.337349236"},{"1":"858","2":"-0.077051982"},{"1":"859","2":"-0.864238864"},{"1":"860","2":"0.234957599"},{"1":"861","2":"0.674942800"},{"1":"862","2":"0.447762722"},{"1":"863","2":"-0.350763027"},{"1":"864","2":"0.919932490"},{"1":"865","2":"-0.058735950"},{"1":"866","2":"0.148779869"},{"1":"867","2":"0.100889847"},{"1":"868","2":"0.443307157"},{"1":"869","2":"0.049563409"},{"1":"870","2":"0.159103022"},{"1":"871","2":"1.140653850"},{"1":"872","2":"-0.716715568"},{"1":"873","2":"0.711394655"},{"1":"874","2":"0.645877075"},{"1":"875","2":"0.052474260"},{"1":"876","2":"-0.592771267"},{"1":"877","2":"0.142287943"},{"1":"878","2":"0.083125249"},{"1":"879","2":"0.340218517"},{"1":"880","2":"-0.040032976"},{"1":"881","2":"0.347478942"},{"1":"882","2":"-0.328249396"},{"1":"883","2":"-0.833759745"},{"1":"884","2":"-0.531378524"},{"1":"885","2":"0.101409868"},{"1":"886","2":"0.828161775"},{"1":"887","2":"0.585718174"},{"1":"888","2":"-0.096721571"},{"1":"889","2":"-0.424256012"},{"1":"890","2":"-0.110487313"},{"1":"891","2":"-0.755252257"},{"1":"892","2":"-0.077212812"},{"1":"893","2":"0.845534804"},{"1":"894","2":"-0.086543108"},{"1":"895","2":"-0.464552218"},{"1":"896","2":"0.280018543"},{"1":"897","2":"-0.119286907"},{"1":"898","2":"0.267388668"},{"1":"899","2":"1.044359429"},{"1":"900","2":"0.530504140"},{"1":"901","2":"0.198386936"},{"1":"902","2":"-0.338508156"},{"1":"903","2":"-0.097141368"},{"1":"904","2":"0.415206537"},{"1":"905","2":"0.346245782"},{"1":"906","2":"0.376927320"},{"1":"907","2":"0.020027050"},{"1":"908","2":"0.430670436"},{"1":"909","2":"-0.151655702"},{"1":"910","2":"0.028519156"},{"1":"911","2":"-0.153605785"},{"1":"912","2":"0.070044340"},{"1":"913","2":"0.003364315"},{"1":"914","2":"0.228797784"},{"1":"915","2":"0.464584300"},{"1":"916","2":"-0.458472029"},{"1":"917","2":"0.320556547"},{"1":"918","2":"0.077463698"},{"1":"919","2":"0.172489587"},{"1":"920","2":"-0.235121923"},{"1":"921","2":"-0.462338668"},{"1":"922","2":"-0.408830020"},{"1":"923","2":"-0.146916033"},{"1":"924","2":"-0.127463473"},{"1":"925","2":"-0.277943645"},{"1":"926","2":"0.255540338"},{"1":"927","2":"-0.598872161"},{"1":"928","2":"-0.580927339"},{"1":"929","2":"0.666118167"},{"1":"930","2":"-0.316312806"},{"1":"931","2":"0.768139246"},{"1":"932","2":"-0.176492620"},{"1":"933","2":"0.079136953"},{"1":"934","2":"0.674402760"},{"1":"935","2":"0.147392292"},{"1":"936","2":"-0.116204076"},{"1":"937","2":"0.052139581"},{"1":"938","2":"-0.398412500"},{"1":"939","2":"0.652713767"},{"1":"940","2":"0.232656922"},{"1":"941","2":"0.163493491"},{"1":"942","2":"0.165047754"},{"1":"943","2":"0.278379922"},{"1":"944","2":"0.441329707"},{"1":"945","2":"-0.076440134"},{"1":"946","2":"0.852878837"},{"1":"947","2":"0.045089307"},{"1":"948","2":"-0.184432197"},{"1":"949","2":"0.042595457"},{"1":"950","2":"-0.098287530"},{"1":"951","2":"-0.233302200"},{"1":"952","2":"0.654555950"},{"1":"953","2":"-0.292173565"},{"1":"954","2":"-0.409533598"},{"1":"955","2":"-0.440082673"},{"1":"956","2":"-0.008045861"},{"1":"957","2":"0.113272692"},{"1":"958","2":"0.193678902"},{"1":"959","2":"0.030089141"},{"1":"960","2":"0.029074791"},{"1":"961","2":"0.086290382"},{"1":"962","2":"-0.708833025"},{"1":"963","2":"0.407647015"},{"1":"964","2":"0.013320233"},{"1":"965","2":"-0.244127743"},{"1":"966","2":"-0.522266592"},{"1":"967","2":"0.088078994"},{"1":"968","2":"0.363748901"},{"1":"969","2":"0.442597090"},{"1":"970","2":"0.191441158"},{"1":"971","2":"0.676713689"},{"1":"972","2":"-0.193628565"},{"1":"973","2":"0.273696680"},{"1":"974","2":"0.790558498"},{"1":"975","2":"-0.275414342"},{"1":"976","2":"-0.319033674"},{"1":"977","2":"1.161872339"},{"1":"978","2":"0.172838230"},{"1":"979","2":"0.446869933"},{"1":"980","2":"-0.039996778"},{"1":"981","2":"-0.211061377"},{"1":"982","2":"0.595677566"},{"1":"983","2":"-0.344605680"},{"1":"984","2":"0.188379377"},{"1":"985","2":"-0.046733556"},{"1":"986","2":"-0.559393057"},{"1":"987","2":"-0.240053907"},{"1":"988","2":"-0.082854610"},{"1":"989","2":"0.168311185"},{"1":"990","2":"-0.436535402"},{"1":"991","2":"0.294716845"},{"1":"992","2":"-0.601406080"},{"1":"993","2":"0.071166505"},{"1":"994","2":"-0.077958129"},{"1":"995","2":"0.929354341"},{"1":"996","2":"-0.279616047"},{"1":"997","2":"-0.430309623"},{"1":"998","2":"0.660500632"},{"1":"999","2":"0.446687345"},{"1":"1000","2":"-0.141201218"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] ] --- # The *infer* Pipeline: Get p Value .left-code[ ### Specify ### Hypothesize ### Generate ### Calculate ### Get p Value .quitesmall[ `%>% get_p_value(obs stat = "", direction = "both")` ] ] .right-plot[ .quitesmall[ - We can calculate the .hi[`\\(p\\)`-value] - the probability of seeing a value at least as large as our `sample_slope` (-2.28) in our simulated null distribution - .hi-purple[Two-sided alternative] `\(H_a: \beta_1 \neq 0\)`, we double the raw `\(p\)`-value ```r CASchool %>% specify(testscr ~ str) %>% hypothesize(null = "independence") %>% generate(reps = 1000, type = "permute") %>% calculate(stat = "slope") %>% * get_p_value(obs_stat = sample_slope, * direction = "both") ``` <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["p_value"],"name":[1],"type":["dbl"],"align":["right"]}],"data":[{"1":"0"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> ] ] --- # The *infer* Pipeline: Visualize I .center[  ] --- # The *infer* Pipeline: Visualize I .left-code[ ### Specify ### Hypothesize ### Generate ### Calculate ### Visualize `%>% visualize()` ] .right-plot[ .smallest[ - Make a histogram of our null distribution of `\(\beta_1\)` - Note it is centered at `\(\beta_1=0\)` because that's `\(H_0\)`! ```r CASchool %>% specify(testscr ~ str) %>% hypothesize(null = "independence") %>% generate(reps = 1000, type = "permute") %>% calculate(stat = "slope") %>% * visualize() ``` <img src="2.7-slides_files/figure-html/unnamed-chunk-16-1.png" width="504" style="display: block; margin: auto;" /> ] ] --- # The *infer* Pipeline: Visualize II .left-code[ ### Specify ### Hypothesize ### Generate ### Calculate ### Visualize `%>% visualize()` ] .right-plot[ .smallest[ - Add our `sample_slope` to show our finding on the null distr. ```r CASchool %>% specify(testscr ~ str) %>% hypothesize(null = "independence") %>% generate(reps = 1000, type = "permute") %>% calculate(stat = "slope") %>% * visualize(obs_stat = sample_slope) ``` <img src="2.7-slides_files/figure-html/unnamed-chunk-17-1.png" width="504" style="display: block; margin: auto;" /> ] ] --- # The *infer* Pipeline: Visualize p-value .left-code[ ### Specify ### Hypothesize ### Generate ### Calculate ### Visualize `%>% visualize()+shade_p_value()` ] .right-plot[ .smallest[ - Add `shade_p_value()` to see what `\(p\)` is ```r CASchool %>% specify(testscr ~ str) %>% hypothesize(null = "independence") %>% generate(reps = 1000, type = "permute") %>% calculate(stat = "slope") %>% visualize(obs_stat = sample_slope)+ * shade_p_value(obs_stat = sample_slope, * direction = "two_sided") ``` <img src="2.7-slides_files/figure-html/unnamed-chunk-18-1.png" width="504" style="display: block; margin: auto;" /> ] ] --- # The *infer* Pipeline: Visualize is a Wrapper of ggplot - `infer`'s `visualize()` function is just a wrapper function for `ggplot()` - you can take your `simulations` `tibble` and just `ggplot` a normal histogram -- .pull-left[ .code50[ ```r simulations %>% ggplot(data = .)+ aes(x = stat)+ geom_histogram(color="white", fill="#e64173")+ geom_vline(xintercept = sample_slope, color = "blue", size = 2, linetype = "dashed")+ annotate(geom = "label", x = -2.28, y = 100, label = expression(paste("Our ", hat(beta[1]))), color = "blue")+ scale_y_continuous(lim=c(0,120), expand = c(0,0))+ labs(x = expression(paste("Sampling distribution of ", hat(beta)[1], " under ", H[0], ": ", beta[1]==0)), y = "Samples")+ theme_classic(base_family = "Fira Sans Condensed", base_size=20) ``` ] ] -- .pull-right[ <img src="2.7-slides_files/figure-html/unnamed-chunk-19-1.png" width="504" style="display: block; margin: auto;" /> ] --- class: inverse, center, middle # Classical Statistical Inference (What R Calculates) --- # What R Does: Classical Statistical Inference I .pull-left[ - R does things the old-fashioned way, using a *theoretical* null distribution instead of *simulating* one - A .hi[`\\(t\\)`-distribution] with `\(n-k-1\)` df<sup>.magenta[†]</sup> - Calculate a `\(t\)`-statistic for `\(\hat{\beta_1}\)`: `$$\text{test statistic} = \frac{\text{estimate} - \text{null hypothesis}}{\text{standard error of estimate}}$$` ] .pull-right[ <img src="2.7-slides_files/figure-html/unnamed-chunk-20-1.png" width="504" style="display: block; margin: auto;" /> ] .footnote[<sup>.magenta[†]</sup> `\\(k\\)` is the number of `\\(X\\)` variables.] --- # What R Does: Classical Statistical Inference II .pull-left[ .smallest[ `$$\text{test statistic} = \frac{\text{estimate} - \text{null hypothesis}}{\text{standard error of estimate}}$$` - `\(t\)` same interpretation as `\(Z\)`: number of std. dev. away from the sampling distribution's expected value `\(E[\hat{\beta_1}]\)`<sup>.magenta[†]</sup> (if `\(H_0\)` were true) - Compares to a .hi-purple[critical value] of `\(t^*\)` (pre-determined by `\(\alpha\)`-level & `\(n-k-1\)` df) - For 95% confidence, `\(\alpha=0.05\)`, `\(t^* \approx 2\)`<sup><span class="magenta">‡</span></sup> ] ] .pull-right[ <img src="2.7-slides_files/figure-html/unnamed-chunk-21-1.png" width="504" style="display: block; margin: auto;" /> ] .footnote[<sup>.magenta[†]</sup> The expected value is 0, because our null hypothesis was `\(\beta_1 =0\)` <sup>.magenta[‡]</sup> Again, the **68-95-99.7%** empirical rule!] --- # What R Does: Classical Statistical Inference III .pull-left[ .quitesmall[ `$$\begin{align*} t &= \frac{\hat{\beta_1}-\beta_{1,0}}{se(\hat{\beta_1})}\\ t &= \frac{-2.28-0}{0.48}\\ t &= -4.75\\ \end{align*}$$` - Our sample slope `\(\hat{\beta_1}\)` is **4.75 standard deviations below** the expected value `\(E[\hat{\beta_1}]\)` (i.e. 0) if `\(H_0\)` were true ] ] .pull-right[ <img src="2.7-slides_files/figure-html/unnamed-chunk-22-1.png" width="504" style="display: block; margin: auto;" /> ] --- # What R Does: Classical Statistical Inference III .pull-left[ .quitesmall[ `$$\begin{align*} t &= \frac{\hat{\beta_1}-\beta_{1,0}}{se(\hat{\beta_1})}\\ t &= \frac{-2.28-0}{0.48}\\ t &= -4.75\\ \end{align*}$$` - .hi[p-value]: prob. of a test statistic at least as large (in magnitude) as ours if the null hypothesis were true - Continuous distribution implies we need probability of area *beyond* our value - p-value is **2-sided** for `\(H_a: \beta_1 \neq 0\)` - `\(2 \times p(t_{418}> \vert -4.75\vert)=0.0000028\)` ] ] .pull-right[ <img src="2.7-slides_files/figure-html/unnamed-chunk-23-1.png" width="504" style="display: block; margin: auto;" /> ] --- # 1-Sided Tests & p-values .pull-left[ `\(H_a: \beta_1<0\)` p-value: `\(p(t \leq t_i)\)` <img src="2.7-slides_files/figure-html/unnamed-chunk-24-1.png" width="504" style="display: block; margin: auto;" /> ] .pull-right[ `\(H_a: \beta_1>0\)` p-value: `\(p(t \geq t_i)\)` <img src="2.7-slides_files/figure-html/unnamed-chunk-25-1.png" width="504" style="display: block; margin: auto;" /> ] --- # 2-Sided Tests and p-values `\(H_a: \beta_1 \neq 0\)` p-value: `\(2 \times p(t \geq |t_i|)\)` <img src="2.7-slides_files/figure-html/unnamed-chunk-26-1.png" width="504" style="display: block; margin: auto;" /> --- # Calculating p-values in R .pull-left[ .smallest[ - `pt()` calculates `p`robabilities on a `t` distribution with arguments: - the t-score - `df = ` the degrees of freedom - `lower.tail = ` - `TRUE` if looking at area to *LEFT* of value - `FALSE` if looking at area to *RIGHT* of value ```r 2 * pt(4.75, # I'll double the right tail df = 418, lower.tail = F) # right tail ``` ``` ## [1] 2.800692e-06 ``` - `\(2 \times p(t_{418}> \vert -4.75\vert)=0.0000028\)` ] ] .pull-right[ <img src="2.7-slides_files/figure-html/unnamed-chunk-28-1.png" width="504" style="display: block; margin: auto;" /> ] --- # Hypothesis Tests in Regression Output I ```r summary(school_reg) ``` ``` ## ## Call: ## lm(formula = testscr ~ str, data = CASchool) ## ## Residuals: ## Min 1Q Median 3Q Max ## -47.727 -14.251 0.483 12.822 48.540 ## ## Coefficients: ## Estimate Std. Error t value Pr(>|t|) ## (Intercept) 698.9330 9.4675 73.825 < 2e-16 *** ## str -2.2798 0.4798 -4.751 2.78e-06 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## Residual standard error: 18.58 on 418 degrees of freedom ## Multiple R-squared: 0.05124, Adjusted R-squared: 0.04897 ## F-statistic: 22.58 on 1 and 418 DF, p-value: 2.783e-06 ``` --- # Hypothesis Tests in Regression Output II - In `broom`'s `tidy()` (with confidence intervals) ```r tidy(school_reg, conf.int=TRUE) ``` <div data-pagedtable="false"> <script data-pagedtable-source type="application/json"> {"columns":[{"label":["term"],"name":[1],"type":["chr"],"align":["left"]},{"label":["estimate"],"name":[2],"type":["dbl"],"align":["right"]},{"label":["std.error"],"name":[3],"type":["dbl"],"align":["right"]},{"label":["statistic"],"name":[4],"type":["dbl"],"align":["right"]},{"label":["p.value"],"name":[5],"type":["dbl"],"align":["right"]},{"label":["conf.low"],"name":[6],"type":["dbl"],"align":["right"]},{"label":["conf.high"],"name":[7],"type":["dbl"],"align":["right"]}],"data":[{"1":"(Intercept)","2":"698.932952","3":"9.4674914","4":"73.824514","5":"6.569925e-242","6":"680.32313","7":"717.542779"},{"1":"str","2":"-2.279808","3":"0.4798256","4":"-4.751327","5":"2.783307e-06","6":"-3.22298","7":"-1.336637"}],"options":{"columns":{"min":{},"max":[10]},"rows":{"min":[10],"max":[10]},"pages":{}}} </script> </div> - p-value on `str` is 0.00000278. --- # Conclusions `$$\begin{align*} H_0: \,& \beta_1=0\\ H_a: \, & \beta_a \neq 0\\ \end{align*}$$` - Because the hypothesis test's `\(p\)`-value `\(<\)` `\(\alpha\)` (0.05)... - .hi-purple[We have sufficient evidence to reject `\\(H_0\\)` in favor of our alternative hypothesis. Our sample suggests that there *is a relationship* between class size and test scores]. -- - Using the confidence intervals: - .hi-purple[We are 95% confident that, from similarly constructed samples, the true marginal effect of class size on test scores is between -3.22 and -1.34.] --- # Hypothesis Testing vs. Confidence Intervals .smaller[ - Confidence intervals are all *two-sided* by nature `$$CI_{0.95}=\left(\left[\hat{\beta_1}-\underbrace{2 \times se(\hat{\beta_1})}_{MOE}\right], \, \left[\hat{\beta_1}+\underbrace{2 \times se(\hat{\beta_1})}_{MOE}\right] \right)$$` - Hypothesis test `\((t\)`-test) of `\(H_0: \, \beta_1=0\)` computes a `\(t\)`-value of<sup>.red[1]</sup> `$$t=\frac{\hat{\beta_1}}{se(\hat{\beta_1})}$$` and `\(p<0.05\)` when `\(t\geq2\)` (approximately) ] .footnote[<sup>.red[1]</sup> Since our null hypothesis is that `\\(\beta_{1,0}=0\\)`, the test statistic simplifies to this neat fraction.] -- .smaller[ - .hi-purple[If our confidence interval contains the `\\(H_0\\)` value (i.e. `\\(0\\)`, for our test), then *we fail to reject* `\\(H_0\\)`.] ] --- class: inverse, center, middle # The Use and Abuse of `\(p\)`-values --- # p-H .center[  ] .source[Source: [XKCD](https://xkcd.com/882/)] --- # p-Hacking .center[  ] .source[Source: [XKCD](https://xkcd.com/882/)] --- # p-Hacking .center[  ] .source[Source: [XKCD](https://xkcd.com/882/)] --- # p-Hacking .center[  ] .source[Source: [XKCD](https://xkcd.com/882/)] --- # p-Hacking .pull-left[ - Consider what 95% confident or `\(\alpha=0.05\)` means - If we repeat a procedure 20 times, we should expect `\(\frac{1}{20}\)` (5%) to produce a fluke result! .source[Image source: [Seeing Theory](https://seeing-theory.brown.edu/frequentist-inference/index.html)] ] .pull-right[ .center[ 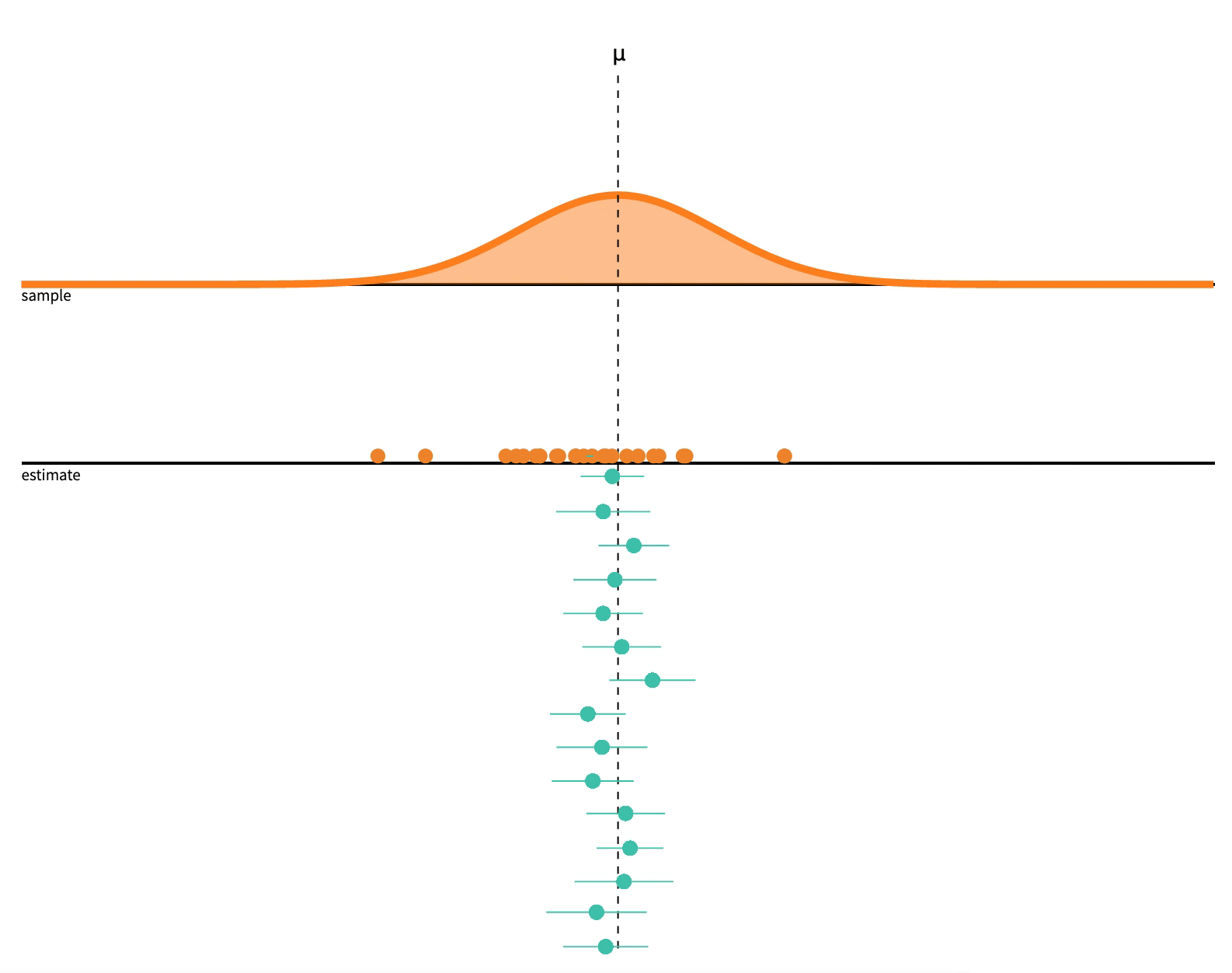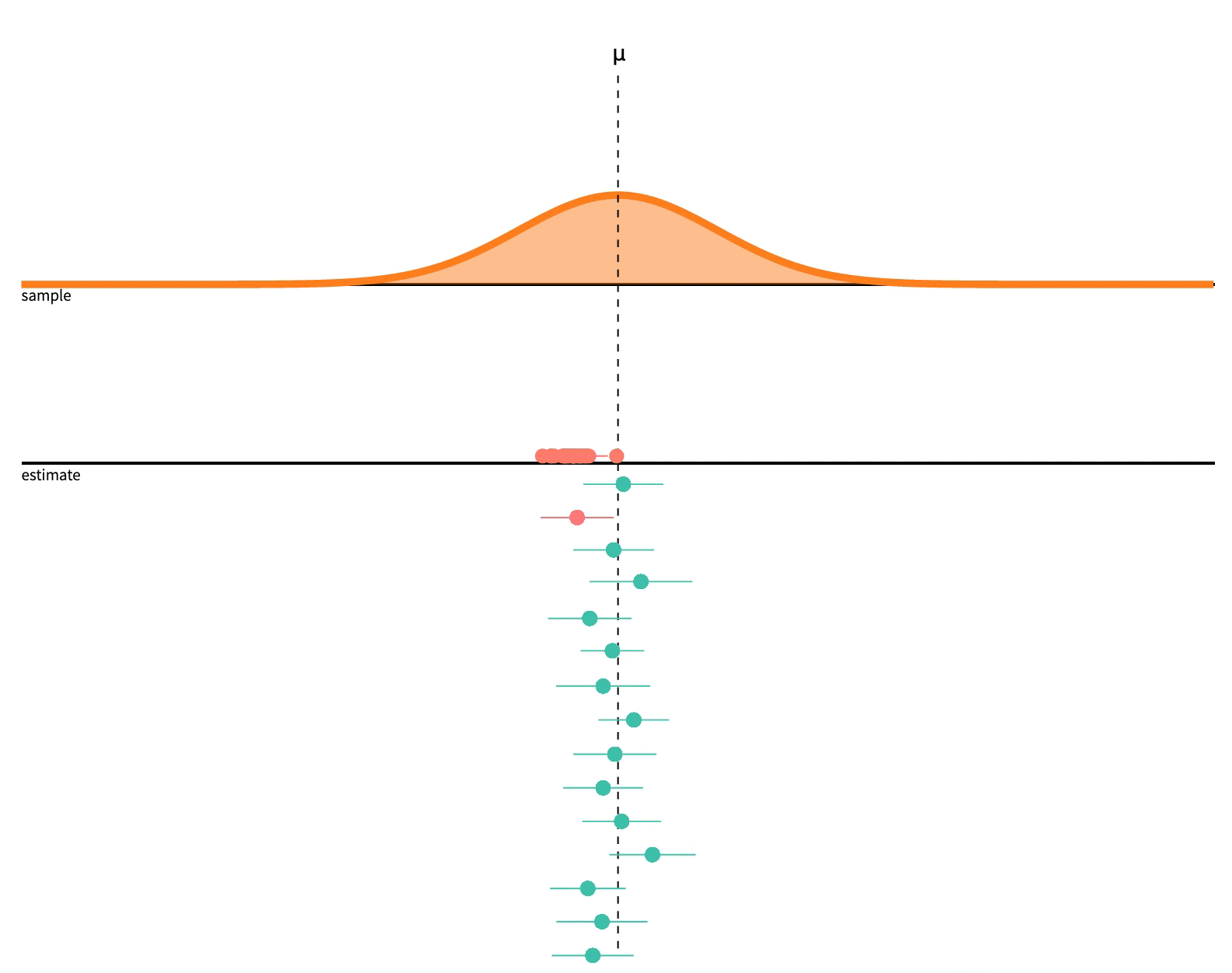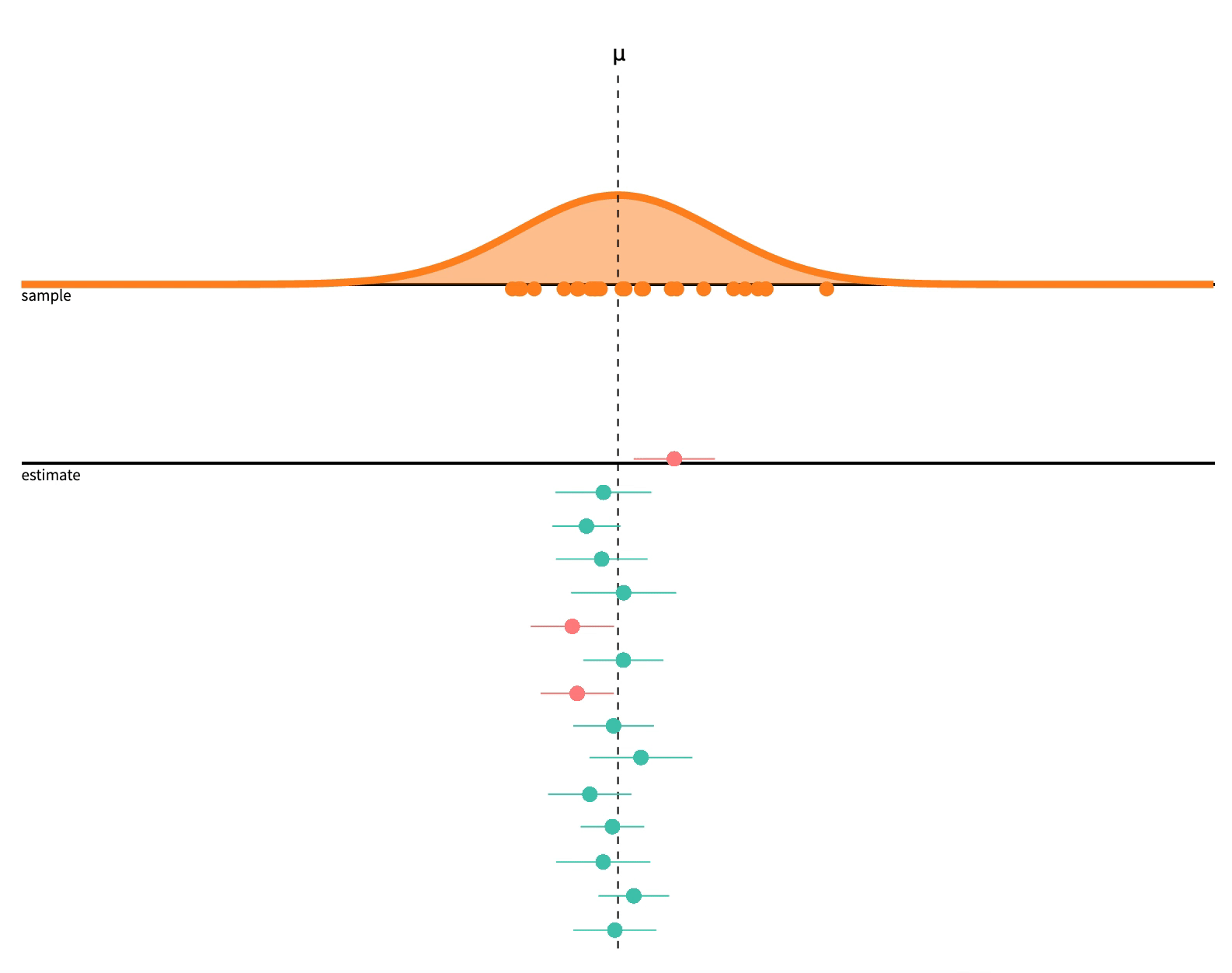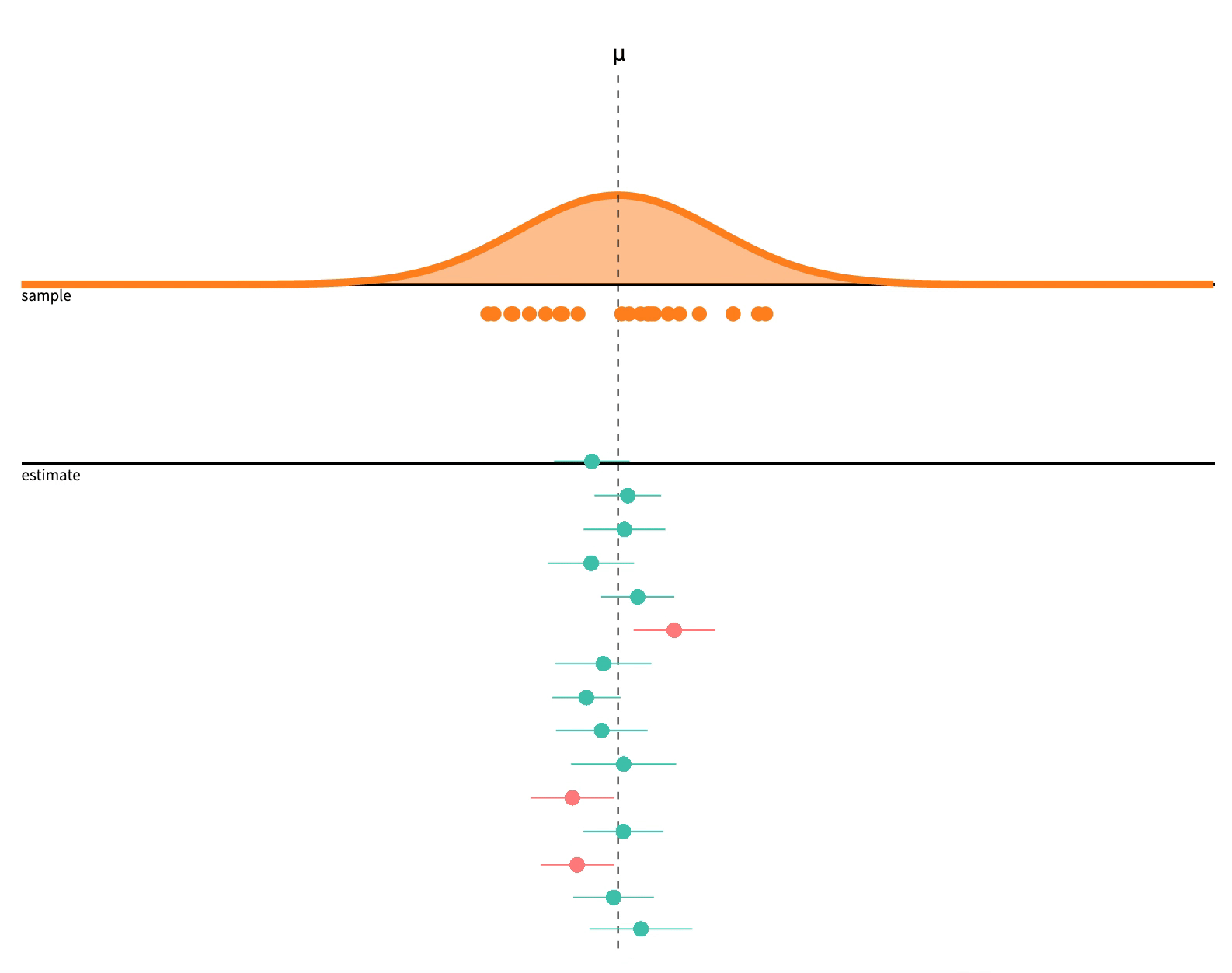 ] ] --- # Abusing p-values and “Science” .center[  ] .source[Source: [Washington Post](https://www.washingtonpost.com/news/morning-mix/wp/2015/05/28/how-and-why-a-journalist-tricked-news-outlets-into-thinking-chocolate-makes-you-thin/?hpid=z5)] --- # Abusing p-Values and “Science” .pull-left[ .center[  ] ] -- .pull-right[ .center[  ] ] .source[Source: [SMBC](http://www.smbc-comics.com/?id=1623)] --- # Abusing p-Values and “Science” .pull-left[ .center[  ] ] .pull-right[ > “The widespread use of 'statistical significance' (generally interpreted as `\((p \leq 0.05)\)` as a license for making a claim of a scientific finding (or implied truth) leads to considerable distortion of the scientific process.” ] .source[Wasserstein, Ronald L. and Nicole A. Lazar, (2016), ["The ASA's Statement on p-Values: Context, Process, and Purpose](http://www.tandfonline.com/doi/full/10.1080/00031305.2016.1154108)," *The American Statistician* 30(2): 129-133] --- # Abusing p-Values and “Science” .left-column[ .center[  ] ] .right-column[ > “No economist has achieved scientific success as a result of a statistically significant coefficient. Massed observations, clever common sense, elegant theorems, new policies, sagacious economic reasoning, historical perspective, relevant accounting, these have all led to scientific success. Statistical significance has not,” (p.112). .source[McCloskey, Dierdre N and Stephen Ziliak, 1996, *The Cult of Statistical Significance*] ] --- # Common Misconceptions about p-values .smallest[ ❌ `\(p\)` **is the probability that the alternative hypothesis is false** - We can never *prove* an alternative hypothesis, only tentatively reject a null hypothesis ] -- .smallest[ ❌ `\(p\)` **is the probability that the null hypothesis is true** - We're not *proving* the `\(H_0\)` is false, only saying that it's very unlikely that if `\(H_0\)` were true, we'd obtain a slope as rare as our sample's slope ] -- .smallest[ ❌ `\(p\)` **is the probability that our observed effects were produced purely by random chance** - `\(p\)` is computed under a specific model (think about our null world) that *assumes* `\(H_0\)` is true ] -- .smallest[ ❌ `\(p\)` **tells us how significant our finding is** - `\(p\)` tells us nothing about the *size* or the *real world significance* of any effect deemed “statistically significant” - it only tells us that the slope is statistically significantly different from 0 (if `\(H_0\)` is `\(\beta_1=0)\)` ] --- # p-value Clarification - Again, .hi[p-value is the probability that, if the null hypothesis were true, we obtain (by pure random chance) a test statistic at least as extreme as the one we estimated for our sample] - A low p-value means either (and we can't distinguish which): 1. `\(H_0\)` is true and a highly improbable event has occurred OR 2. `\(H_0\)` is false --- # Significance In Regression Tables .pull-left[ .font80[ .regtable[ <table class="huxtable" style="border-collapse: collapse; border: 0px; margin-bottom: 2em; margin-top: 2em; ; margin-left: auto; margin-right: auto; " id="tab:unnamed-chunk-31"> <col><col><tr> <th style="vertical-align: top; text-align: center; white-space: normal; border-style: solid solid solid solid; border-width: 0.8pt 0pt 0pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;"></th><th style="vertical-align: top; text-align: center; white-space: normal; border-style: solid solid solid solid; border-width: 0.8pt 0pt 0.4pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">Test Score</th></tr> <tr> <th style="vertical-align: top; text-align: left; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">Intercept</th><td style="vertical-align: top; text-align: right; white-space: normal; border-style: solid solid solid solid; border-width: 0.4pt 0pt 0pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">698.93 ***</td></tr> <tr> <th style="vertical-align: top; text-align: left; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;"></th><td style="vertical-align: top; text-align: right; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">(9.47) </td></tr> <tr> <th style="vertical-align: top; text-align: left; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">STR</th><td style="vertical-align: top; text-align: right; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">-2.28 ***</td></tr> <tr> <th style="vertical-align: top; text-align: left; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;"></th><td style="vertical-align: top; text-align: right; white-space: normal; border-style: solid solid solid solid; border-width: 0pt 0pt 0.4pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">(0.48) </td></tr> <tr> <th style="vertical-align: top; text-align: left; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">N</th><td style="vertical-align: top; text-align: right; white-space: normal; border-style: solid solid solid solid; border-width: 0.4pt 0pt 0pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">420 </td></tr> <tr> <th style="vertical-align: top; text-align: left; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">R-Squared</th><td style="vertical-align: top; text-align: right; white-space: normal; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">0.05 </td></tr> <tr> <th style="vertical-align: top; text-align: left; white-space: normal; border-style: solid solid solid solid; border-width: 0pt 0pt 0.8pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">SER</th><td style="vertical-align: top; text-align: right; white-space: normal; border-style: solid solid solid solid; border-width: 0pt 0pt 0.8pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;">18.58 </td></tr> <tr> <th colspan="2" style="vertical-align: top; text-align: left; white-space: normal; border-style: solid solid solid solid; border-width: 0.8pt 0pt 0pt 0pt; padding: 6pt 6pt 6pt 6pt; font-weight: normal;"> *** p < 0.001; ** p < 0.01; * p < 0.05.</th></tr> </table> ] ] ] .pull-right[ - Statistical significance is shown by asterisks, common (but not always!) standard: - 1 asterisk: significant at `\(\alpha=0.10\)` - 2 asterisks: significant at `\(\alpha=0.05\)` - 3 asterisks: significant at `\(\alpha=0.01\)` - Rare, but sometimes regression tables include `\(p\)`-values for estimates ]